Search Count: 17
All
Selected
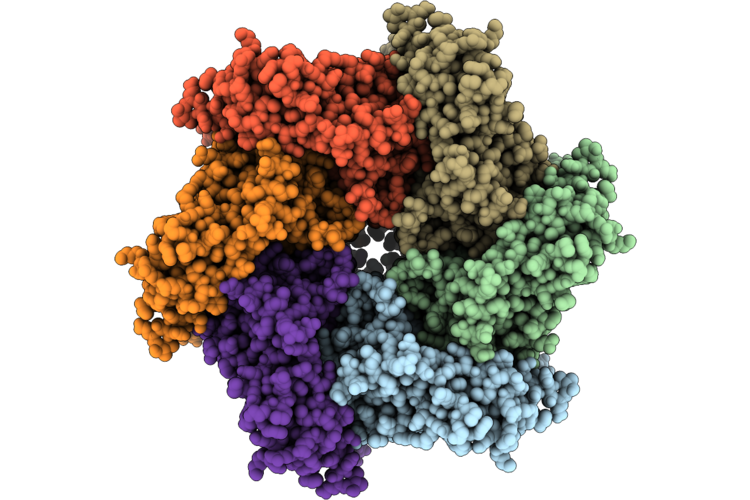 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ADP |
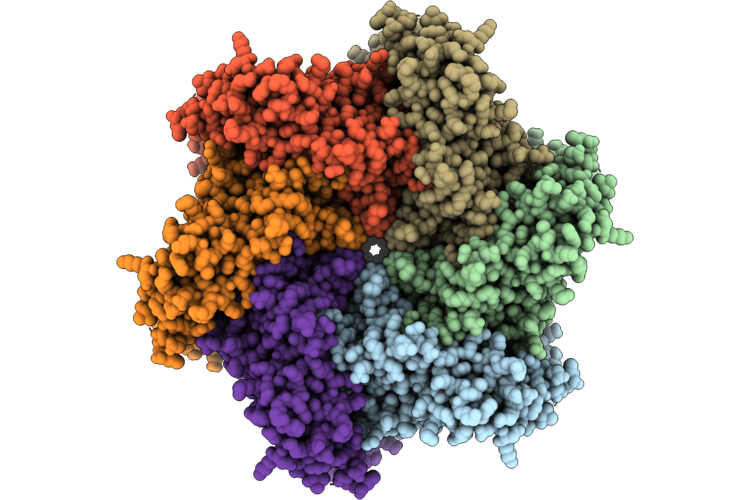 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.55 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: AGS, MG |
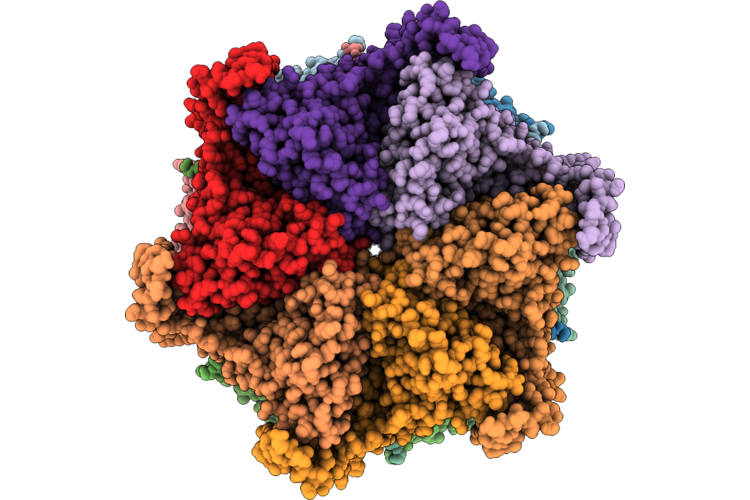 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.58 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: JDP |
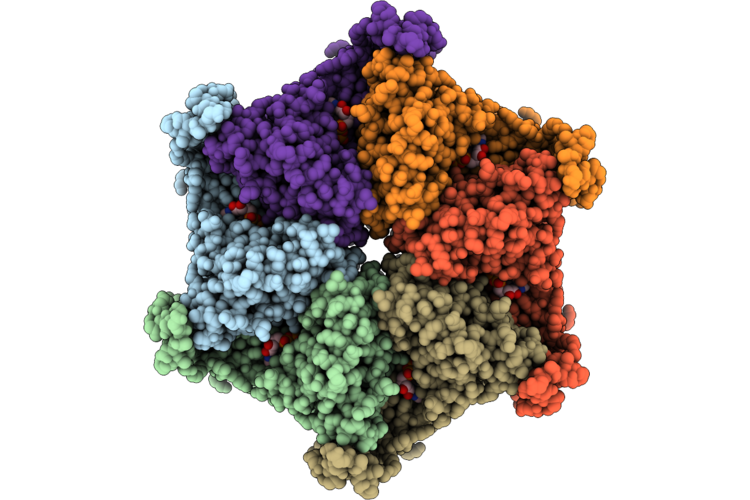 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.74 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ADP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.41 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: AGS, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.78 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ADP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: AGS, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.61 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ADP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.61 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: AGS, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2019-06-05 Classification: OXIDOREDUCTASE |
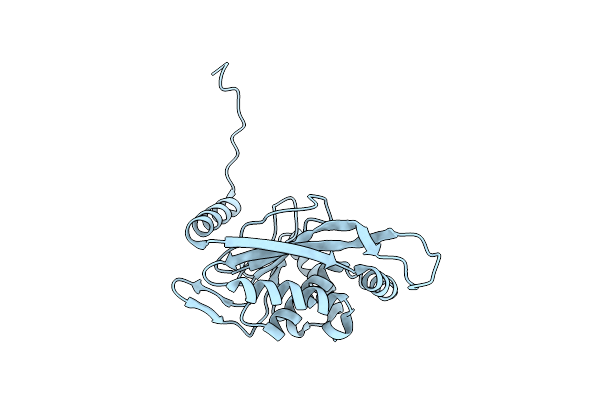 |
Pseudo-Atomic Structural Model Of The E3Bp Component Of The Human Pyruvate Dehydrogenase Multienzyme Complex
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2019-06-05 Classification: OXIDOREDUCTASE |
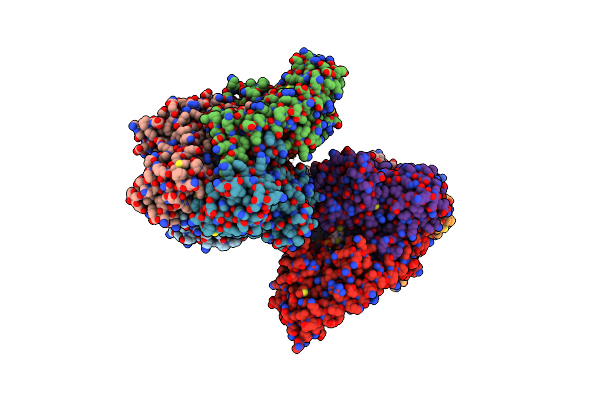 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2018-07-11 Classification: OXIDOREDUCTASE Ligands: TPP, MG |
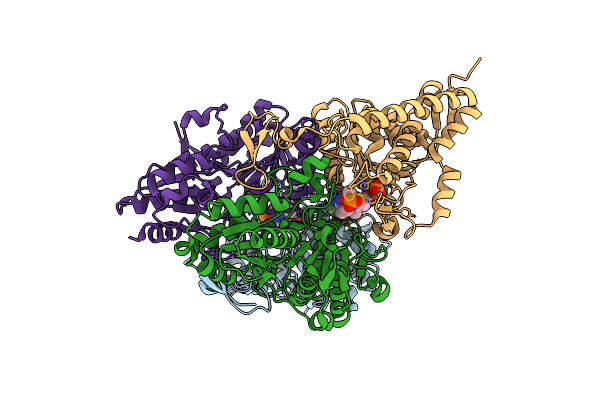 |
Human Pyruvate Dehydrogenase E1 Component Complex With Covalent Tdp Adduct Acetyl Phosphinate
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2018-07-11 Classification: OXIDOREDUCTASE Ligands: A5X, MG |
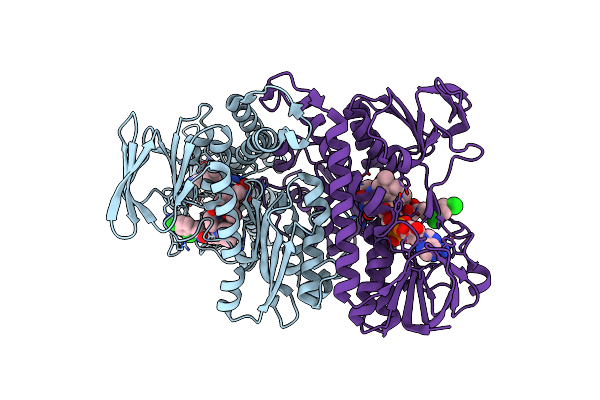 |
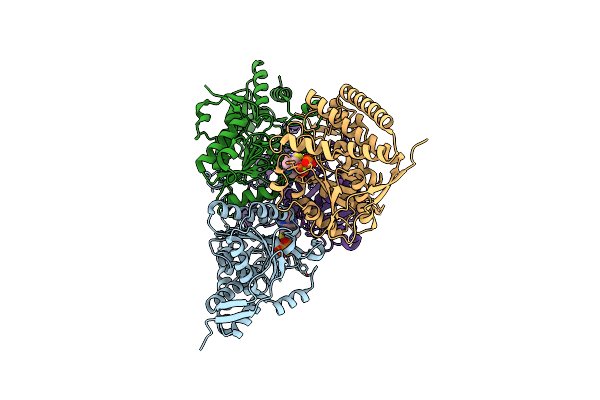
Structure Of Mycobacterial Lipoamide Dehydrogenase Bound To A Triazaspirodimethoxybenzoyl Inhibitor
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2010-01-26 Classification: OXIDOREDUCTASE Ligands: FAD, 3II |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2007-05-22 Classification: OXIDOREDUCTASE Ligands: MG, TPP, K |
 |
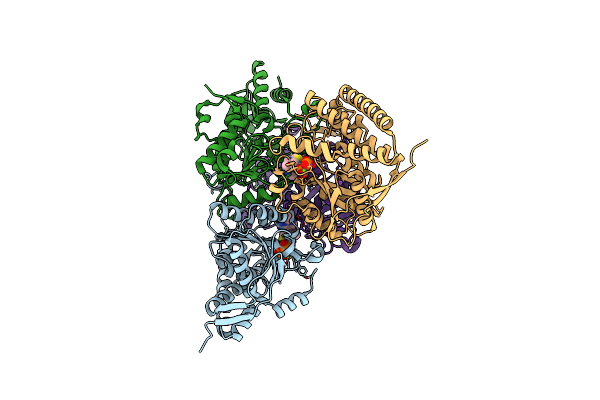
The Crystal Structure Of Dihydrolipoamide Dehydrogenase And Dihydrolipoamide Dehydrogenase-Binding Protein (Didomain) Subcomplex Of Human Pyruvate Dehydrogenase Complex.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2005-11-15 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2003-06-17 Classification: OXIDOREDUCTASE Ligands: MG, TPP, K |

