Search Count: 29
All
Selected
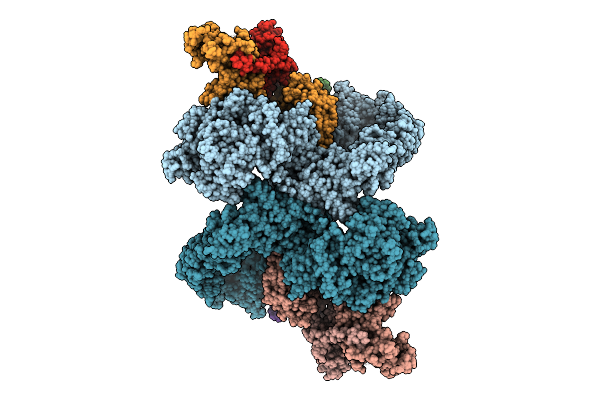 |
Cryo-Em Structure Of The Pi4Ka Complex Bound To An Efr3 Interfering Nanobody (F3In)
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2025-12-03 Classification: SIGNALING PROTEIN |
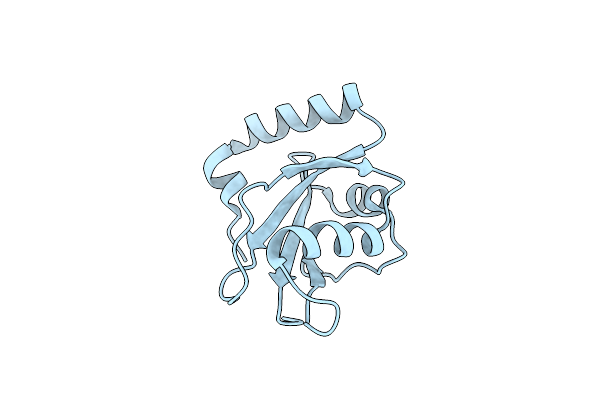 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2025-01-22 Classification: SIGNALING PROTEIN Ligands: CL |
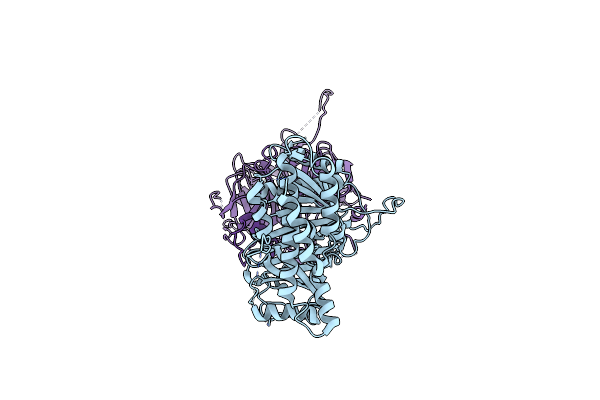 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: CYTOSOLIC PROTEIN |
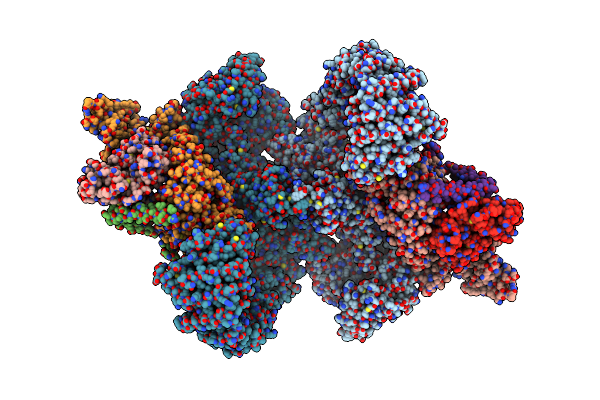 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-27 Classification: SIGNALING PROTEIN |
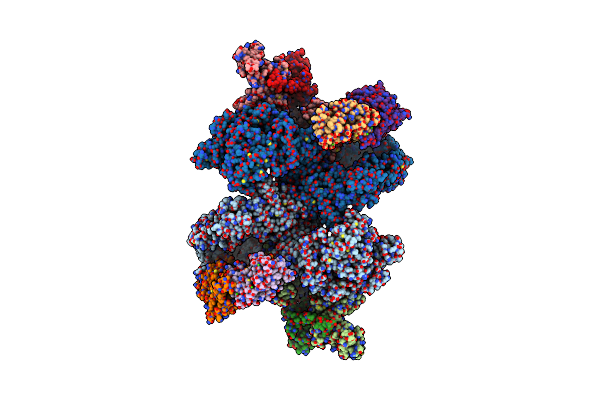 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-09-11 Classification: SIGNALING PROTEIN Ligands: CA |
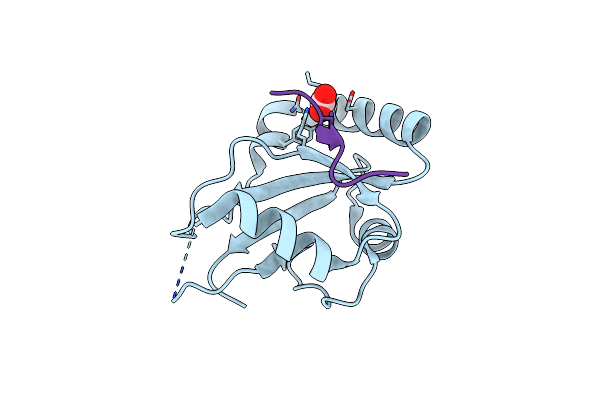 |
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2024-07-17 Classification: SIGNALING PROTEIN Ligands: ACT |
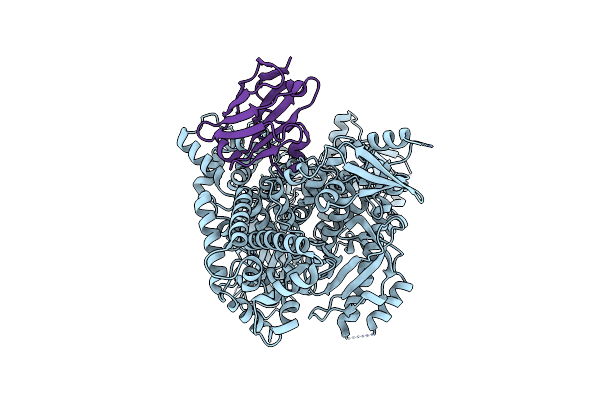 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2023-07-19 Classification: TRANSFERASE/IMMUNE SYSTEM |
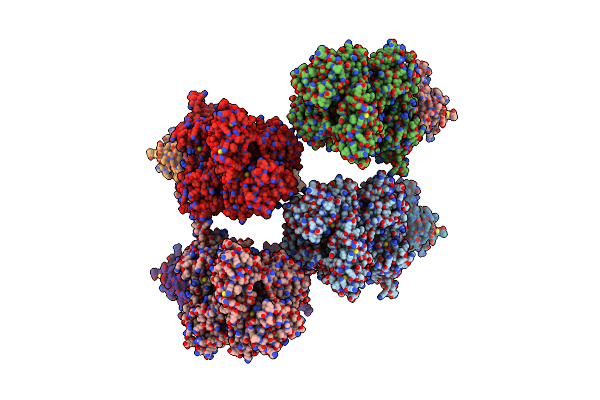 |
Organism: Sus scrofa, Mus musculus
Method: X-RAY DIFFRACTION Resolution:8.50 Å Release Date: 2023-03-22 Classification: SIGNALING PROTEIN |
 |
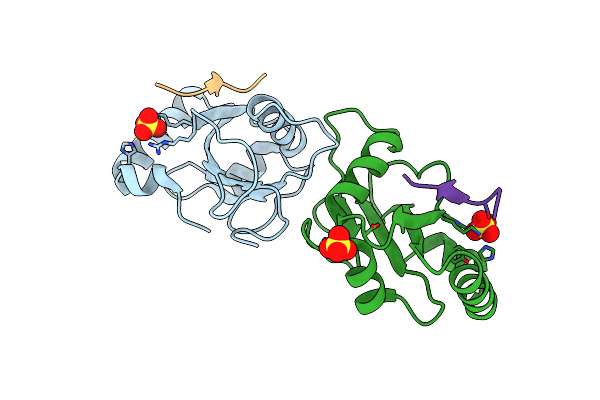
Structure Of The Phosphoinositide 3-Kinase P110 Gamma (Pik3Cg) P101 (Pik3R5) Complex
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2021-07-14 Classification: TRANSFERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2021-03-17 Classification: SIGNALING PROTEIN Ligands: SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2018-04-04 Classification: HORMONE RECEPTOR Ligands: C6V, GOL |
 |
Organism: Human rhinovirus 2
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2016-06-22 Classification: TRANSFERASE Ligands: GOL, PG4, PGE, HOV |
 |
Organism: Human rhinovirus 2
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2016-06-22 Classification: TRANSFERASE Ligands: 6OY |
 |
Organism: Schizophyllum commune h4-8
Method: SOLUTION NMR Release Date: 2016-03-16 Classification: PROTEIN FIBRIL |
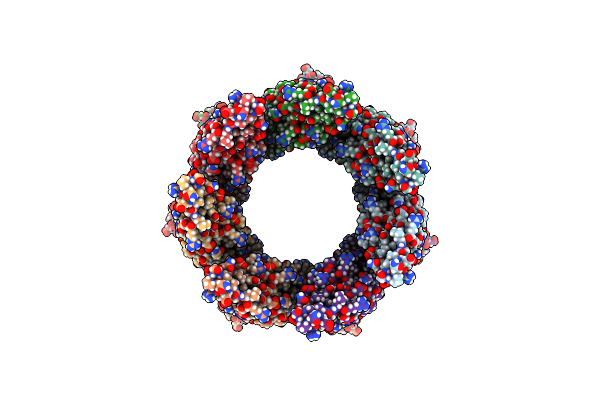 |
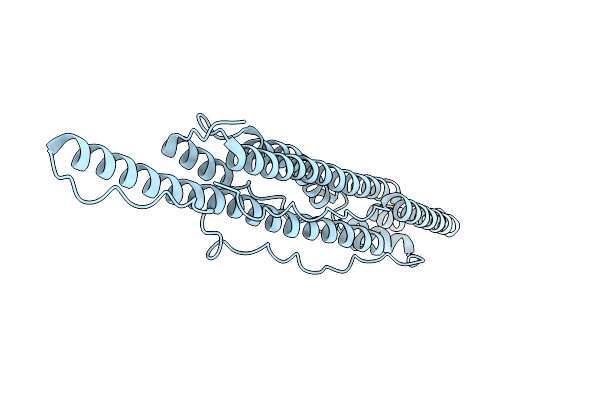
Organism: Mycobacterium tuberculosis h37rv
Method: ELECTRON MICROSCOPY Release Date: 2015-03-11 Classification: PROTEIN BINDING |
 |
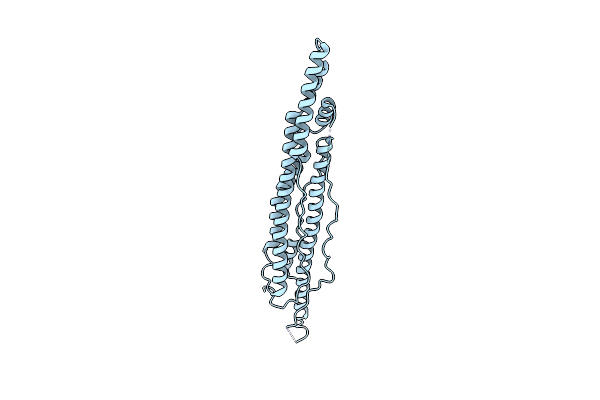
Organism: Mycobacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2015-02-25 Classification: PROTEIN TRANSPORT |
 |
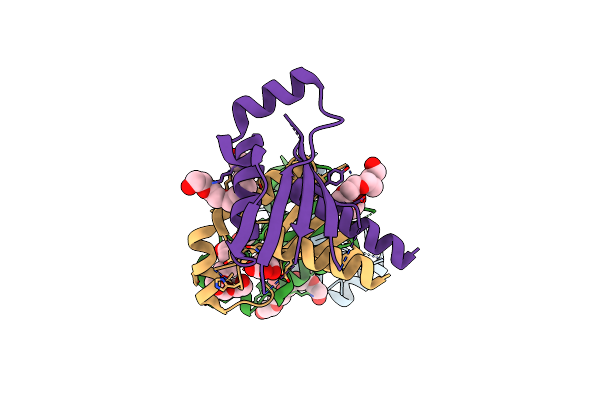
Organism: Mycobacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2015-02-25 Classification: UNKNOWN FUNCTION |
 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2010-09-01 Classification: REPLICATION Ligands: 1PE, MLI |
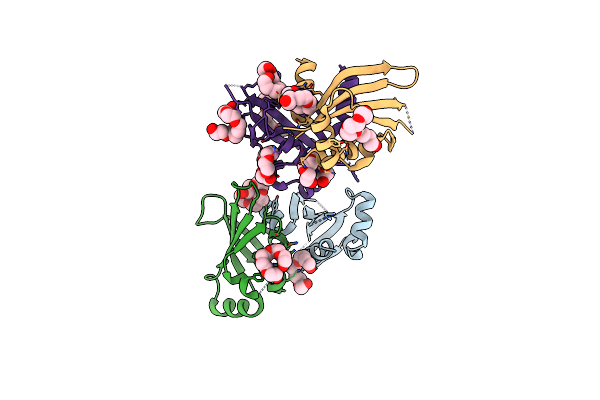 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2010-09-01 Classification: REPLICATION Ligands: 1PE |
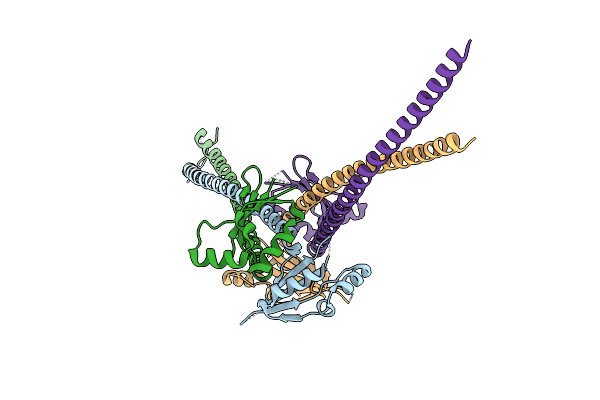 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:3.41 Å Release Date: 2010-09-01 Classification: REPLICATION |

