Search Count: 63
All
Selected
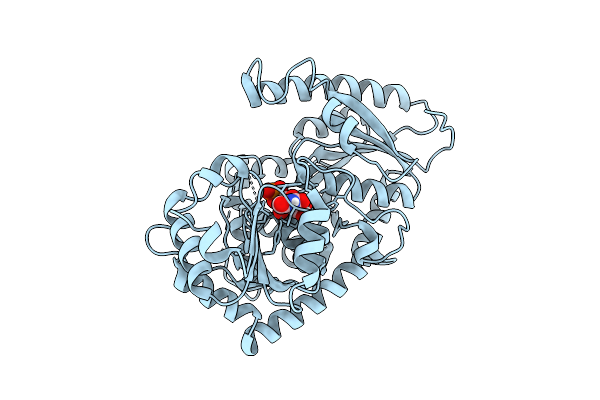 |
Crystal Structure Of The Glycosyltransferase Qsfuct From Quillaja Saponaria
Organism: Quillaja saponaria
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP |
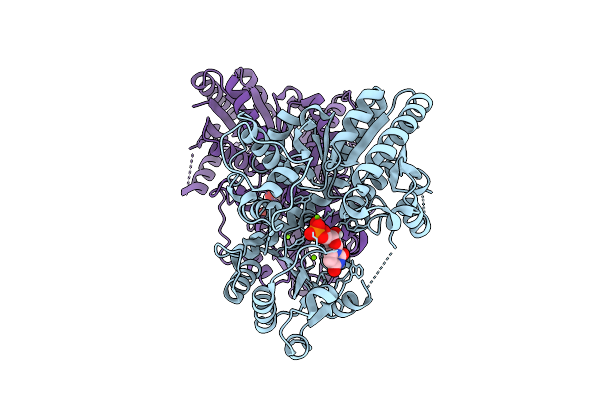 |
Crystal Structure Of The Glycosyltransferase Svfuct From Saponaria Vaccaria
Organism: Gypsophila vaccaria
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP, MG |
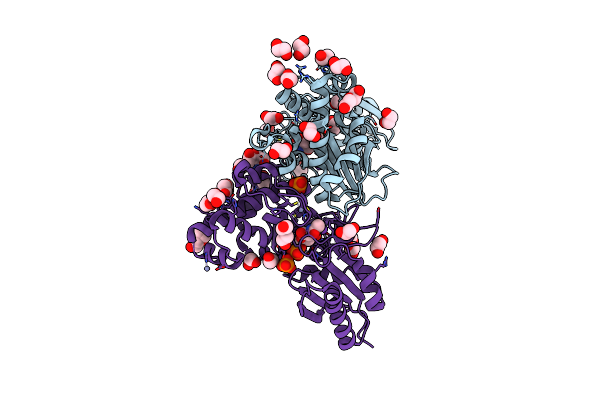 |
N-Acetylglucosamine Kinase From Plesiomonas Shigelloides Compexed With Alpha-N-Acetylglucosamine-6-Phosphate
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2022-08-10 Classification: SUGAR BINDING PROTEIN Ligands: EDO, PEG, 4QY, ZN, K, PO4, CL, TRS |
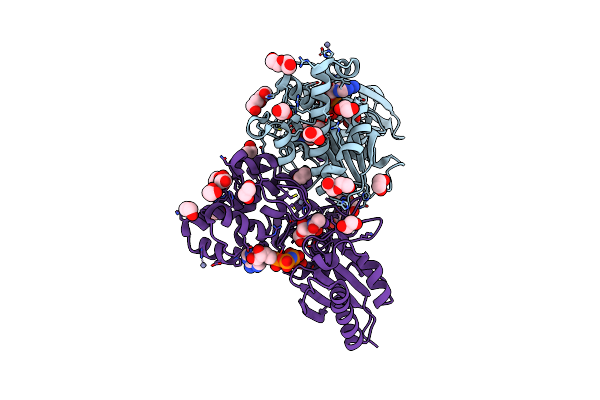 |
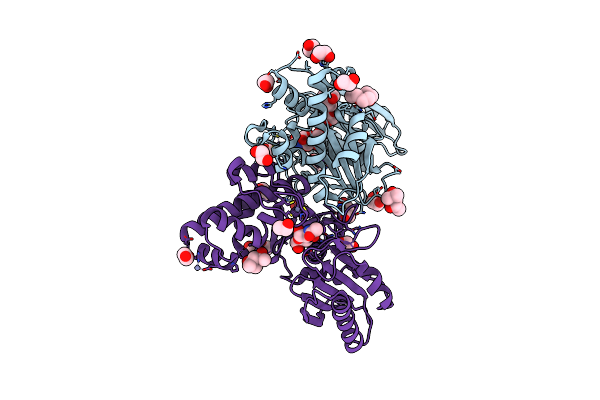
N-Acetylglucosamine Kinase From Plesiomonas Shigelloides Compexed With Alpha-N-Acetylglucosamine And Amp-Pnp Inhibitor
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2022-08-10 Classification: SUGAR BINDING PROTEIN Ligands: ZN, ANP, PEG, PGE, NDG, EDO, K |
 |
N-Acetylglucosamine Kinase From Plesiomonas Shigelloides Compexed With Alpha-N-Acetylglucosamine
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2022-08-10 Classification: SUGAR BINDING PROTEIN Ligands: MPD, EDO, PGE, NDG, ZN, K, CL, PEG |
 |
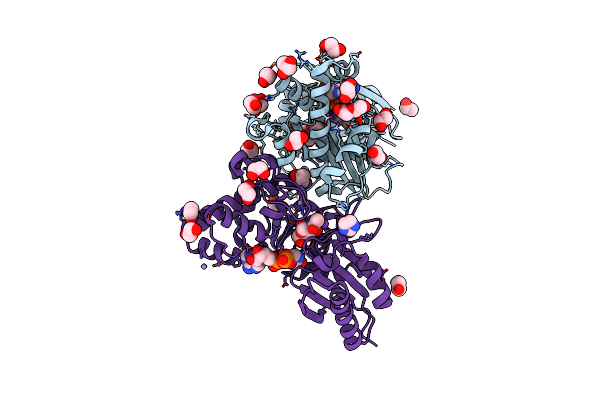
Structure Of N-Acetylglucosamine Kinase From Plesiomonas Shigelloides In Complex With Amp-Pnp In The Absence Of N-Acetylglucoseamine Substrate
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2022-08-10 Classification: SUGAR BINDING PROTEIN Ligands: PEG, ANP, ZN, PGE, EDO |
 |
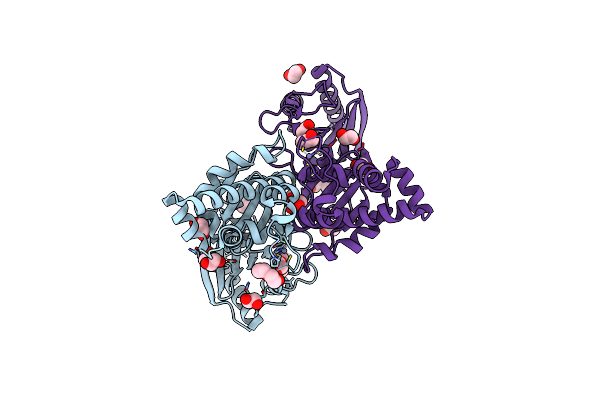
N-Acetylglucosamine Kinase From Plesiomonas Shigelloides Compexed With Alpha-N-Acetylglucosamine And Adp
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2022-08-03 Classification: SUGAR BINDING PROTEIN Ligands: ADP, GOL, IMD, NDG, ZN, EDO, PEG, K, IPA |
 |
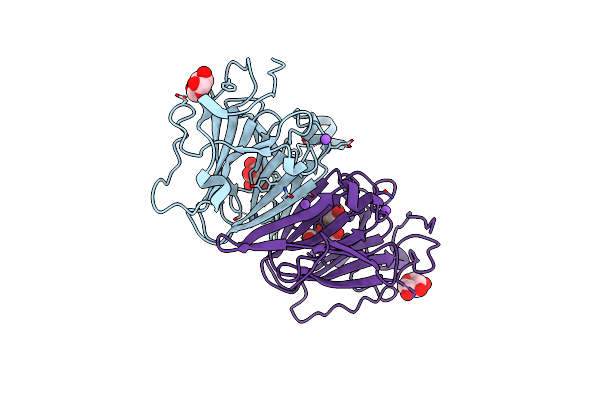
Native Structure Of N-Acetylglucosamine Kinase From Plesiomonas Shigelloides
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c), Plesiomonas shigelloides 302-73
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-07-27 Classification: SUGAR BINDING PROTEIN Ligands: PEG, TRS, ZN, PGE, EDO, CL |
 |
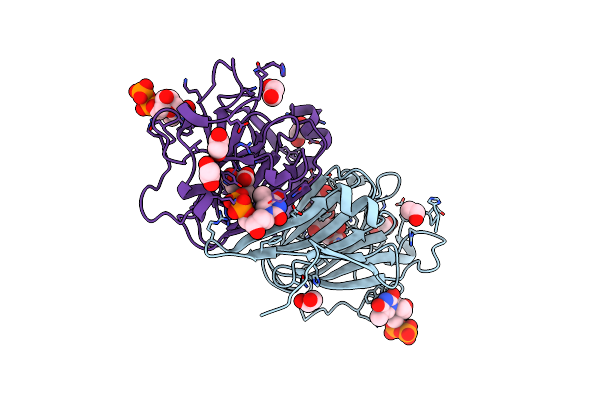
Organism: Coxiella burnetii (strain rsa 493 / nine mile phase i)
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2022-04-20 Classification: SUGAR BINDING PROTEIN Ligands: XYP, XYS, NA, FLC |
 |
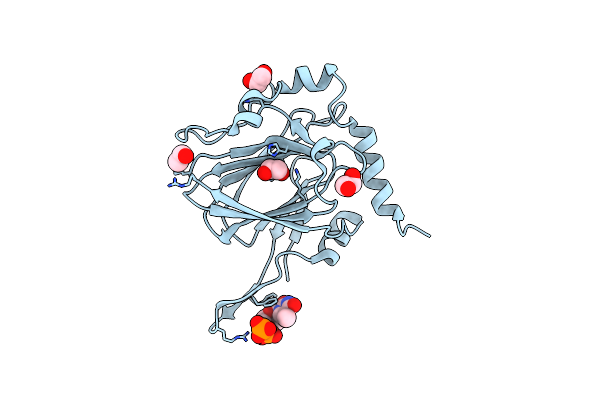
Organism: Coxiella burnetii (strain rsa 493 / nine mile phase i)
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2022-04-20 Classification: SUGAR BINDING PROTEIN Ligands: TYD, CIT, EDO |
 |
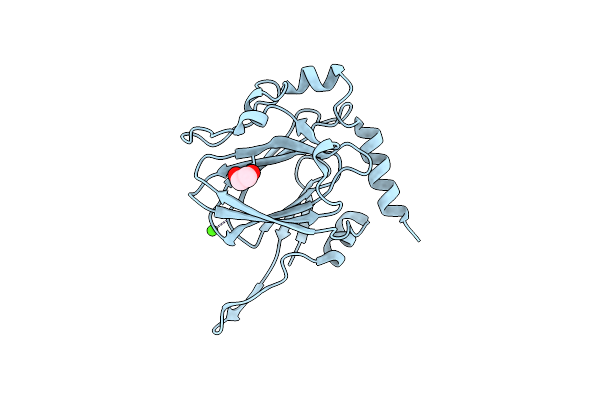
Organism: Streptomyces griseus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-04-20 Classification: SUGAR BINDING PROTEIN Ligands: EDO, TYD, CL |
 |
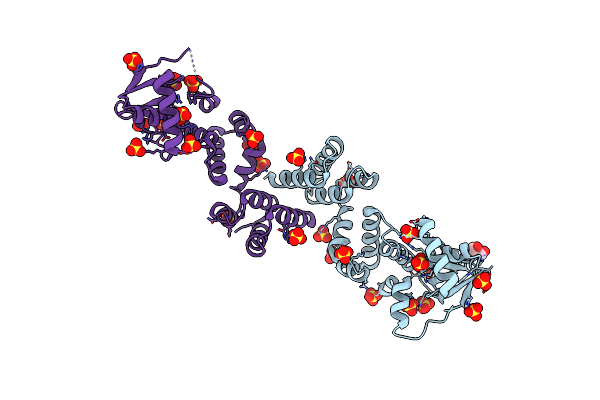
Organism: Streptomyces griseus
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2022-04-20 Classification: SUGAR BINDING PROTEIN Ligands: EDO, CA |
 |
Crystal Structure Of C-Terminally Truncated Geobacillus Thermoleovorans Nucleoid Occlusion Protein Noc
Organism: Geobacillus thermoleovorans ccb_us3_uf5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2021-02-17 Classification: DNA BINDING PROTEIN Ligands: SO4, GOL |
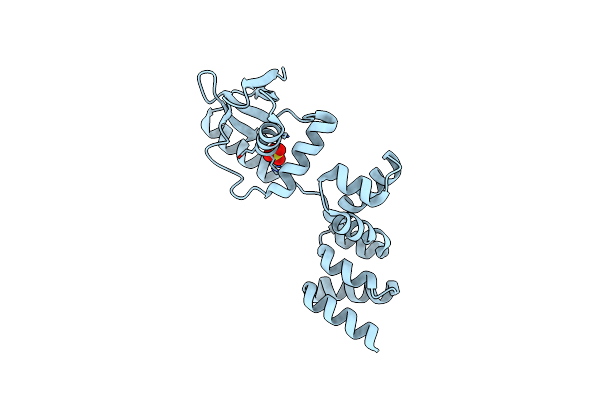 |
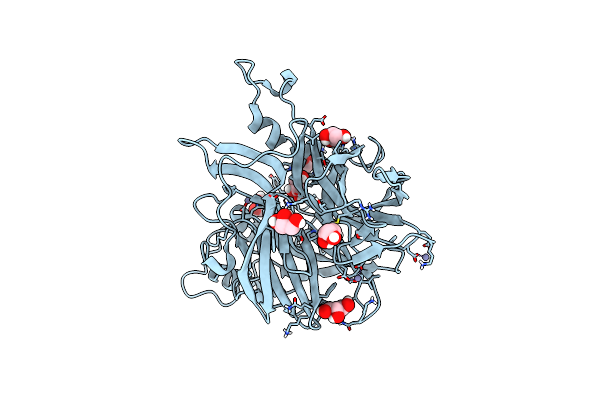
Crystal Structure Of N- And C-Terminally Truncated Geobacillus Thermoleovorans Nucleoid Occlusion Protein Noc
Organism: Geobacillus thermoleovorans ccb_us3_uf5
Method: X-RAY DIFFRACTION Resolution:2.95 Å Release Date: 2021-02-17 Classification: DNA BINDING PROTEIN Ligands: SO4 |
 |
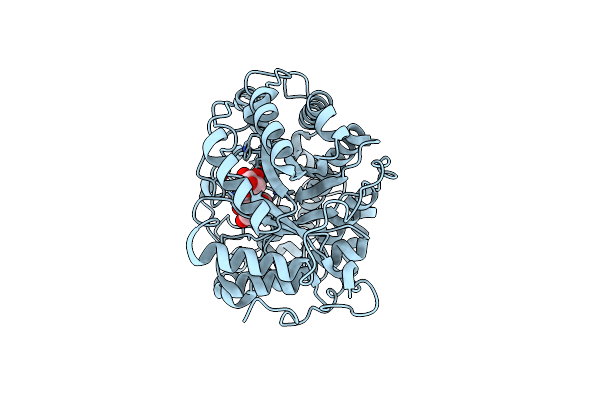
Organism: Erwinia tasmaniensis
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2019-02-06 Classification: TRANSFERASE Ligands: GOL, ZN |
 |
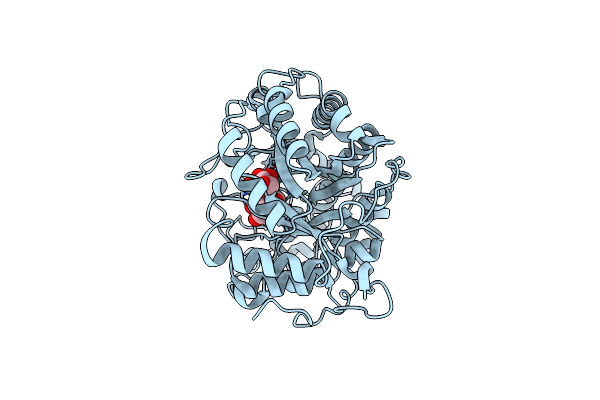
Crystal Structure Of Barley Beta-Amylase Complexed With 4-S-Alpha-D-Glucopyranosyl-(1,4-Dideoxy-4-Thio-Nojirimycin)
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2019-01-30 Classification: HYDROLASE Ligands: SGJ, GLC, CL |
 |
Crystal Structure Of Barley Beta-Amylase Complexed With 4-O-Alpha-D-Mannopyranosyl-(1-Deoxynojirimycin)
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2019-01-30 Classification: HYDROLASE Ligands: CL |
 |
Crystal Structure Of Barley Beta-Amylase Complexed With 3-Deoxy-3-Fluoro-Maltose
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2019-01-30 Classification: HYDROLASE Ligands: CL |
 |
Crystal Structure Of Sorbitol-6-Phosphate 2-Dehydrogenase Srld From Erwinia Amylovora
Organism: Erwinia amylovora cfbp1430
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2018-06-13 Classification: OXIDOREDUCTASE Ligands: CL |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25177 / h37ra)
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2018-05-02 Classification: TRANSFERASE Ligands: MG, G1P, ACO, PEG, EDO |

