Search Count: 35,396
All
Selected
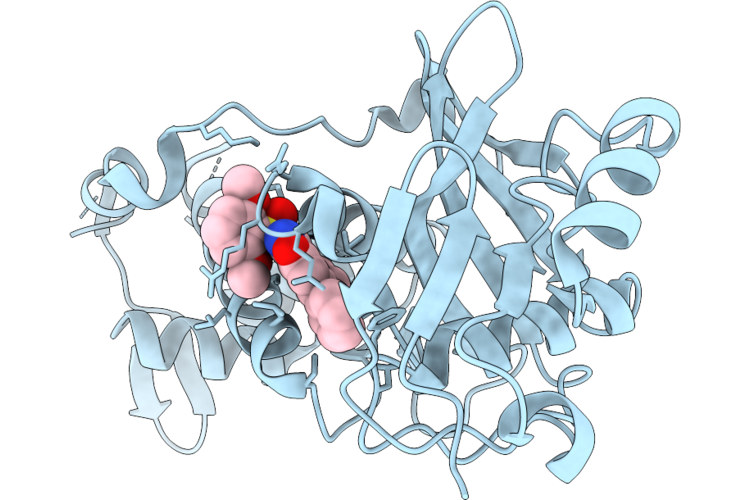 |
Crystal Structure Of Myst Histone Acetyltransferase Kat6A In Complex With Inhibitor Compound 9
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-05-06 Classification: TRANSCRIPTION Ligands: A1MGC |
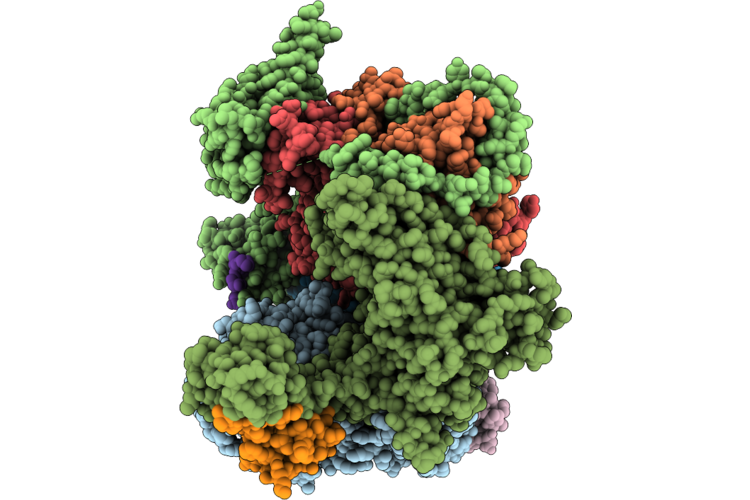 |
Cryo-Em Structure Of The Human Holo-Tfiih And Xpc Initial Encounter Complex
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
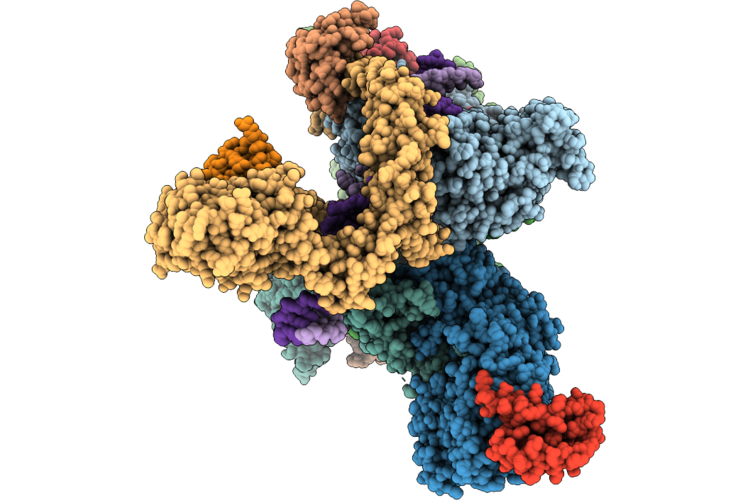 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
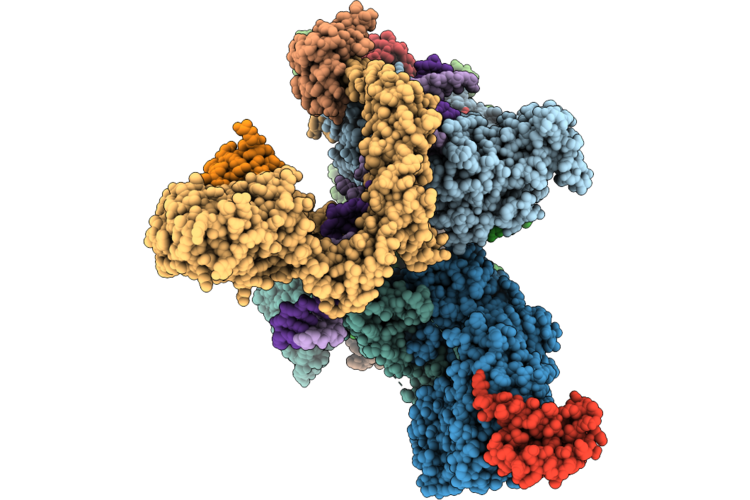 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
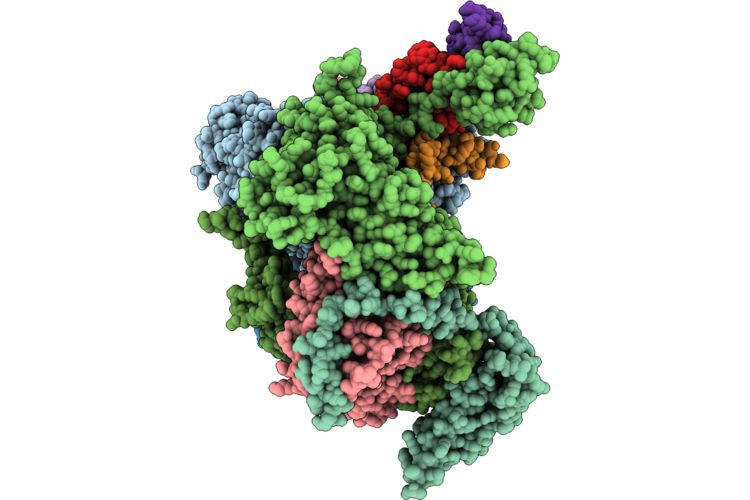 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc-Xpa Complex Bound To Bulky Lesion-Mimic Dna
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
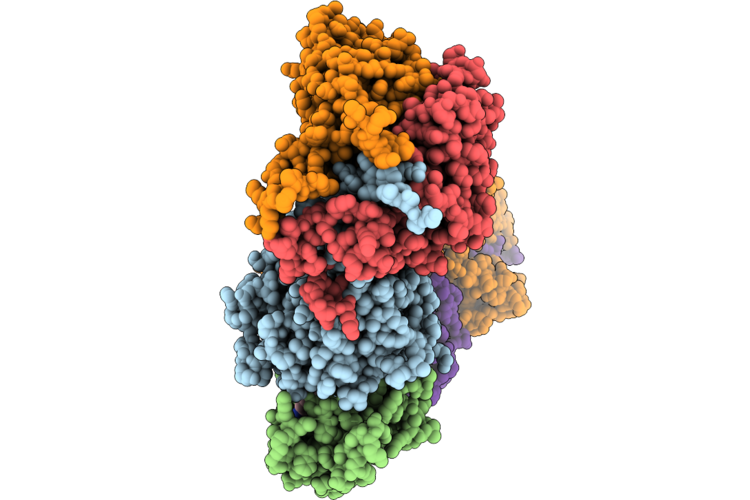 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10088
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
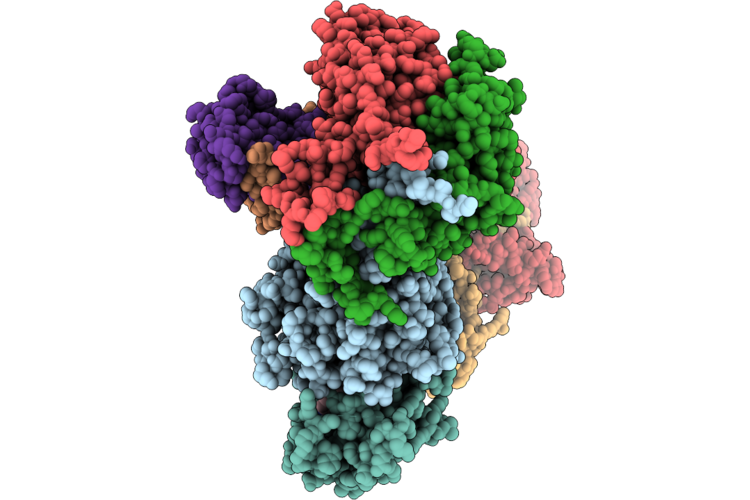 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2, Unc10088 And Bach1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
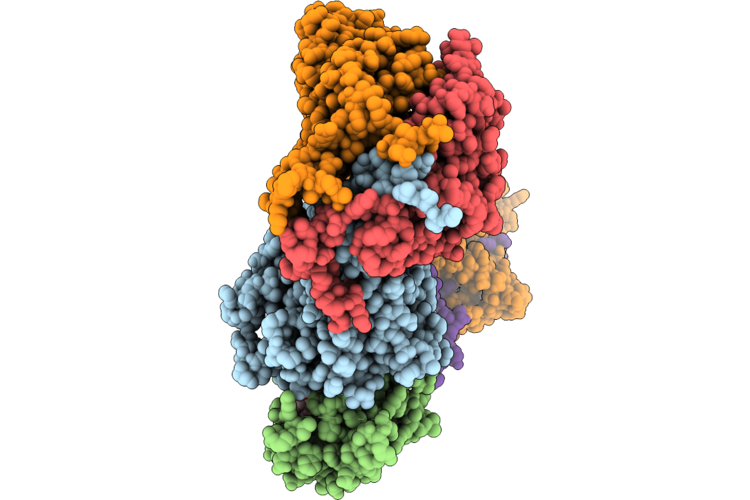 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10415667
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J21 |
 |
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Listeria monocytogenes egd-e
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
Organism: Listeria monocytogenes egd-e
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
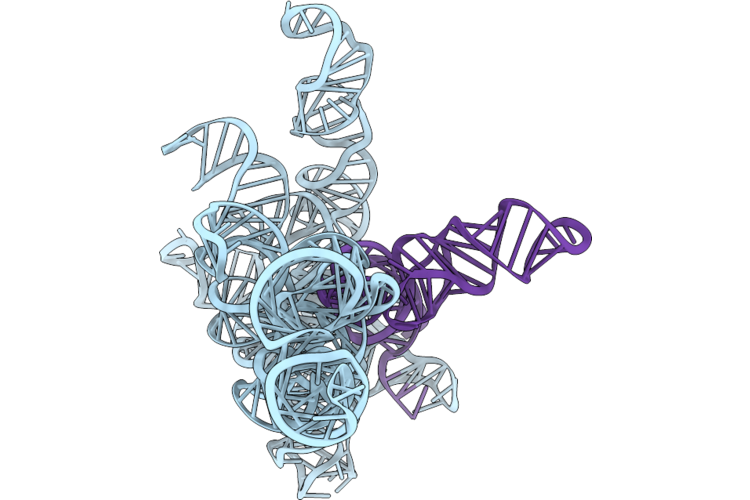 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna With Non-Complementary 5' Leader (Consensus)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna With Loop-Back 5' Leader
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RNA Ligands: CA |
 |
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TOXIN |
 |
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:3.01 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN |
 |
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: CD |
 |
Organism: Xenopus laevis, Homo sapiens, Artificial sequences
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA |

