Search Count: 29
All
Selected
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
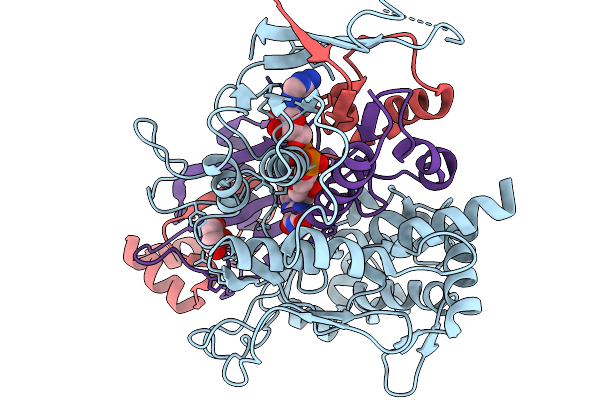 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
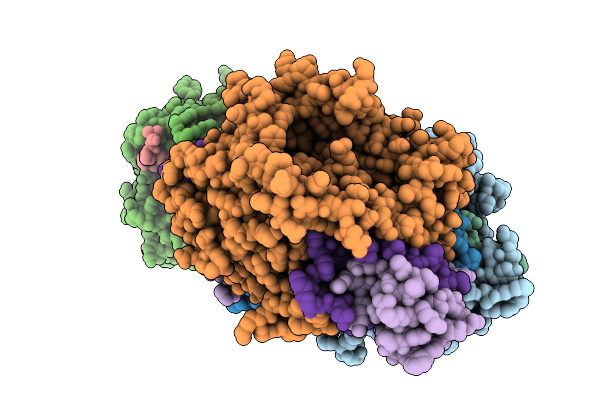 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
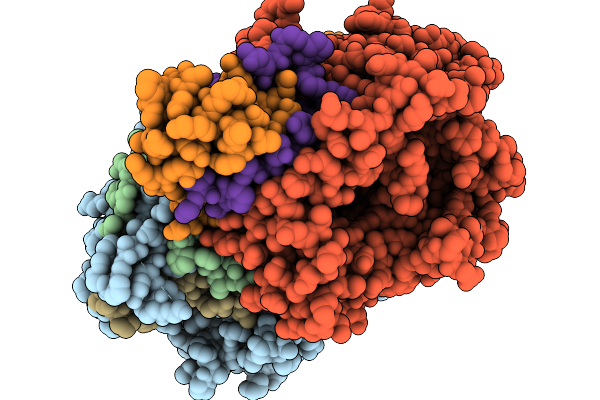 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:3.27 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
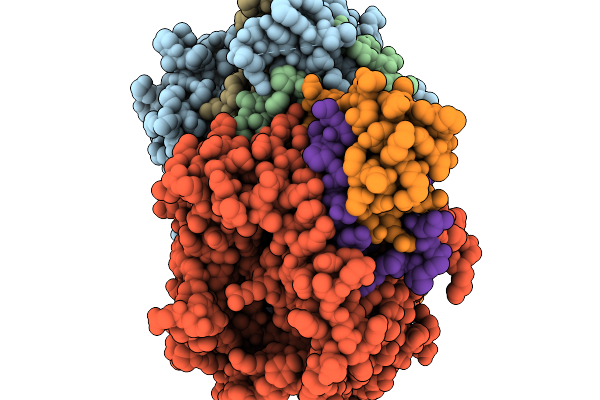 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Cecembia lonarensis lw9
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-01-18 Classification: HYDROLASE |
 |
Organism: Cecembia lonarensis lw9
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2023-01-18 Classification: HYDROLASE |
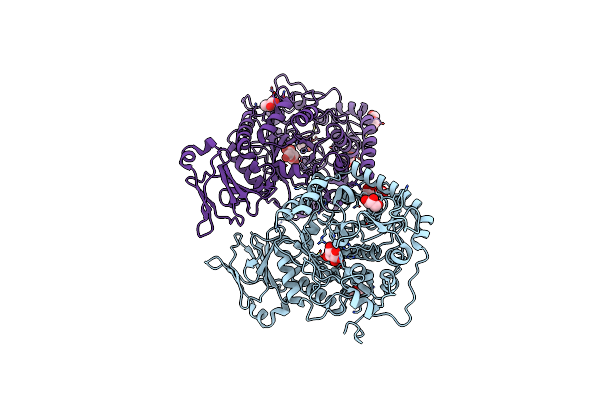 |
Organism: Cecembia lonarensis lw9
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2023-01-18 Classification: HYDROLASE Ligands: Z8T, NA, GOL, CIT, PEG |
 |
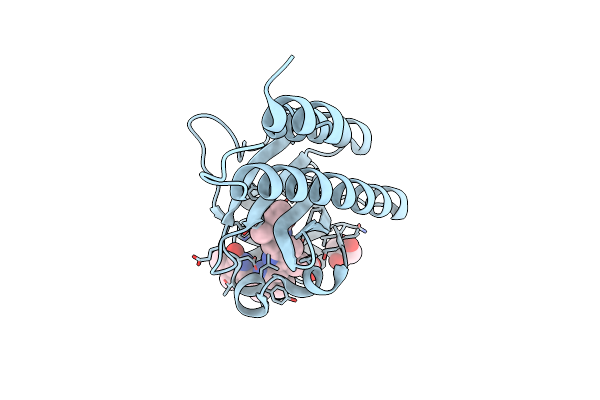
Crystal Structure Of Rorgt With 3-Cyano-N-(3-{[(3S)-4-(Cyclopentanecarbonyl)-3-Methylpiperazin-1-Yl]Methyl}-5-Fluoro-2-Methylphenyl)Benzamide (Compound 1)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2020-03-04 Classification: PROTEIN BINDING Ligands: LKY, EDO |
 |
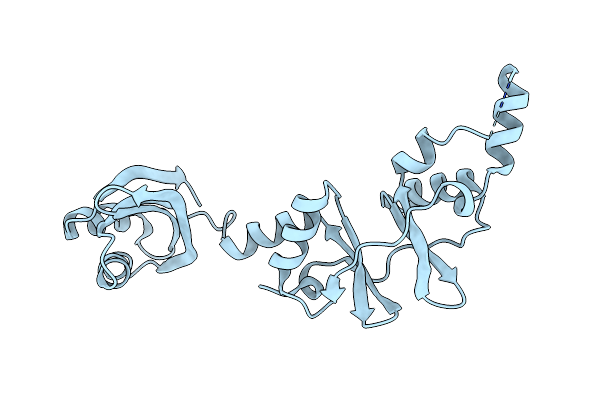
Crystal Structure Of A Far-Red Light-Absorbing Form Of Anpixjg2_Bv4 In Complex With Biliverdin
Organism: Nostoc sp.
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2019-04-17 Classification: SIGNALING PROTEIN Ligands: BLA, GOL |
 |
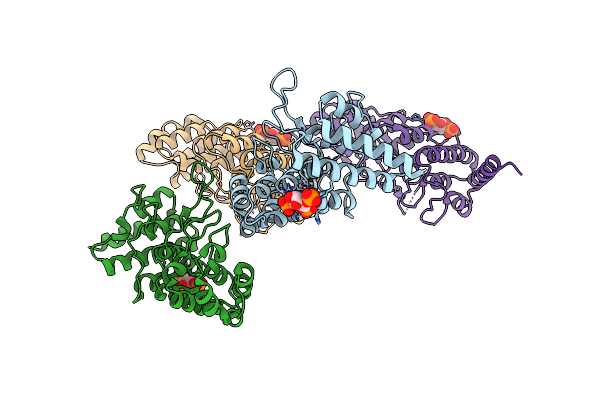
Crystal Structure Of A Hypothetical Protein, Ttha0829 From Thermus Thermophilus Hb8, Composed Of Cystathionine-Beta-Synthase (Cbs) And Aspartate-Kinase Chorismate-Mutase Tyra (Act) Domains
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2016-05-18 Classification: STRUCTURAL GENOMICS, UNKNOWN FUNCTION |
 |
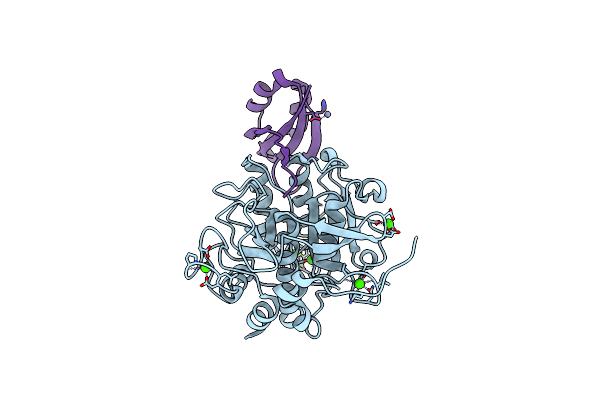
Crystal Structure Of The Ankyrin Repeat Domain Of Chicken Trpv4 In Complex With Ip3
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2014-04-09 Classification: TRANSPORT PROTEIN Ligands: I3P |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2014-04-09 Classification: TRANSPORT PROTEIN |
 |
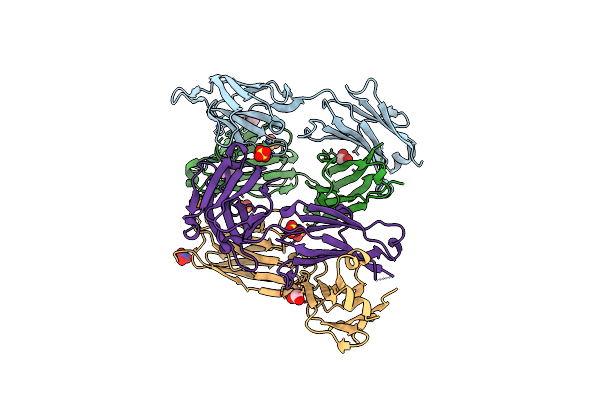
Crystal Structure Of Complex Form Between S324A-Subtilisin And Mutant Tkpro
Organism: Thermococcus kodakarensis
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2013-03-06 Classification: HYDROLASE Ligands: CA, CL, ZN |
 |
Structural Characteristics Of Enantiomorphic Dna: Crystal Analysis Of Racemates Of The D(Cgcgcg) Duplex
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2011-07-13 Classification: DNA |
 |
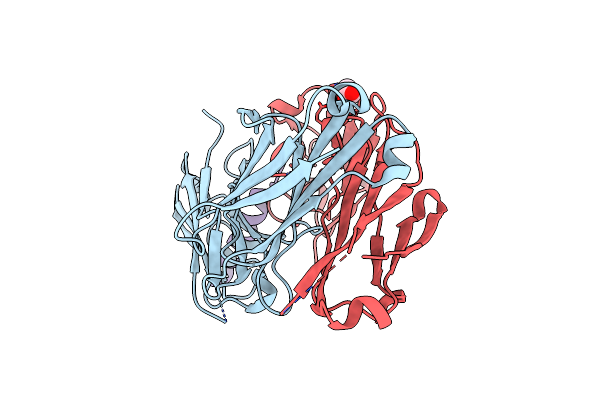
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2010-11-17 Classification: IMMUNE SYSTEM Ligands: SO4, EDO, CL, NO3 |
 |
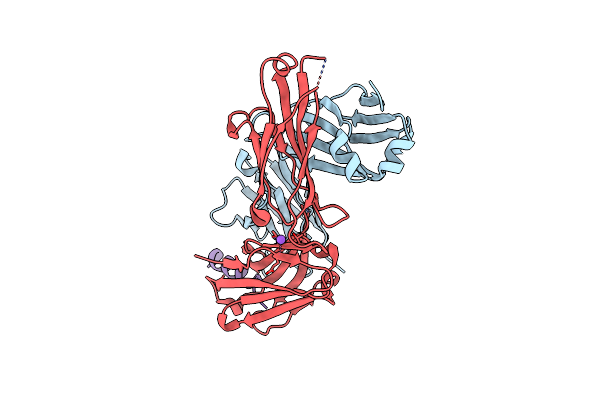
Crystal Structure Of The Non-Neutralizing Hiv Antibody 13H11 Fab Fragment With A Gp41 Mper-Derived Peptide In A Helical Conformation
Organism: Mus musculus, Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2010-11-17 Classification: IMMUNE SYSTEM Ligands: EDO |
 |
Crystal Structure Of The Non-Neutralizing Hiv Antibody 13H11 Fab Fragment With A Gp41 Mper-Derived Peptide Bearing Ala Substitutions In A Helical Conformation
Organism: Mus musculus, Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2010-11-17 Classification: IMMUNE SYSTEM |
 |
Organism: Mus musculus,homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2010-11-17 Classification: IMMUNE SYSTEM Ligands: ACT, SO4 |
 |
Crystal Structure Of The Neutralizing Hiv Antibody 2F5 Fab Fragment (Recombinantly Produced Fab) With 17 Aa Gp41 Mper-Derived Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2010-11-17 Classification: IMMUNE SYSTEM |

