Search Count: 184
 |
Organism: Micromonospora ureilytica
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-06-03 Classification: HYDROLASE Ligands: GOL, EDO |
 |
Organism: Allorhizocola rhizosphaerae
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-06-03 Classification: HYDROLASE Ligands: GOL, EDO |
 |
Organism: Saccharothrix sp. cb00851
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-06-03 Classification: HYDROLASE Ligands: GOL, EDO |
 |
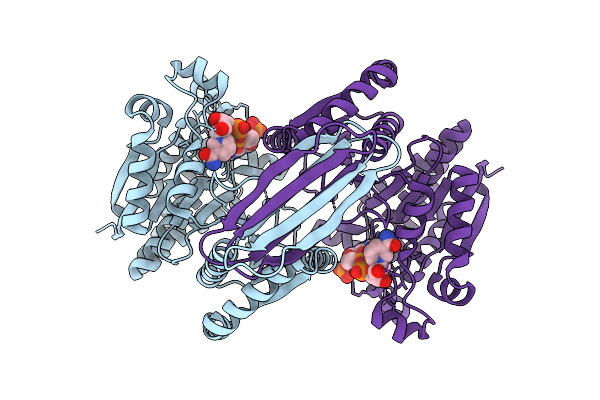
Organism: Micromonospora ureilytica
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-06-03 Classification: HYDROLASE Ligands: SO4, GOL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CYTOSOLIC PROTEIN Ligands: NAP |
 |
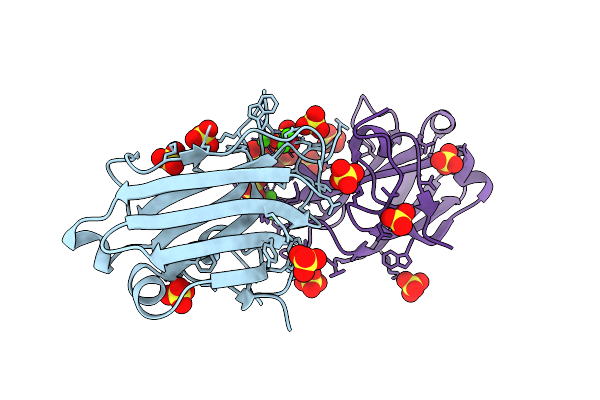
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CYTOSOLIC PROTEIN Ligands: NAP |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2026-01-21 Classification: TRANSFERASE Ligands: CA, SO4 |
 |
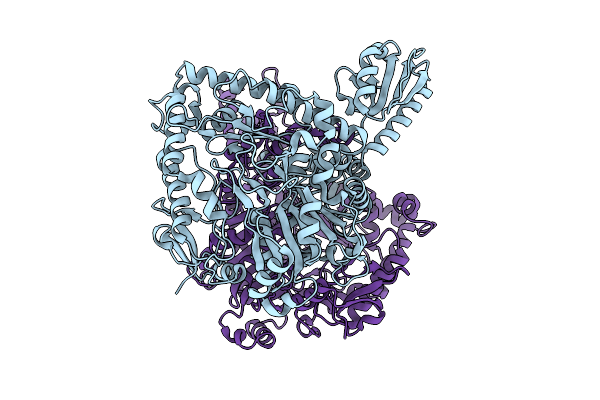
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2025-12-24 Classification: LYASE |
 |
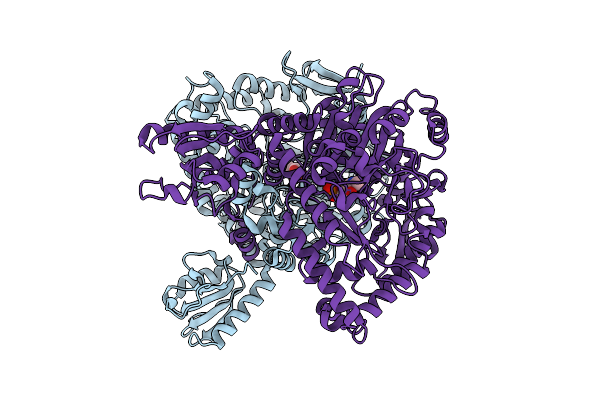
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2025-12-24 Classification: LYASE |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2025-12-24 Classification: LYASE Ligands: PLP |
 |
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-12-24 Classification: LYASE |
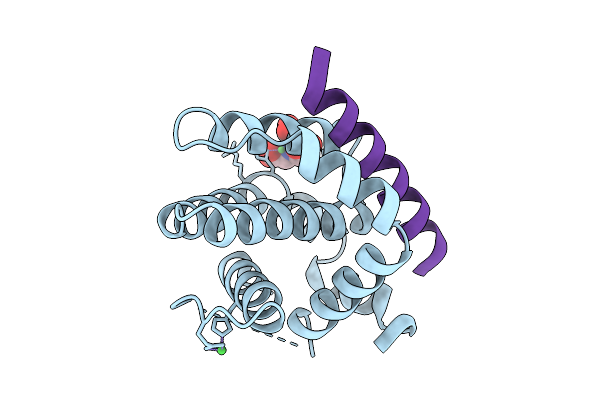 |
Crystal Structure Of Bcl-2 In Complex With A Stapled Bad Bh3 Peptide Bad Sahb 4.2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-10-08 Classification: APOPTOSIS Ligands: NI, NTA |
 |
Crystal Structure Of Bcl-2 (G101V) Mutant In Complex With A Stapled Bad Bh3 Peptide Bad Sahb 4.2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2025-10-08 Classification: APOPTOSIS |
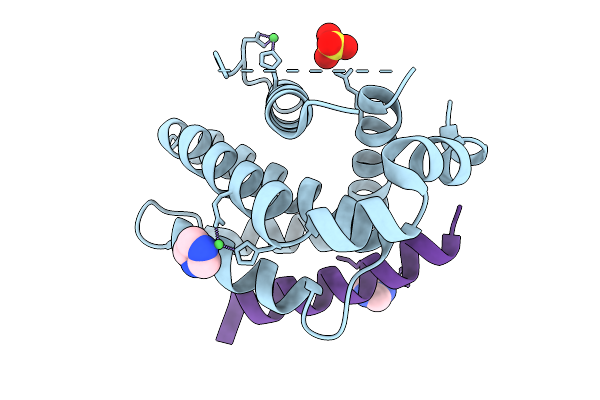 |
Crystal Structure Of Human Bcl-2 (R129L) Mutant In Complex With A Stapled Bad Bh3 Peptide Bad Sahb 4.2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-10-08 Classification: APOPTOSIS Ligands: IMD, SO4, NI |
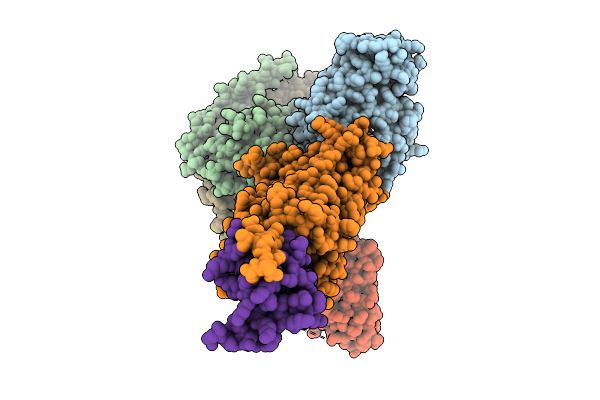 |
Organism: Homo sapiens, Human gammaherpesvirus 8, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.98 Å Release Date: 2025-09-03 Classification: VIRAL PROTEIN/SIGNALING PROTEIN |
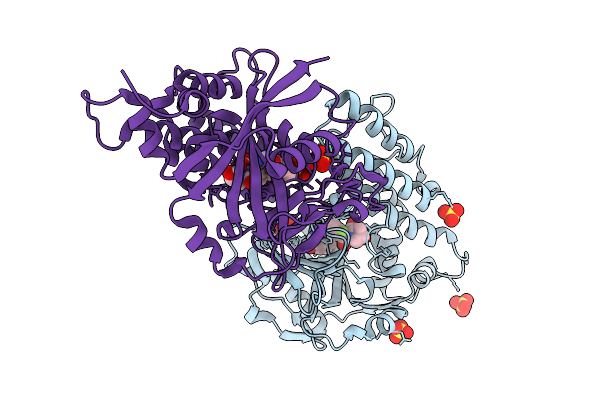 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-08-27 Classification: TRANSFERASE/INHIBITOR Ligands: A1AEO, SO4 |
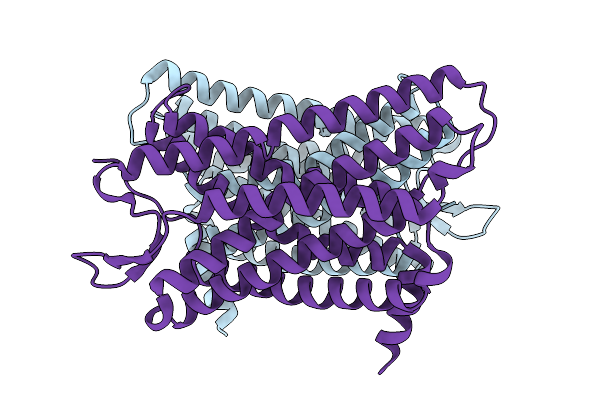 |
Organism: Human gammaherpesvirus 8
Method: ELECTRON MICROSCOPY Release Date: 2025-08-27 Classification: VIRAL PROTEIN |
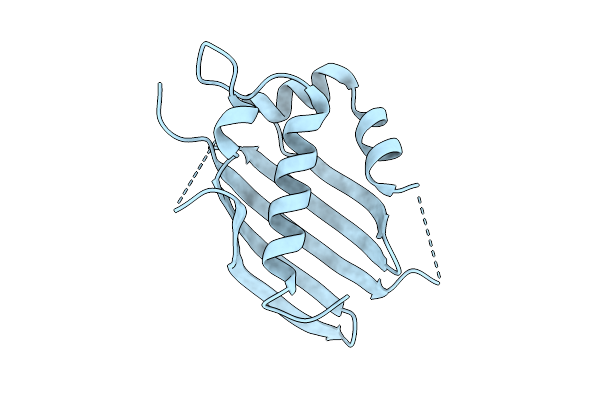 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2025-07-23 Classification: PROTEIN BINDING Ligands: HOH |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: NUCLEAR PROTEIN Ligands: ATP, MG, ADP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-11 Classification: NUCLEAR PROTEIN Ligands: ZN, ATP, MG, ADP |

