Search Count: 555
All
Selected
 |
Epitope And Functional Classification Of Human Neutralizing Antibodies Against Sftsv Gn
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.85 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN |
 |
Epitope And Functional Classification Of Human Neutralizing Antibodies Against Sftsv Gn
Organism: Homo sapiens, Severe fever with thrombocytopenia syndrome virus
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN |
 |
Epitope And Functional Classification Of Human Neutralizing Antibodies Against Sftsv Gn
Organism: Homo sapiens, Severe fever with thrombocytopenia syndrome virus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN |
 |
Epitope And Functional Classification Of Human Neutralizing Antibodies Against Sftsv Gn
Organism: Homo sapiens, Severe fever with thrombocytopenia syndrome virus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN |
 |
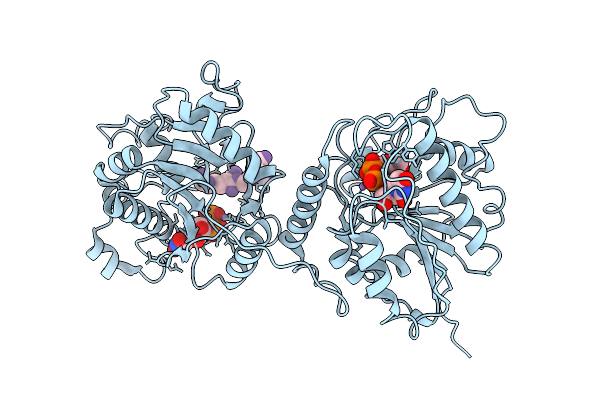
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.52 Å Release Date: 2026-04-22 Classification: RECOMBINATION Ligands: A1EVT |
 |
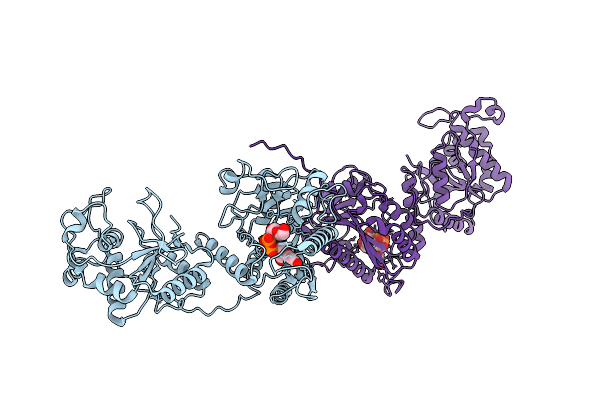
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.08 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN, UDP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
 |
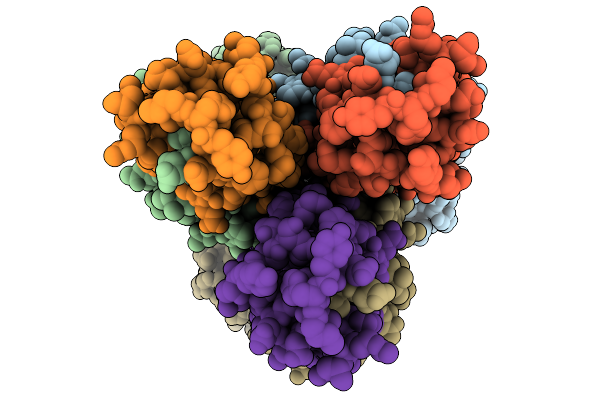
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GDU, MN |
 |
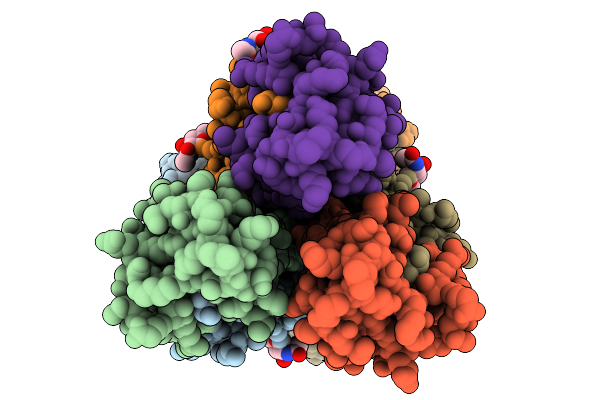
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: VIRAL PROTEIN |
 |
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN |
 |
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: NAG |
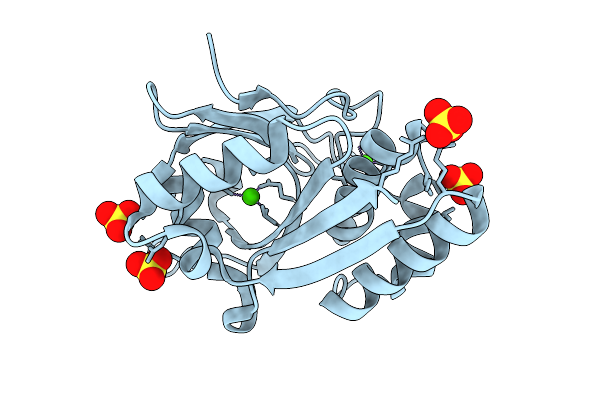 |
Organism: Metagenomes
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: LYASE Ligands: CA, SO4 |
 |
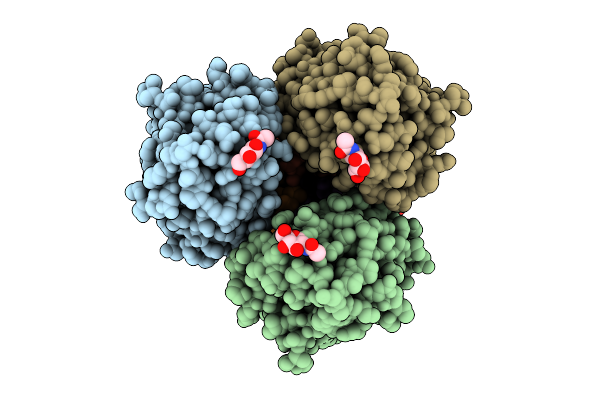
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
 |
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN Ligands: NAG |
 |
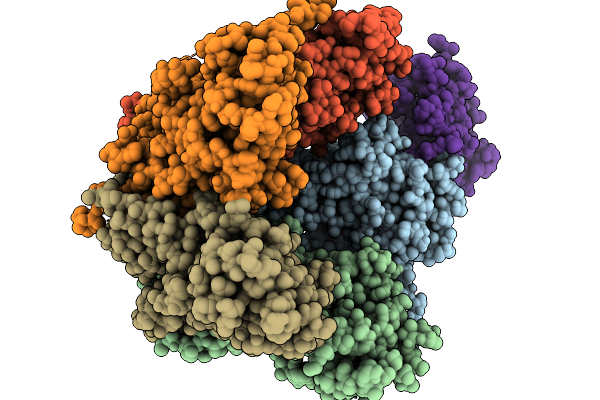
Organism: Severe acute respiratory syndrome coronavirus 2, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.33 Å Release Date: 2026-03-18 Classification: VIRAL PROTEIN |
 |
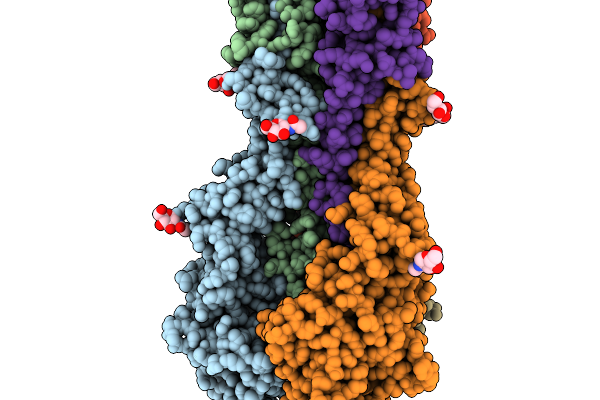
Organism: Human papillomavirus 45
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN |
 |
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Crystal Structure Of H1 Haemagglutinin Hn/18-Ha2-L113S From Influenza A Virus
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: NAG |

