Search Count: 167
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-27 Classification: RNA |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2026-05-27 Classification: RNA |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:4.80 Å Release Date: 2026-05-27 Classification: RNA |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:4.70 Å Release Date: 2026-05-27 Classification: RNA |
 |
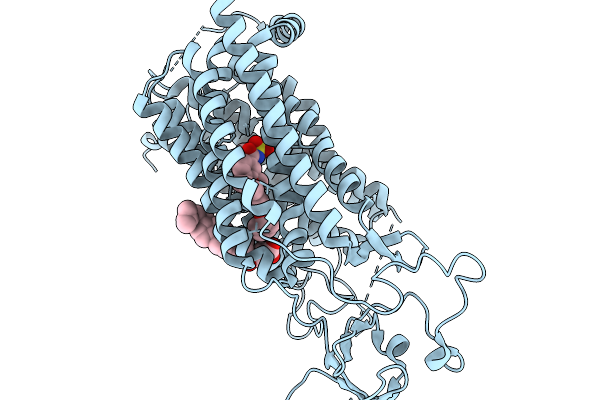
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: Y01 |
 |
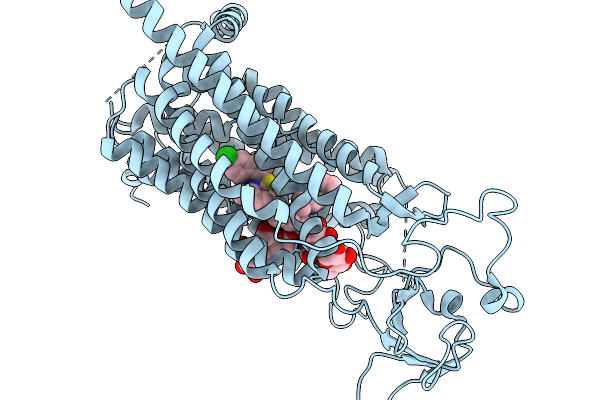
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: ZLK, Y01 |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: A1JBL, AV0 |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: LSN |
 |
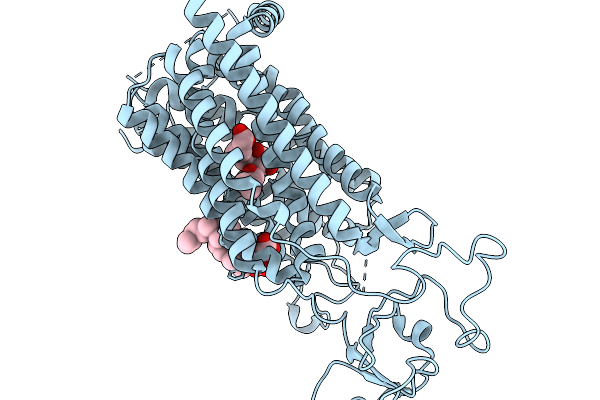
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: P2E, Y01 |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: UGU, Y01 |
 |
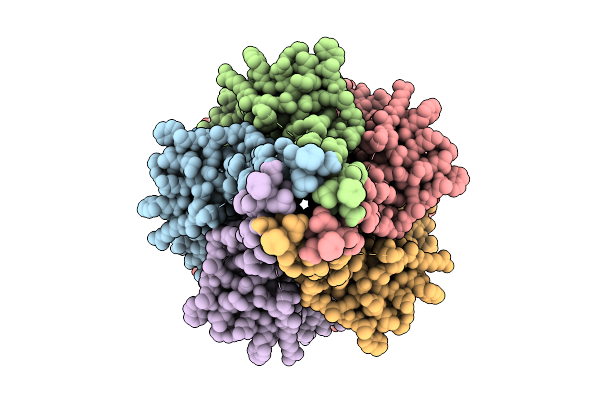
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: TCW |
 |
Alpha-7 Nicotinic Acetylcholine Receptor Bound To Inhibitory Bicyclic Peptide Kp2007 In A Resting State.
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: NAG |
 |
Organism: Ostrinia furnacalis
Method: SOLID-STATE NMR Release Date: 2025-10-22 Classification: STRUCTURAL PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: CYTOSOLIC PROTEIN |
 |
Human Phosphorylase Kinase Alpha/Gamma/Delta Subcomplex - Phosphorylation And Ca2+ Bound State
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: CYTOSOLIC PROTEIN Ligands: ATP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-04-23 Classification: CYTOSOLIC PROTEIN Ligands: ATP |
 |
Organism: Mycobacterium tuberculosis variant bovis bcg str. pasteur 1173p2
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: RIBOSOME/ANTIBIOTIC Ligands: ZN, MG, A1L32 |
 |
Organism: Macaca fascicularis, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2024-11-13 Classification: TRANSPORT PROTEIN/IMMUNE SYSTEM Ligands: TCH |

