Search Count: 20
All
Selected
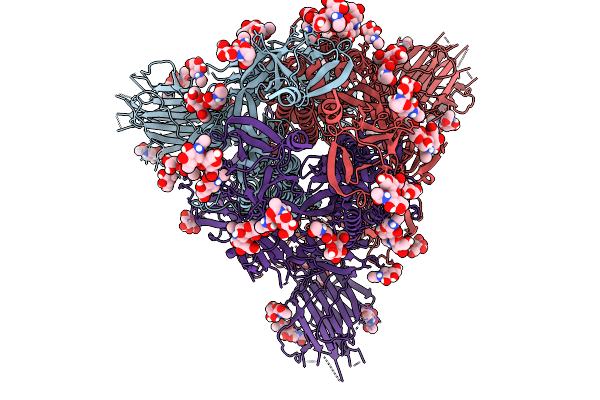 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
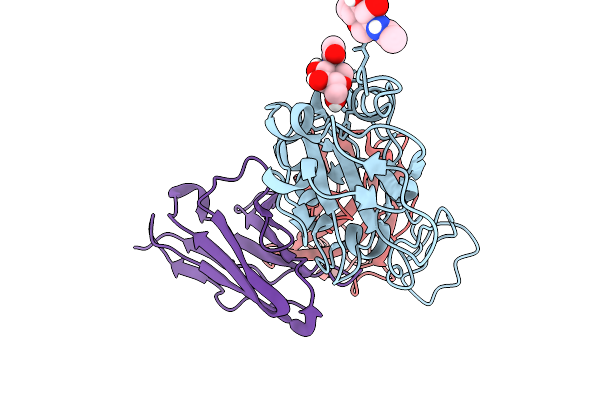 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 1 Up & 2 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
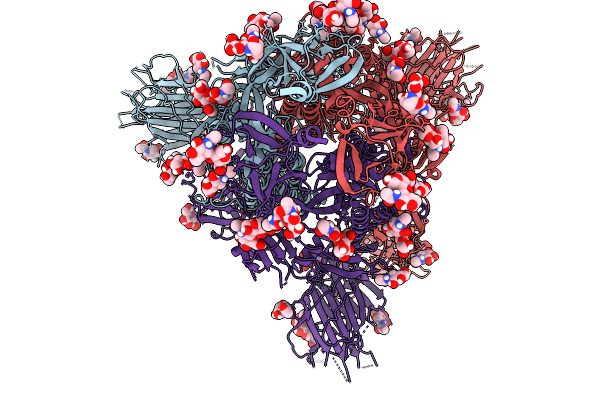 |
Cryo-Em Consensus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
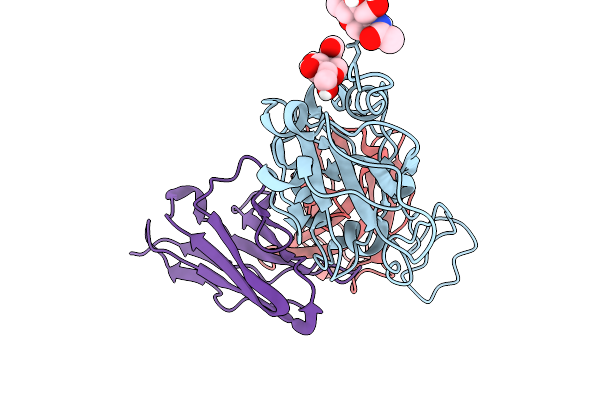 |
Cryo-Em Focus Map Of Prefusion Sars-Cov-2 Spike (Rbds: 2 Up & 1 Down) Bound To Rbd-Targeting Mo176-117 Antibody
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: NAG |
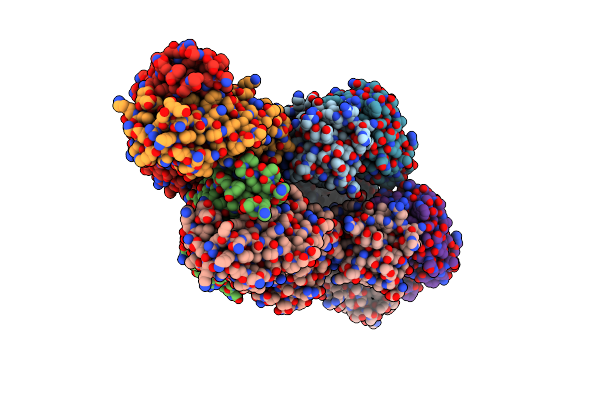 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-04-24 Classification: IMMUNE SYSTEM |
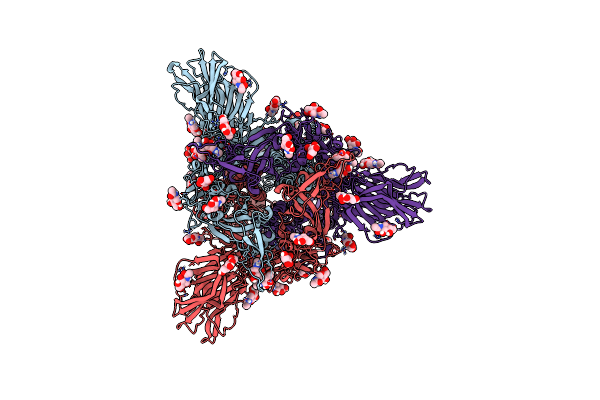 |
Organism: Severe acute respiratory syndrome coronavirus, Tequatrovirus t4
Method: ELECTRON MICROSCOPY Release Date: 2023-08-16 Classification: BIOSYNTHETIC PROTEIN Ligands: NAG |
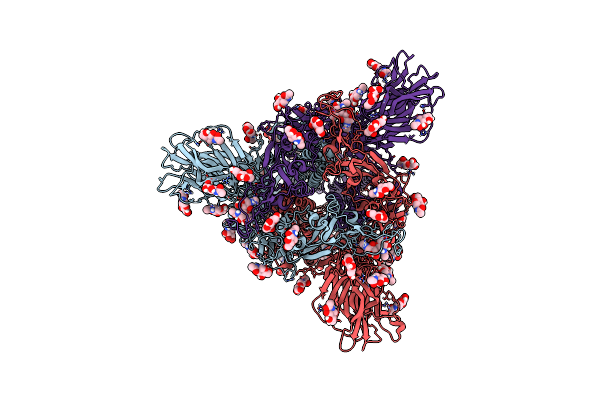 |
Organism: Severe acute respiratory syndrome coronavirus, Enterobacteria phage t4 (bacteriophage t4)
Method: ELECTRON MICROSCOPY Release Date: 2023-08-16 Classification: BIOSYNTHETIC PROTEIN Ligands: NAG |
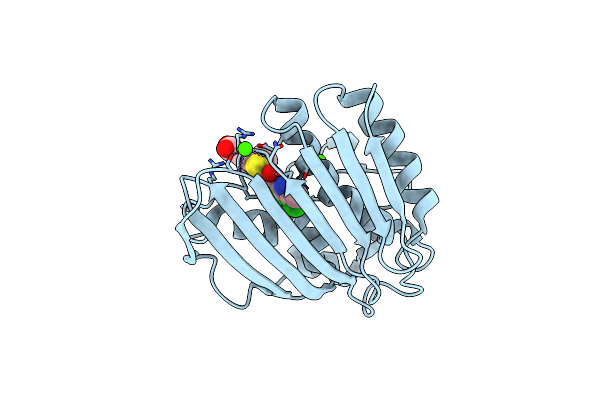 |
Pseudomonas Aeruginosa Dna Gyrase B 24Kda Atpase Subdomain Complexed With Ebl3021
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-03-29 Classification: DNA BINDING PROTEIN Ligands: CA, R53 |
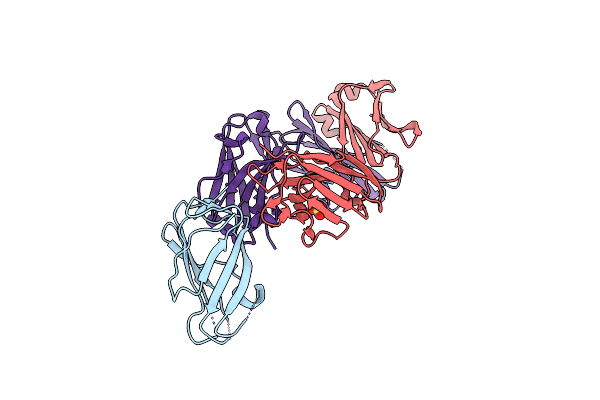 |
Crystal Structure Of Complex Between Recombinant Der P 2.0103 And Fab Fragment Of 7A1
Organism: Dermatophagoides pteronyssinus, Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2019-08-28 Classification: allergen/immune system Ligands: SO4 |
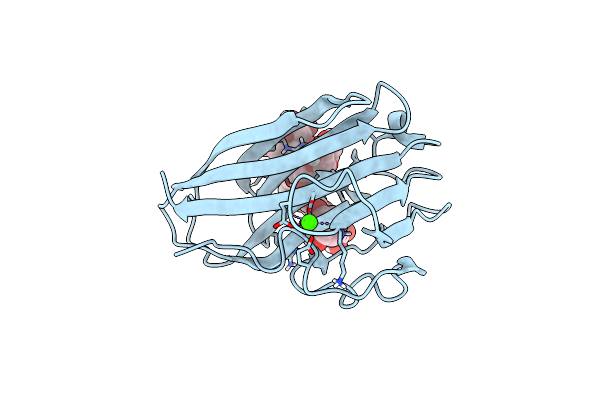 |
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.6 Å, 1.606 Å Release Date: 2015-10-28 Classification: SUGAR BINDING PROTEIN Ligands: CA, DOD |
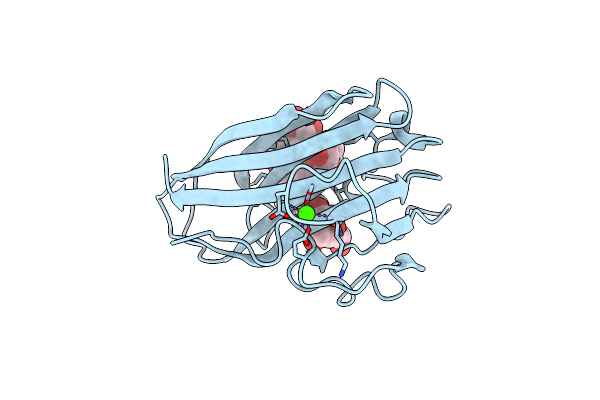 |
Xyloglucan Binding Module (Cbm4-2 X2-L110F) In Complex With Branched Xyloses
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2014-04-23 Classification: HYDROLASE Ligands: CA, GLC |
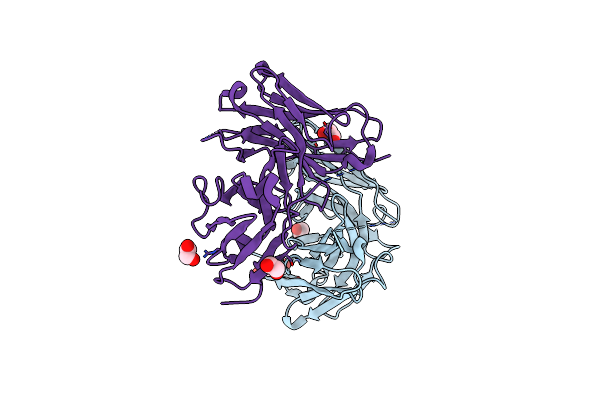 |
Human Ige Against The Major Allergen Bet V 1 - Crystal Structure Of Clone M0418 Scfv
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2013-11-20 Classification: IMMUNE SYSTEM Ligands: EDO, PEG |
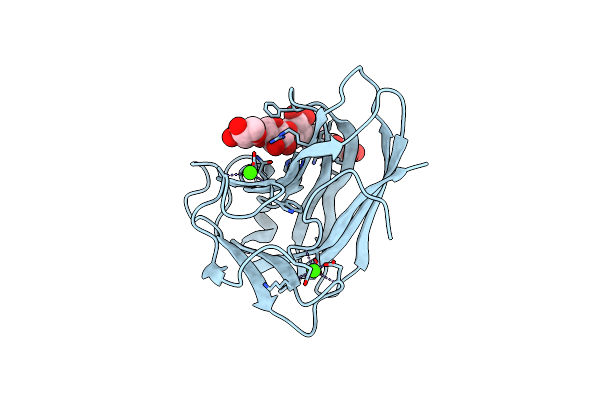 |
Xylopentaose Binding Mutated (X-2 L110F) Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA |
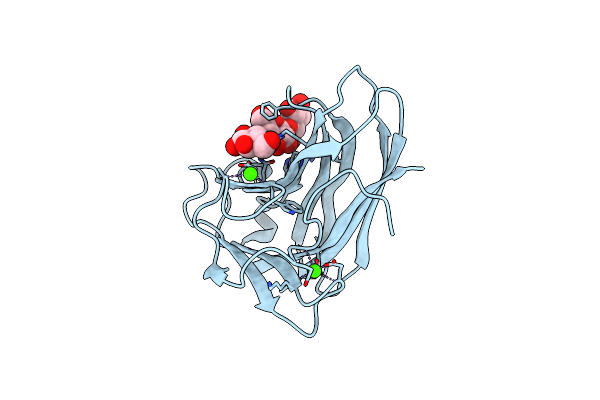 |
Cellopentaose Binding Mutated (X-2 L110F) Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA |
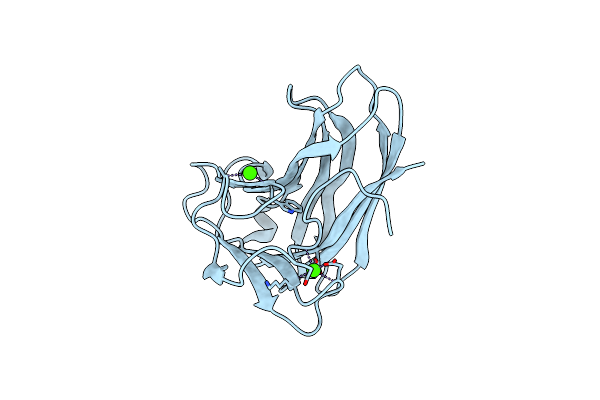 |
X-2 L110F Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.08 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA |
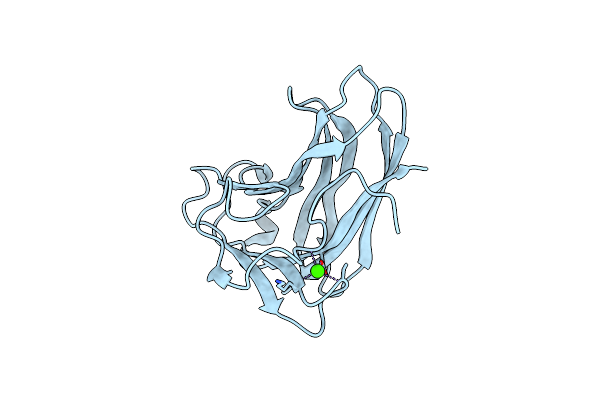 |
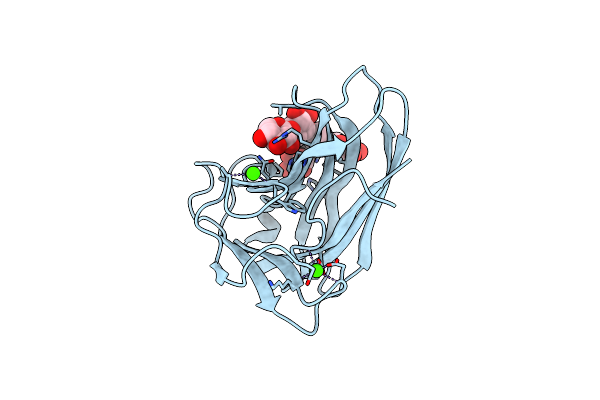
X-2 Engineered Mutated Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA |
 |
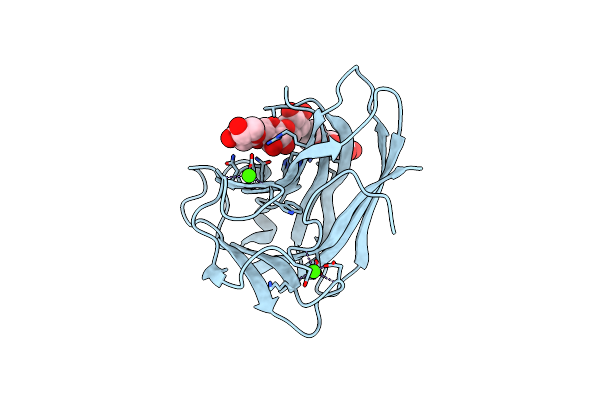
Xylotetraose Bound To X-2 Engineered Mutated Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.36 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA, CIT |
 |
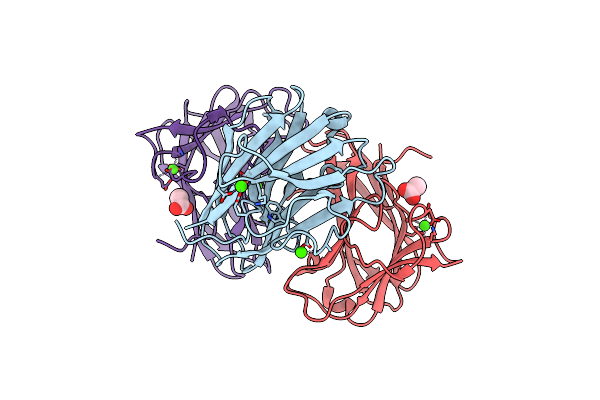
Xylopentaose Binding X-2 Engineered Mutated Cbm4-2 Carbohydrate Binding Module From A Thermostable Rhodothermus Marinus Xylanase
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2012-03-07 Classification: HYDROLASE Ligands: CA |
 |
Organism: Rhodothermus marinus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2009-10-27 Classification: hydrolase, carbohydrate-binding domain Ligands: CA, ACT |
 |
Three-Dimensional Structure Of A Human Fab With High Affinity For Tetanus Toxoid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 1998-02-04 Classification: IMMUNOGLOBULIN |

