Search Count: 8,003
All
Selected
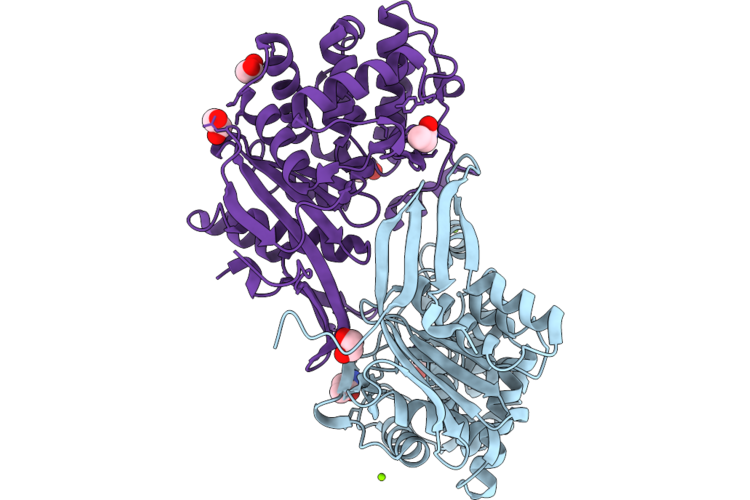 |
Reaiv Structure Containing An Acrylamide Adduct At Cys183 And An Orthoborate Ester At Ser47
Organism: Rhizobium etli
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-05-27 Classification: HYDROLASE Ligands: ZN, CL, EDO, MG, ROP |
 |
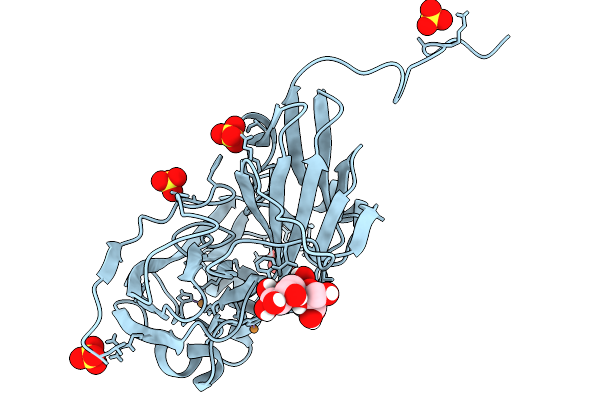
Atomic Resolution (1.02 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, NO2, GLC, FRU, SO4 |
 |
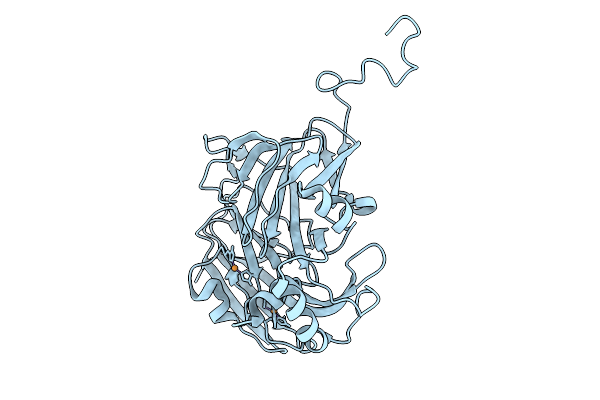
Atomic Resolution (1.00 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At High Ph (7.3) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4 |
 |
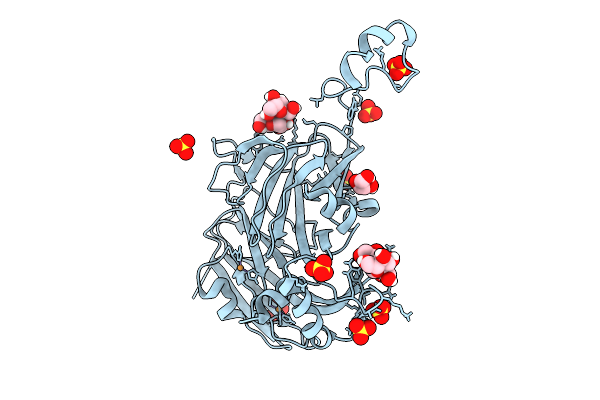
Atomic Resolution (1.05 A) Xfel Structure Of Chemically-Reduced Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Atomic Resolution (1.15 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At Low Ph (5.5) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4, GOL |
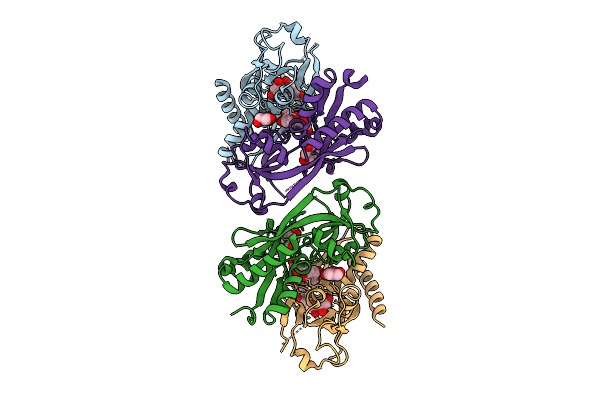 |
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) In Complex With Hesperetin From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION Ligands: GOL, 6JP |
 |
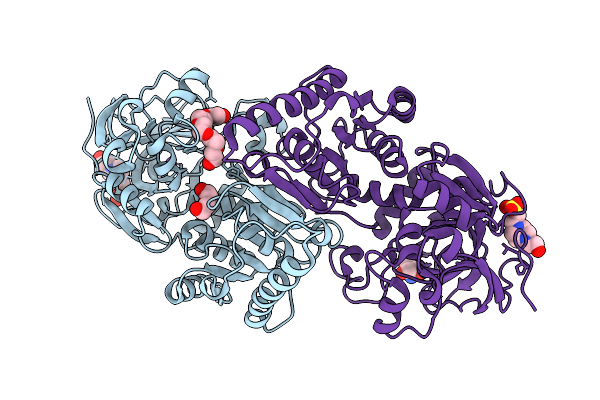
Crystal Structure Of Nodd-Ebd (Effector Binding Domain) From Rhizobium Leguminosarum Bv. Vicae 3841
Organism: Rhizobium leguminosarum
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2025-12-24 Classification: TRANSCRIPTION |
 |
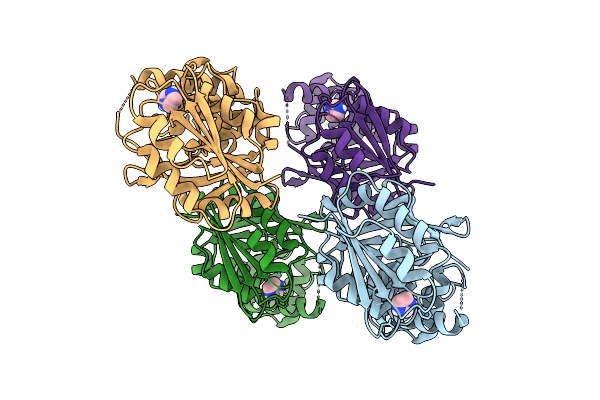
Organism: Mesorhizobium metallidurans
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: P6G, PEG, EPE, NA, TRS |
 |
Organism: Sinorhizobium meliloti
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: IMMUNE SYSTEM Ligands: CA |
 |
Organism: Sinorhizobium meliloti
Method: ELECTRON MICROSCOPY Resolution:4.40 Å Release Date: 2025-12-10 Classification: IMMUNE SYSTEM |
 |
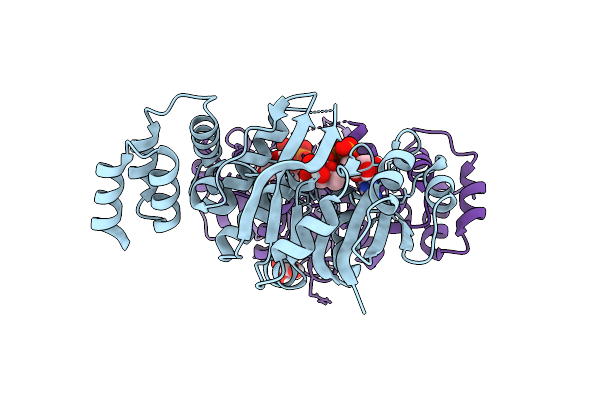
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: IMD, CL |
 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: GDP, GOL, SO4 |
 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: GFB, GOL |
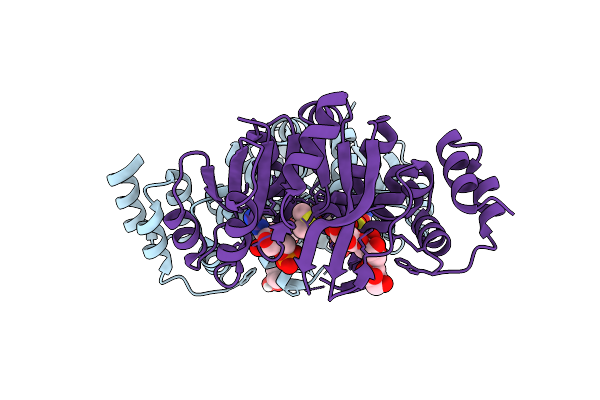 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: 5GP, DMS, FUC, A1CCV, BGC |
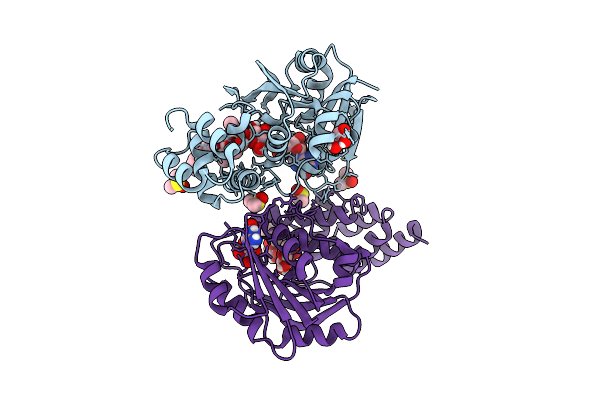 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: 5GP, GOL, DMS, BGC, A1CCW, FUC, OC9 |
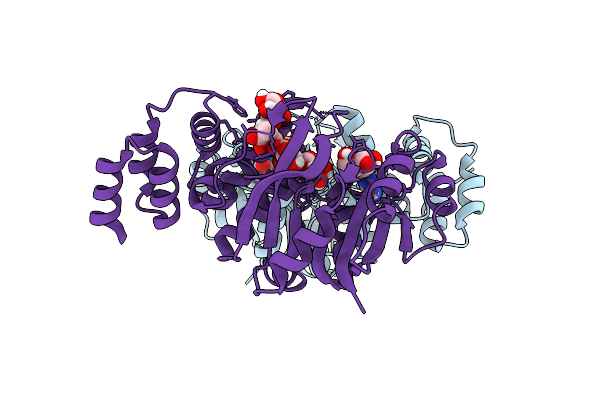 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: 5GP, DMS, GMP, FUC, A1CCW |
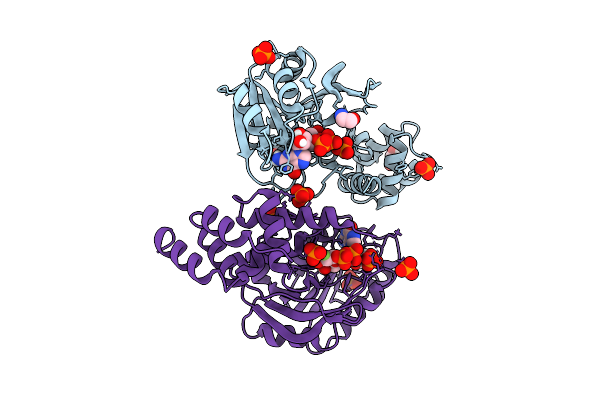 |
Organism: Rhizobium tropici ciat 899
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-10-29 Classification: TRANSFERASE Ligands: A1CCU, GLY, GOL, PO4 |
 |
Structure Of Phosphonate Monoester Hydrolase From Rhizobium Leguminosarum With Vanadate
Organism: Rhizobium johnstonii 3841
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: MN, VO4, NA |
 |
Structure Of Phosphonate Monoester Hydrolase From Rhizobium Leguminosarum With Phenyl Phosphonate
Organism: Rhizobium johnstonii 3841
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2025-10-08 Classification: HYDROLASE Ligands: MN, SV7, ACT, NA |
 |
The Outward-Open Structure Of Bjsemisweet In Native Cellular Membranes Determined By In Situ Solid-State Nmr
Organism: Bradyrhizobium diazoefficiens usda 110
Method: SOLID-STATE NMR Release Date: 2025-09-10 Classification: MEMBRANE PROTEIN |

