Search Count: 227
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
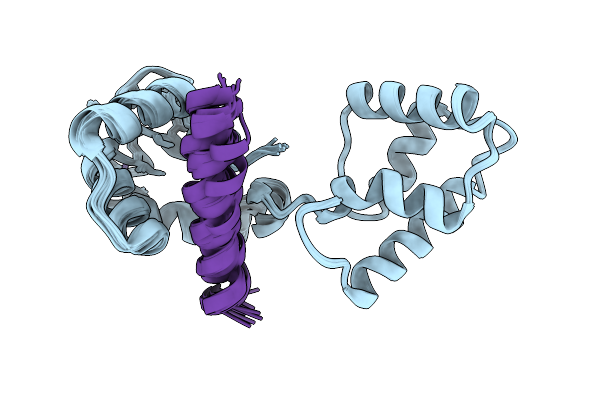 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-04-08 Classification: METAL BINDING PROTEIN Ligands: CA |
 |
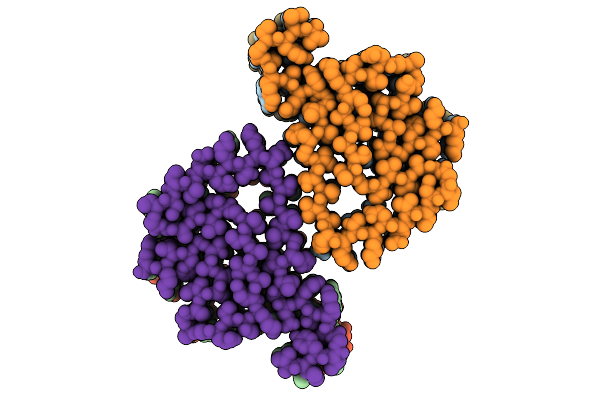
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-08-27 Classification: IMMUNE SYSTEM Ligands: A1BLD |
 |
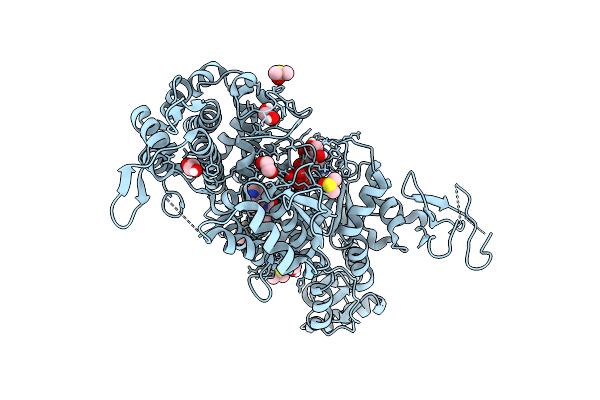
Structure Of In-Vivo Formed Alpha-Synuclein Fibrils Purified From A M83+/- Mouse Brain Injected With Recombinant 1B Fibrils
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2025-07-30 Classification: PROTEIN FIBRIL |
 |
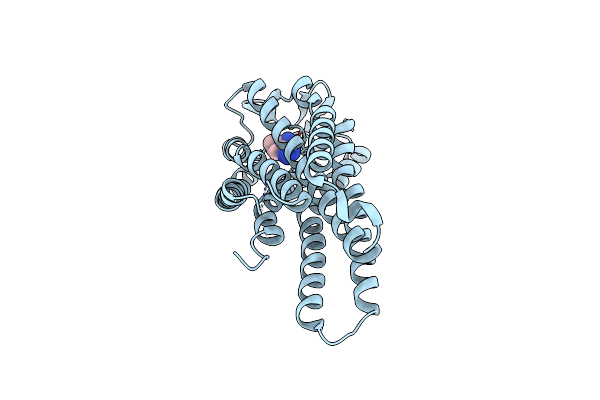
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2025-07-09 Classification: MOTOR PROTEIN Ligands: ADP, VO4, MG, DMS, EDO, A1IGE, CL |
 |
Cryoem Structure Of Hca2 Dreadd Gi1 In Complex With Fch-2296413 (Local Refinement)
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: XI9 |
 |
Solution Nmr Structure Of The Delta60 Domain Of The Hepatitis Delta Virus Small Antigen Shdag
Organism: Escherichia coli bl21
Method: SOLUTION NMR Release Date: 2025-04-16 Classification: VIRAL PROTEIN |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: SIGNALING PROTEIN Ligands: A1AIW |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: SIGNALING PROTEIN Ligands: KCA |
 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: MEMBRANE PROTEIN Ligands: XOT |
 |
Organism: Homo sapiens, Lama glama, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: MEMBRANE PROTEIN Ligands: XI9 |
 |
Structure Of Recombinant Alpha-Synuclein Fibrils 1B Capable Of Seeding Gcis In Vivo
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-06-05 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2024-03-06 Classification: LYASE Ligands: ZN, MBO, BEZ |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2024-03-06 Classification: LYASE Ligands: MBO, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2024-03-06 Classification: LYASE Ligands: ZN, MBO |
 |
Structure Of N-Terminal Of Schistosoma Japonicum Asparaginyl-Trna Synthetase
Organism: Schistosoma japonicum
Method: SOLUTION NMR Release Date: 2023-09-06 Classification: LIGASE |
 |
Crystal Structure Of Carbonmonoxy Hemoglobin S (Liganded Sickle Cell Hemoglobin) Complexed With Gbt021601
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2023-04-05 Classification: OXYGEN TRANSPORT Ligands: HEM, FOR, OHF |
 |
Neurotensin Receptor Allosterism Revealed In Complex With A Biased Allosteric Modulator
Organism: Escherichia coli, Homo sapiens, Lama glama, Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2023-03-29 Classification: MEMBRANE PROTEIN |
 |
Organism: Rattus norvegicus, Homo sapiens, Escherichia coli, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2023-03-29 Classification: MEMBRANE PROTEIN Ligands: SRW |
 |
Organism: Rattus norvegicus, Spodoptera frugiperda, Homo sapiens, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-03-29 Classification: MEMBRANE PROTEIN |

