Search Count: 74
All
Selected
 |
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN |
 |
Prenylated-Fmn Maturase Phdc From Mycolicibacterium Fortuitum Bound To Flavin Mononucleotide
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN Ligands: EDO, NA, FMN, SO4 |
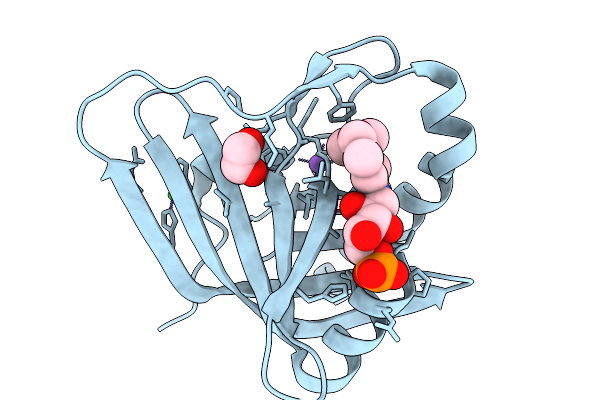 |
Prenylated-Fmn Maturase Phdc From Mycolicibacterium Fortuitum Bound To Prenylated Flavin Mononucleotide
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN Ligands: 4LU, NI, NA, EDO |
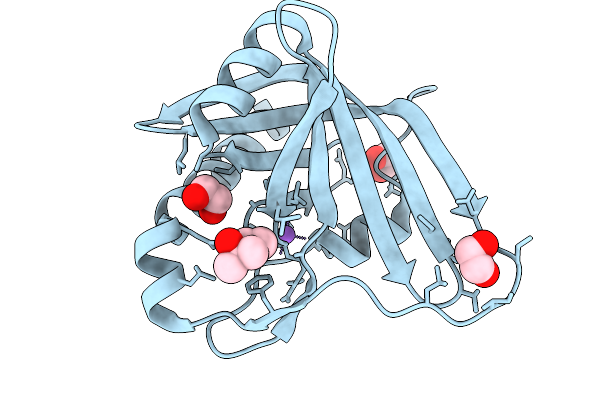 |
Prenylated-Fmn Maturase Phdc E45A Mutant From Mycolicibacterium Fortuitum (Apo)
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN |
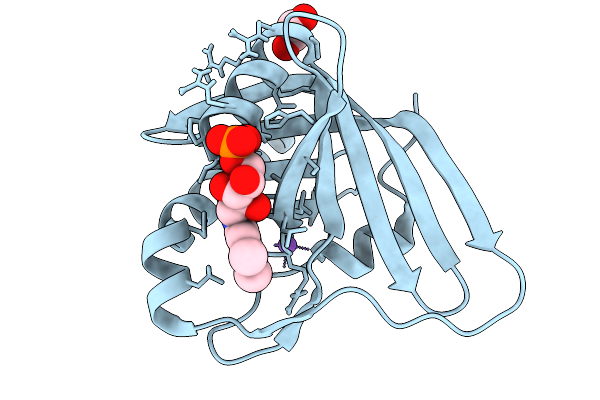 |
Prenylated-Fmn Maturase Phdc E45A Mutant From Mycolicibacterium Fortuitum Bound To Flavin Mononucleotide
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN Ligands: EDO, FMN, NA |
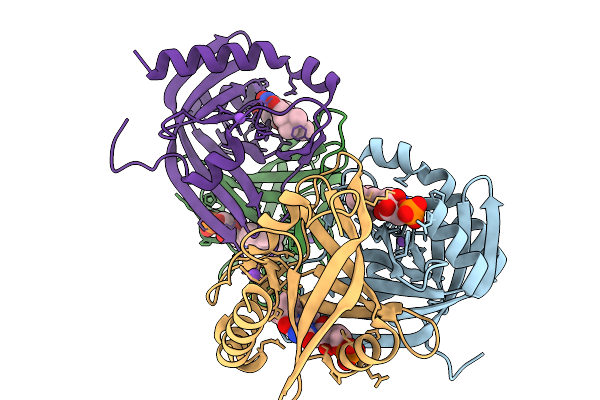 |
Prenylated-Fmn Maturase Phdc From Mycolicibacterium Fortuitum Bound To Prenylated Flavin Mononucleotide In The P1 Space Group
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN Ligands: 4LU, NA |
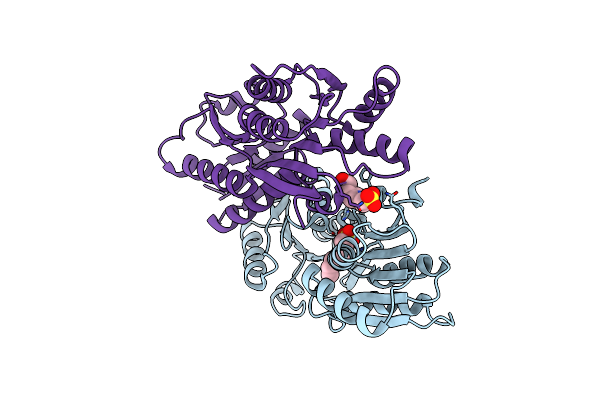 |
Crystal Structure Of Catabolite Repressor Acivator From E. Coli In Complex With Hepes
Organism: Escherichia coli 536
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2024-04-24 Classification: GENE REGULATION Ligands: EPE |
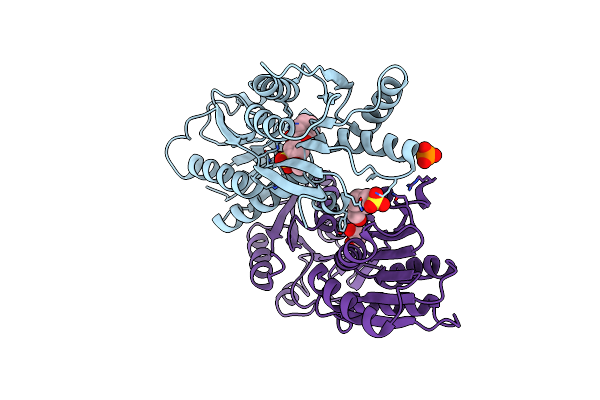 |
Crystal Structure Of Catabolite Repressor Acivator From E. Coli In Complex With Sulisobenzone
Organism: Escherichia coli 536
Method: X-RAY DIFFRACTION Resolution:3.05 Å Release Date: 2024-04-24 Classification: TRANSCRIPTION Ligands: I4Y, PO4, EPE, EDO |
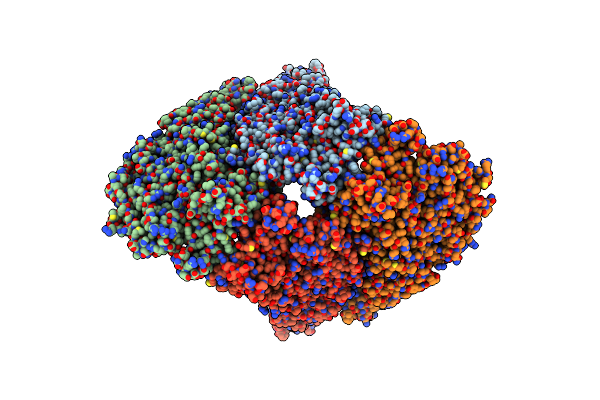 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
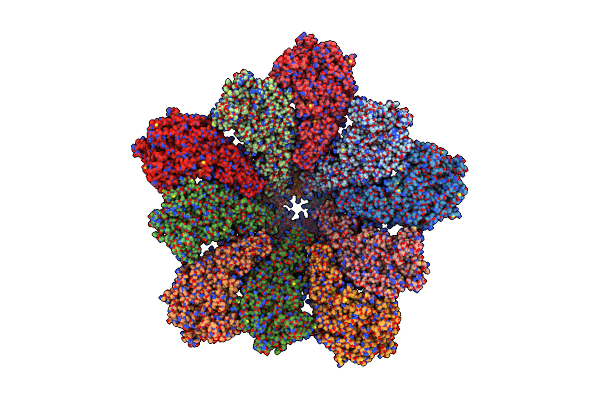 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Bluetongue virus (serotype 1 / isolate south africa)
Method: ELECTRON MICROSCOPY Release Date: 2024-04-24 Classification: VIRAL PROTEIN |
 |
Organism: Gallus gallus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2023-07-26 Classification: PROTEIN BINDING Ligands: K, PO4 |
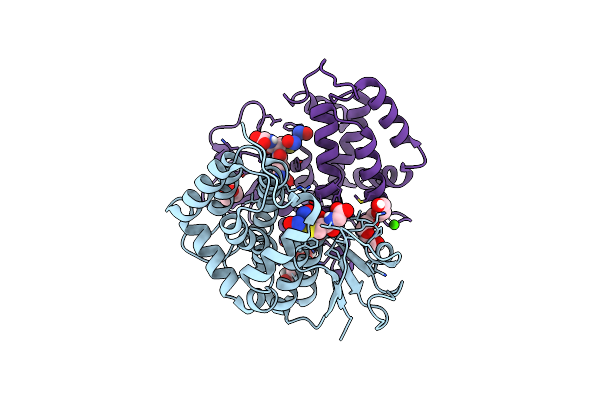 |
Crystal Structure Of Trametes Versicolor Glutathione Transferase Omega 3S In Complex With Dinitrosyl Glutathionyl Iron Complex (Dngic)
Organism: Trametes versicolor
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2023-03-01 Classification: TRANSFERASE Ligands: PG4, GOL, FE, NO, GSH, CA, PEG |
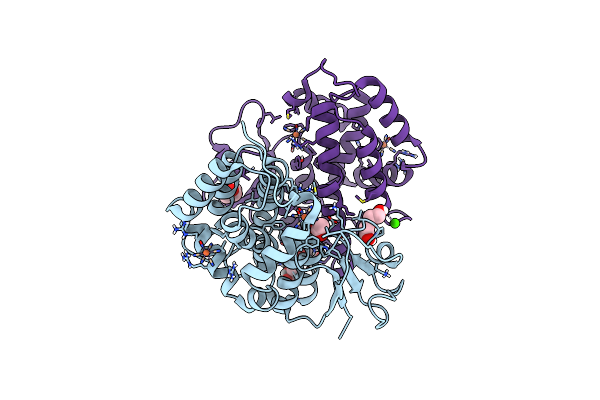 |
Crystal Structure Of Trametes Versicolor Glutathione Transferase Omega 3S In Complex With Sodium Nitroprusside
Organism: Trametes versicolor
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-03-01 Classification: TRANSFERASE Ligands: NT3, ACT, GOL, PEG, CA |
 |
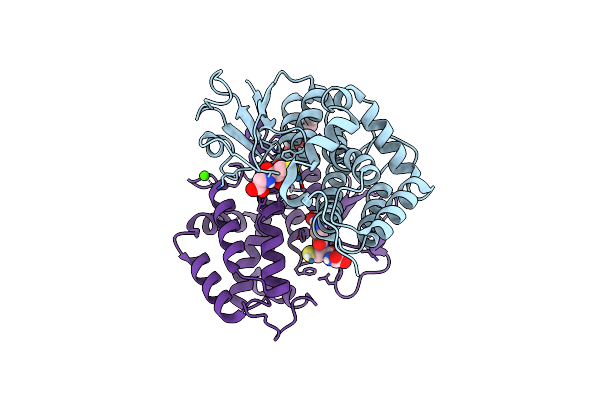
Crystal Structure Of Trametes Versicolor Glutathione Transferase Omega 3S In Complex With Hydroxy-Tetranitro-Nitrosyl-Ruthenate
Organism: Trametes versicolor
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-03-01 Classification: TRANSFERASE Ligands: ODU, ACT, GOL, RU, CA |
 |
Crystal Structure Of Trametes Versicolor Glutathione Transferase Omega 3S In Complex With Glutathione And Pentachloro-Nitrosyl-Osmate
Organism: Trametes versicolor
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2023-03-01 Classification: TRANSFERASE Ligands: OS, GOL, GSH, CA |

