Search Count: 302
All
Selected
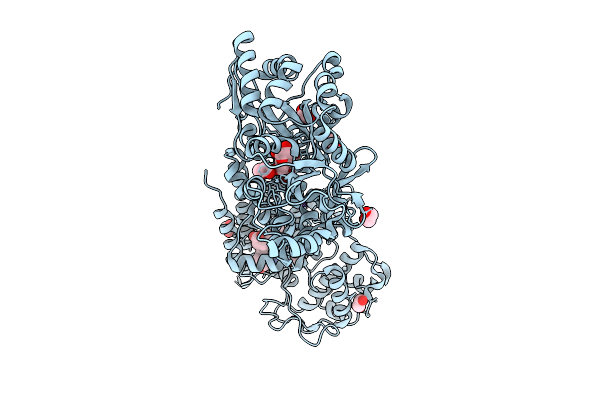 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) H143Y Mutant Complexed With 5-Methoxy-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: A1A2Y, EDO, HEM, CYN, NA |
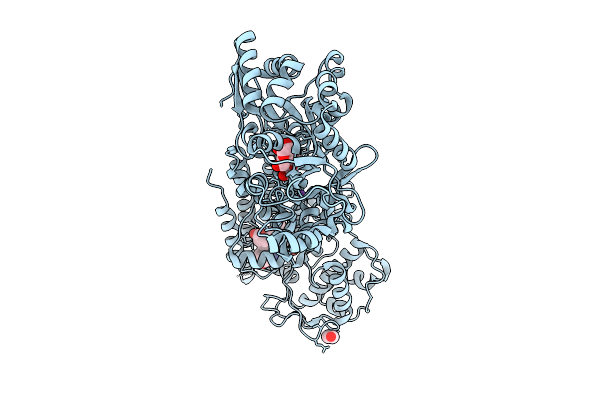 |
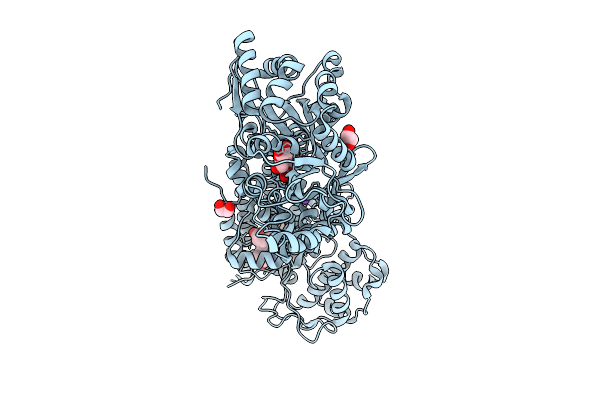
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With 5-Hydroxy-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, HEM, CYN, 4PQ, NA |
 |
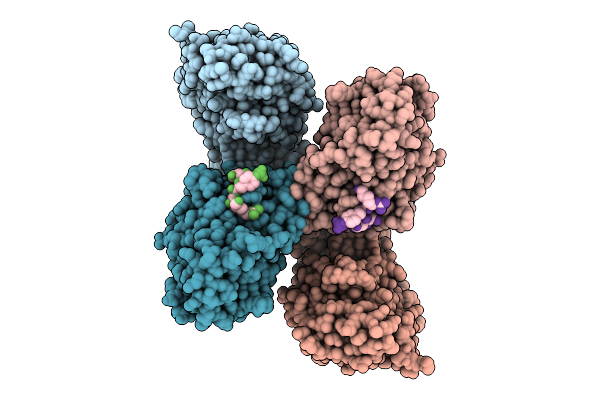
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) H143Y Mutant Complexed With 5-Methyl-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, HEM, CYN, D0Q, NA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE |
 |
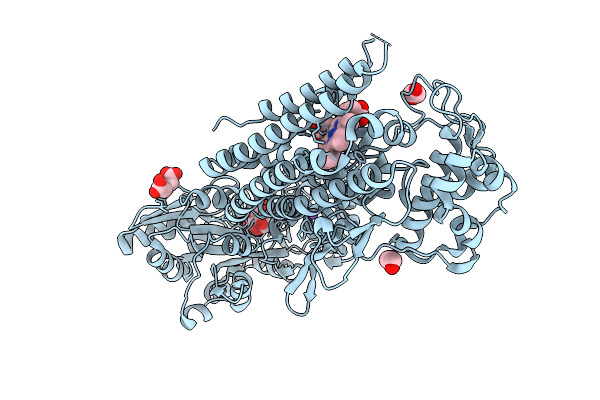
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: PEG, EDO, HEM, TRP, CYN, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With 5-Ht
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: SRO, EDO, PEG, HEM, CYN, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) As Resting State
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, CIT, HEM, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) H143Y Mutant Complexed With L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: TRP, PEG, CIT, EDO, HEM, CYN, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With Cyanide Ion
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: CIT, EDO, HEM, CYN, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With D-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: HEM, CYN, DTR, EDO, NA |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With 5-Methoxy-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: A1A2Y, EDO, HEM, CYN |
 |
Crystal Structure Of The Indoleamine 2,3-Dioxygenagse 2 (Ido2) Complexed With 5-Methyl-L-Trp
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: EDO, HEM, CYN, D0Q, NA |
 |
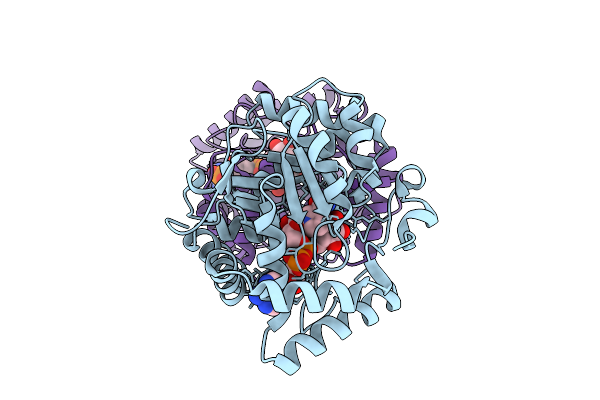
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Galactose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: GIV, NAP |
 |
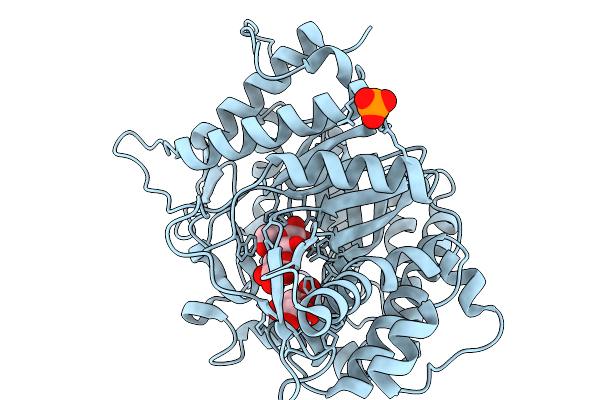
Crystal Structure Of L-Galactose Dehydrogenase From Luteolibacter Sp. Strain Lg18 In Complex With L-Glucose And Nadp+
Organism: Luteolibacter sp. lg18
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: Z8T, NAP |
 |
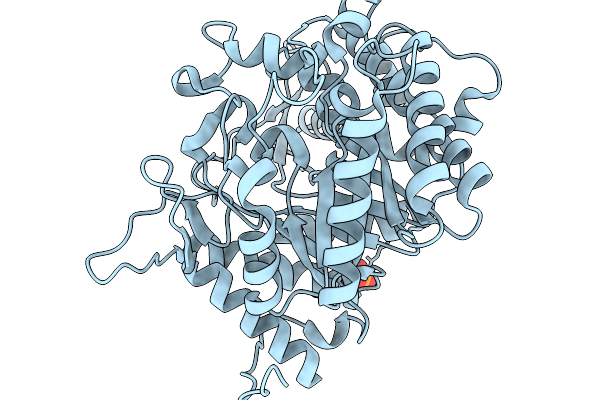
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: GLC, BGC, PO4 |
 |
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
Crystal Structure Of Glucose Tolerant Beta-Glucosidase Unbgl1 Mutant (C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: CL, PO4 |
 |
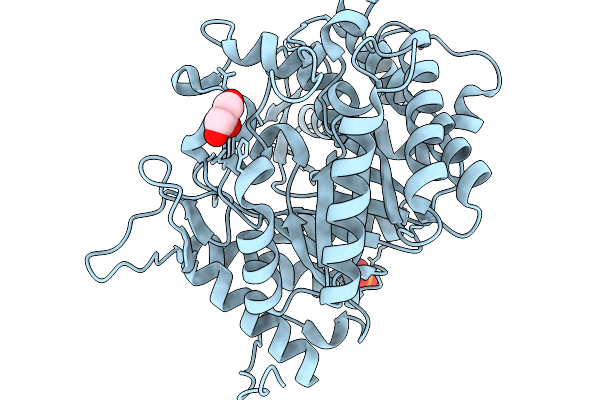
Crystal Structure Of Glucsoe Bound Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_C188V)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, PO4 |
 |
Crystal Structure Of Glucose Tolerant Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: PO4, GOL, CL |
 |
Crystal Structure Of Glucose Bound Gh1 Beta-Glucosidase Mutant (Unbgl1_H261W)
Organism: Soil metagenome
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-02-04 Classification: HYDROLASE Ligands: BGC, GOL, PO4 |

