Search Count: 18,445
All
Selected
 |
Crystal Structure Of Thioredoxin Gluthathione Reductase From Schistosoma Japonicum Sjtgr-Wt
Organism: Schistosoma japonicum
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of Thioredoxin Gluthathione Reductase From Schistosoma Japonicum With The U597C Mutation Sjtgr-U597C
Organism: Schistosoma japonicum
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of Thioredoxin Gluthathione Reductase From Schistosoma Japonicum With The U597C Mutation In Complex With Nadph
Organism: Schistosoma japonicum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, ATR, MLA |
 |
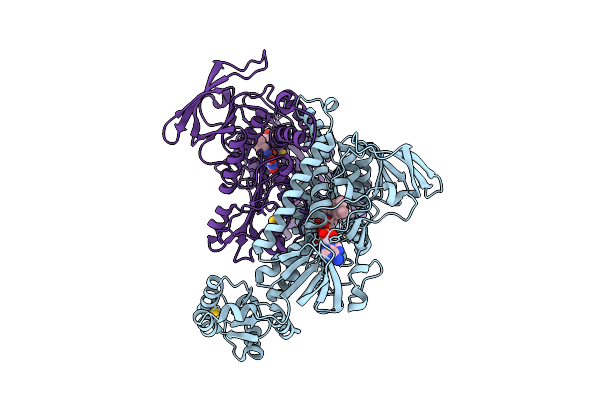
Crystal Structure Of Thioredoxin Gluthathione Reductase From Schistosoma Japonicum With The U597C Mutation In Complex With Gsh
Organism: Schistosoma japonicum
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GSH, SIN, NA |
 |
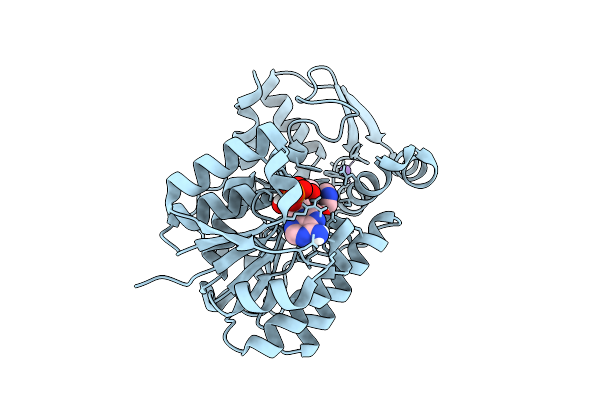
Crystal Structure Of Thioredoxin Gluthathione Reductase From Schistosoma Japonicum With The U597C Mutation In Complex With Auranofin
Organism: Schistosoma japonicum
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, AU |
 |
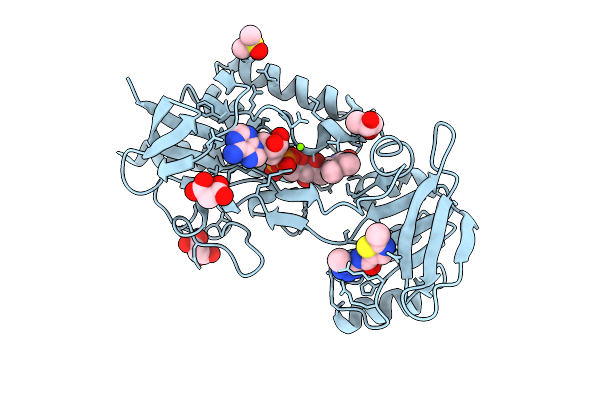
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN Ligands: NAP, NCA, NA |
 |
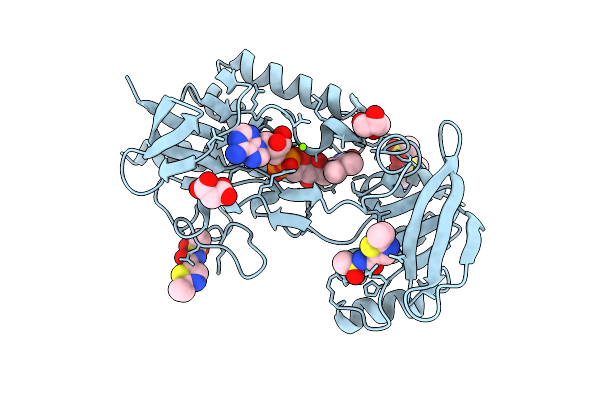
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2997505083
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1JAC, DMS |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z854627136
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I94 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z30008604
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I96 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2858787682
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I97 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z1280094148
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I95, DMS |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z166687084
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, DMS, A1I98 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z741560256
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I99 |
 |
Crystal Structure Of An Nta622L Variant In Complex With Nadp+ And Nicotinic Acid
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN Ligands: NIO, NAP |
 |
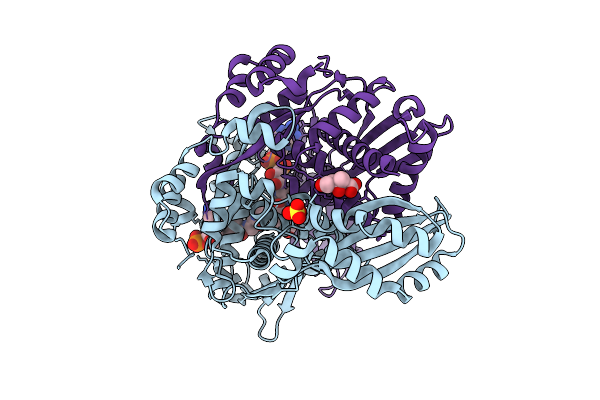
Crystal Structure Of (R)-Carbonyl Reductase Mutant I51L/Y61F/D147E From Candida Parapsilosis In Complex With Nad
Organism: Candida parapsilosis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-15 Classification: BIOSYNTHETIC PROTEIN Ligands: NAD, ZN |
 |
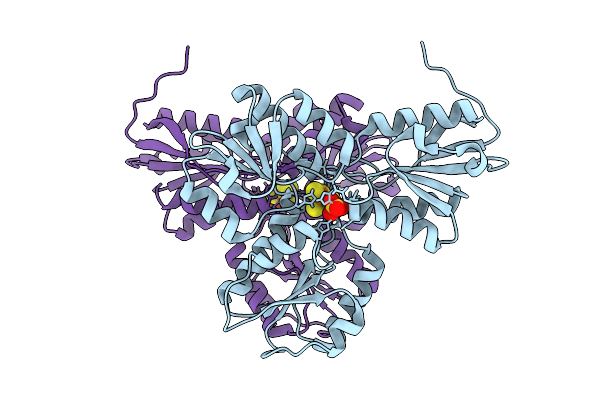
Structure Of Pmhmgr Bound To Mevalonate, Coa And Nad 2 Minutes 20 Seconds After Reaction Initiation At Ph 9
Organism: Pseudomonas sp. 'mevalonii'
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE/SUBSTRATE/INTERMEDIATE Ligands: COA, SO4, A1AFY, ZKE, NAI, MEV |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-08 Classification: OXIDOREDUCTASE Ligands: SF4, SO4 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
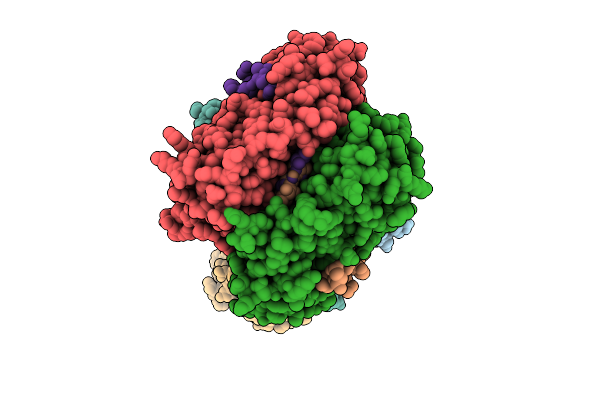 |
Cryoem Structure Of Human Mata2 In Complex With Mat2B Isoform V1 At 2.6 A Resolution
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
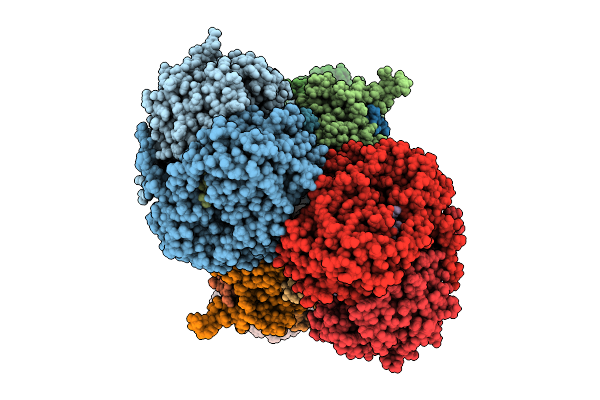 |
Cryo-Em Structure Of Complex Iii On The Bovine Heart Submitochondrial Particles, Iii-1
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MOTOR PROTEIN Ligands: HEM, HEC, FES |

