Search Count: 266
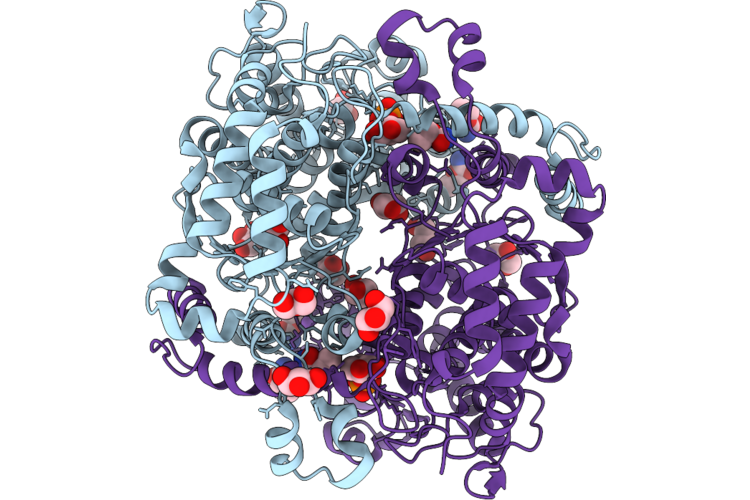 |
Organism: Candida albicans
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: GOL, A1JCW, A1JCV, CL, NA |
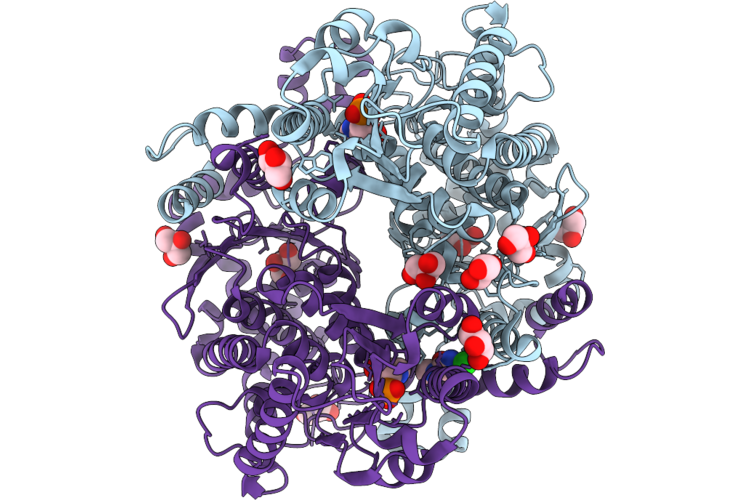 |
Organism: Candida albicans
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: GOL, A1JCV, CL, XKD |
 |
Organism: Candida albicans
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: GOL, A1JCV, XH1, CL |
 |
The Structure Of Aspergillus Fumigatus Phosphoglucose Isomerase In Complex With Fragments
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-05-13 Classification: ISOMERASE Ligands: XKD, GOL, PA5, CL, NA |
 |
The Structure Of Aspergillus Fumigatus Phosphoglucose Isomerase In Complex With An Inhibitor
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-05-13 Classification: ISOMERASE Ligands: GOL, PA5 |
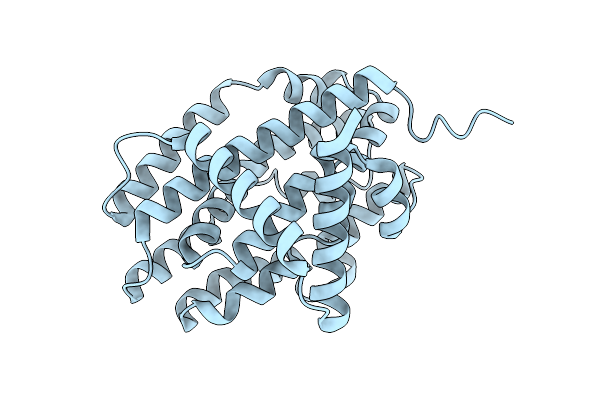 |
Crystal Structure Of Selenomethionine-Substituted N-Oxygenase Rohs From Pseudomonas Syringae Pv. Tomato Str. Dc3000 (Pstorohs)
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN |
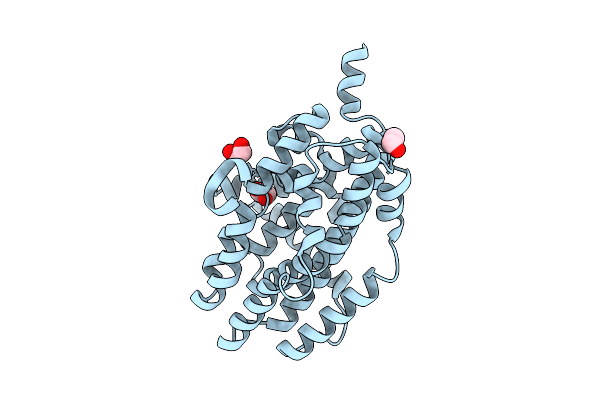 |
Crystal Structure Of Native Hdo-Family N-Oxygenase Rohs From Pseudomonas Brassicacearum Strain Df41 (Pbrrohs)
Organism: Pseudomonas brassicacearum
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO |
 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: GN1, CL |
 |
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (2.5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, HEM, ONH |
 |
N-Hydroxylamine Dehydratase (Nohd) T98A/K167A Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine (5H Soak)
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, ONH |
 |
N-Hydroxylamine Dehydratase (Nohd) R144Y/V98A (A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144A (P1/P2/A1) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/R144Y/V96A (P1/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, GOL, SO4 |
 |
N-Hydroxylamine Dehydratase (Nohd) A59N/R144Y/V96A (P2/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144Y/V96A (P1/P2/A1/A2) Mutant Crystal Structure With Heme
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, EDO |
 |
N-Hydroxylamine Dehydratase (Nohd) H2F/F4P/P5S/R6Y/A59N/R144Y/V96A/D172E/L172Q/D173P/R176V (P1/P2/A1/A2/A3) Mutant Crystal Structure With Heme And N-Hydroxylated Ornithine
Organism: Actinomadura luzonensis
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-12-03 Classification: BIOSYNTHETIC PROTEIN Ligands: HEM, BTB, ONH |
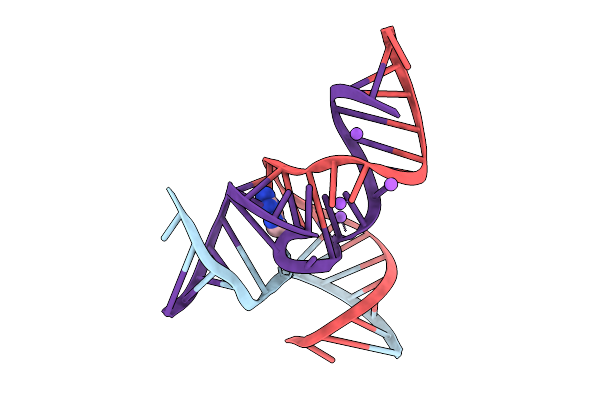 |
Crystal Structure Of The Methyltransferase-Ribozyme 1 Bound To Dna Substrate (With 1-Methyl-Adenosine)
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2025-09-17 Classification: DNA-RNA HYBRID |
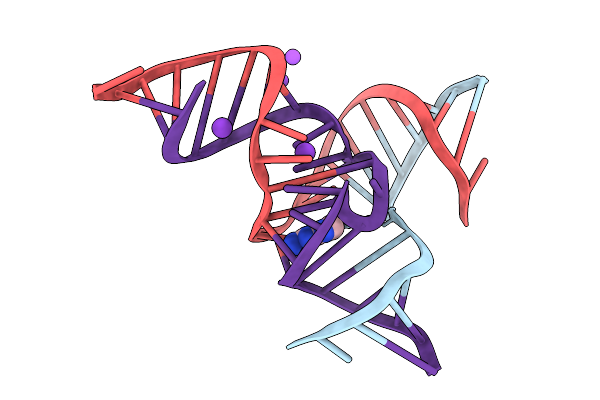 |
Crystal Structure Of The Methyltransferase-Ribozyme 1 Bound To Dna Substrate (1-Benzyl-Adenosine Derivative)
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2025-09-17 Classification: DNA-RNA HYBRID |
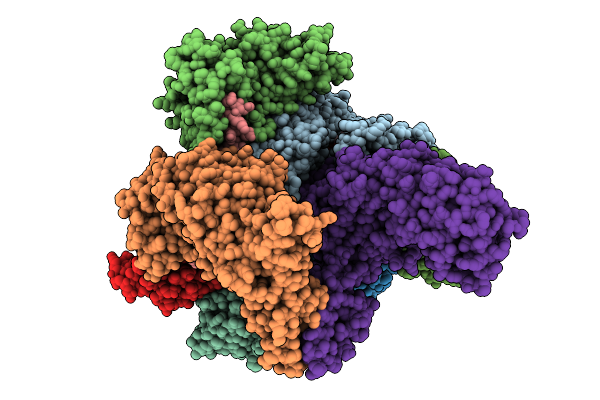 |
Structure Of Spin90 Dimer-Arp2/3 Complex-Nucleated Actin Filament (Singlet Complex)
Organism: Homo sapiens, Bos taurus, Gallus gallus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: CYTOSOLIC PROTEIN Ligands: MG, ADP |

