Search Count: 277
All
Selected
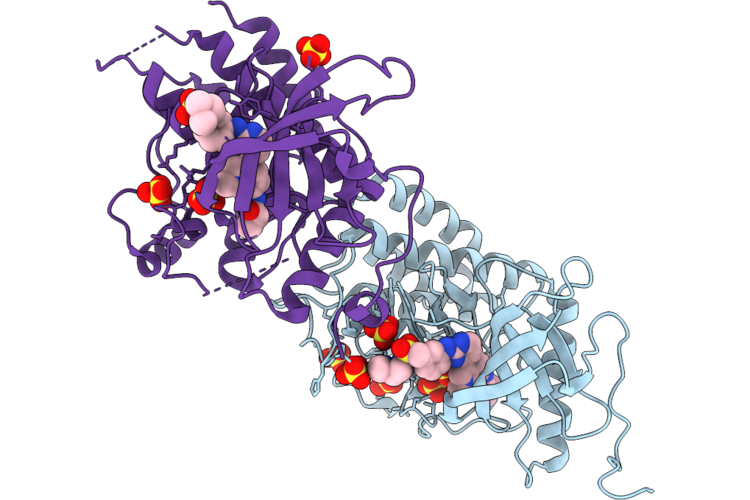 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1ESP, SO4 |
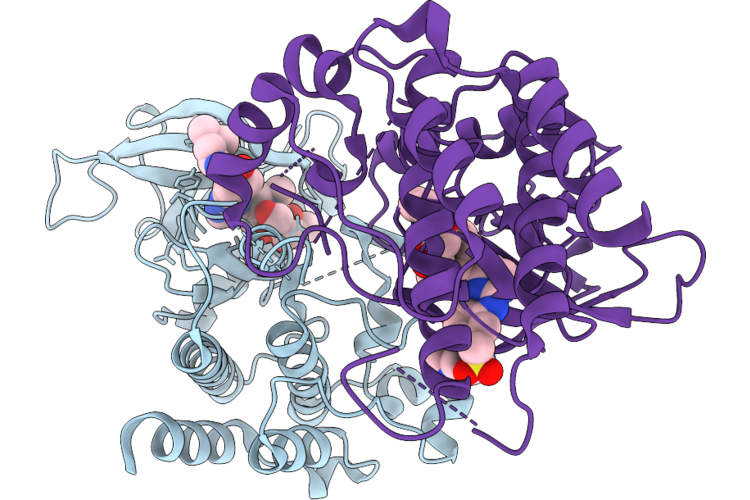 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1ESP, GOL |
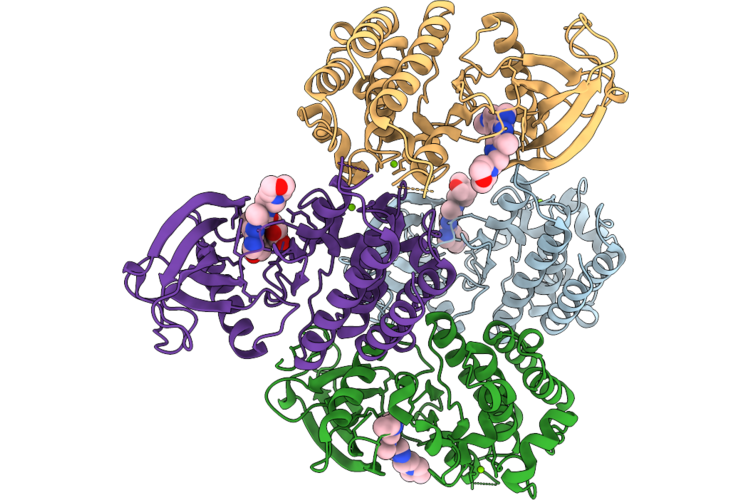 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1ESW, MG, GOL |
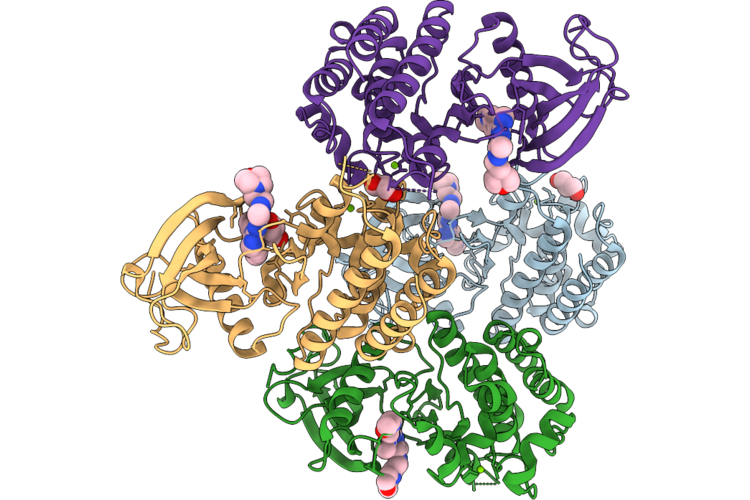 |
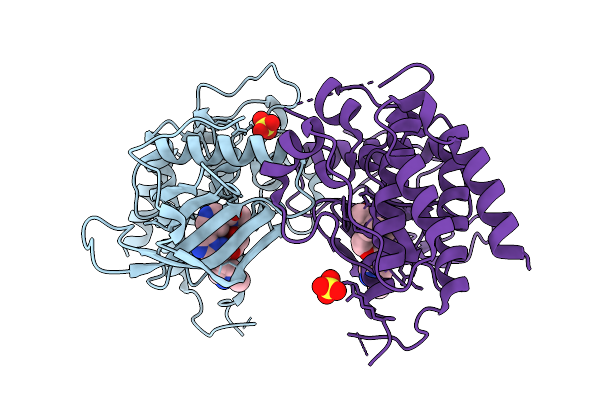
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1ESX, MG, GOL |
 |
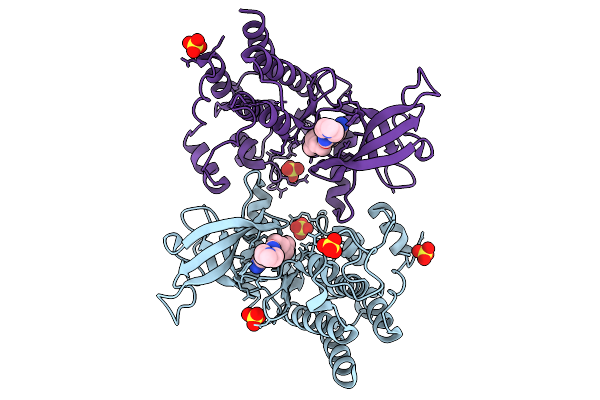
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-21 Classification: TRANSFERASE Ligands: A1EOH, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-01-21 Classification: TRANSFERASE Ligands: A1EOJ, SO4 |
 |
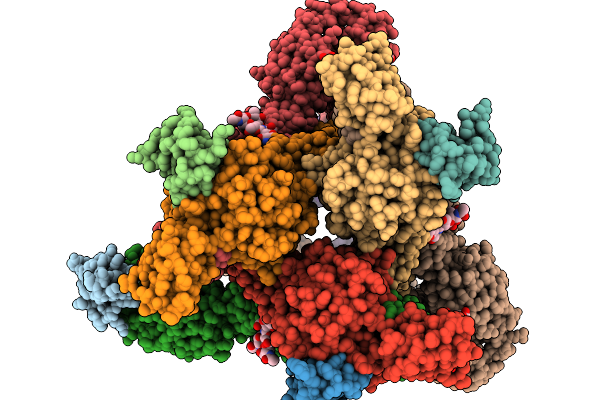
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-01-14 Classification: SIGNALING PROTEIN |
 |
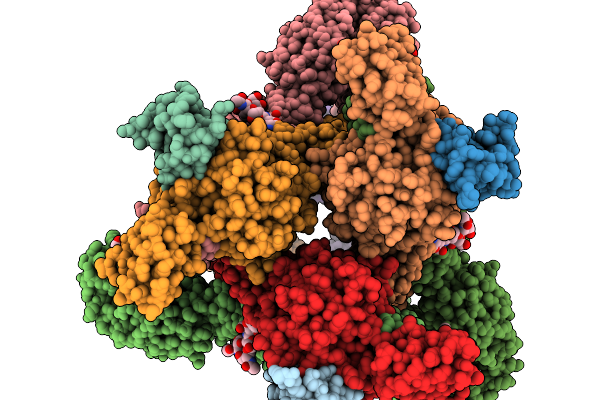
Organism: Homo sapiens, Mus musculus, Semliki forest virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS Ligands: CA |
 |
Organism: Semliki forest virus
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: VIRUS |
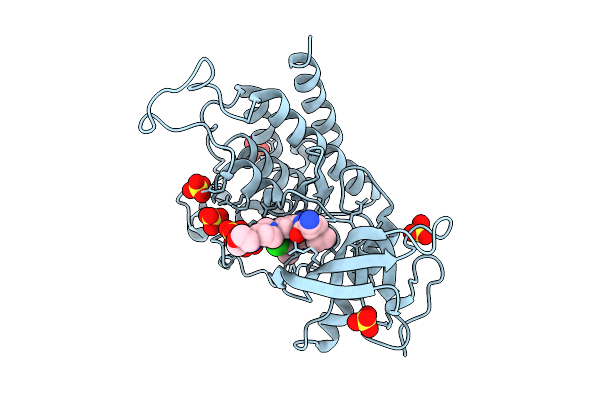 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2025-10-22 Classification: TRANSFERASE Ligands: A1EEW, SO4 |
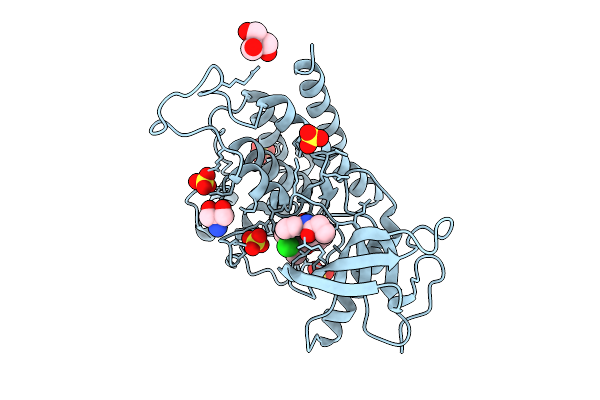 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2025-10-22 Classification: TRANSFERASE Ligands: A1EEX, SO4, TRS |
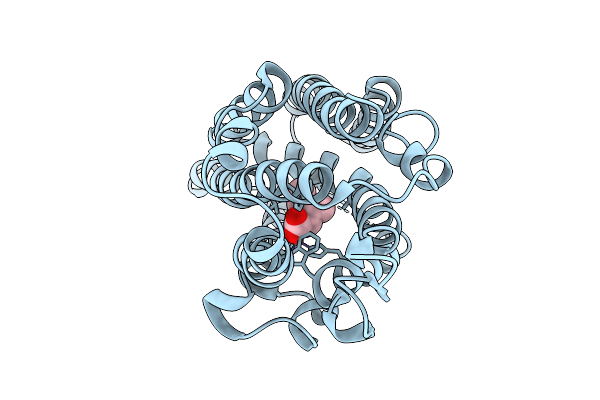 |
Organism: Streptomyces armeniacus
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, ILE |
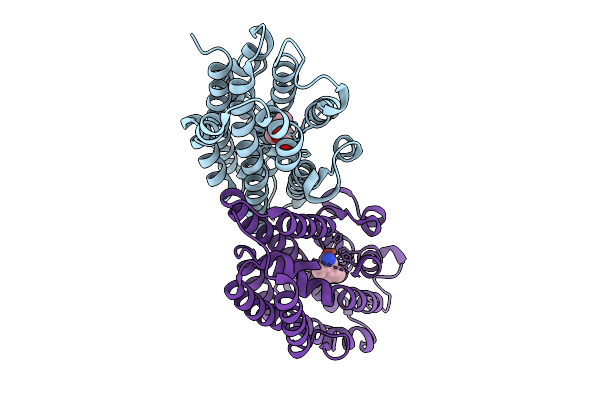 |
Organism: Streptomyces sp. z26
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, A1EAJ |
 |
Organism: Streptomyces sp. z26
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, A1EAI |
 |
Organism: Streptomyces sp. z26
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, A1EAJ |
 |
Organism: Streptomyces sp. z26
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, ILE |
 |
Organism: Streptomyces sp. z26
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, ILE |
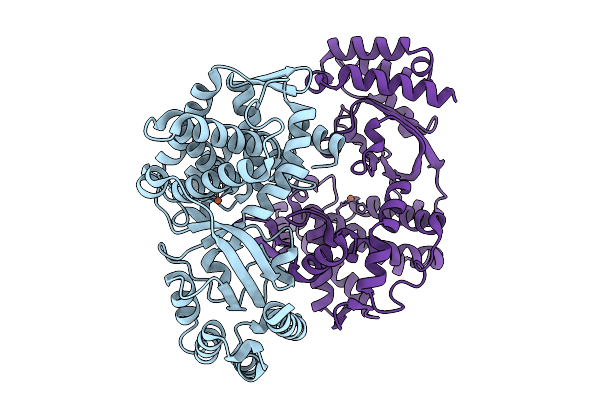 |
Organism: Streptomyces sp. nth33
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2 |
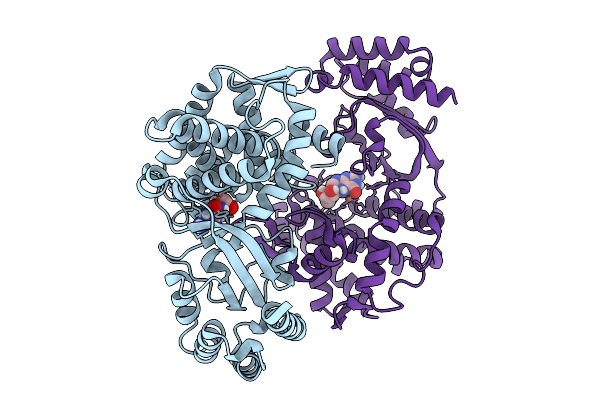 |
Organism: Streptomyces sp. nth33
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, H4B |
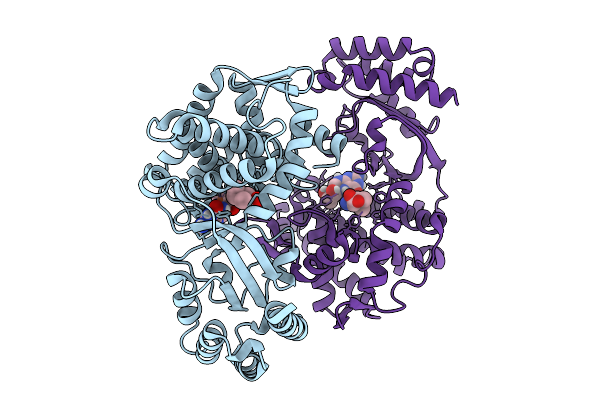 |
Organism: Streptomyces sp. nth33
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-10-08 Classification: OXIDOREDUCTASE Ligands: FE2, ILE, H4B |

