Search Count: 158
All
Selected
 |
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: DE NOVO PROTEIN |
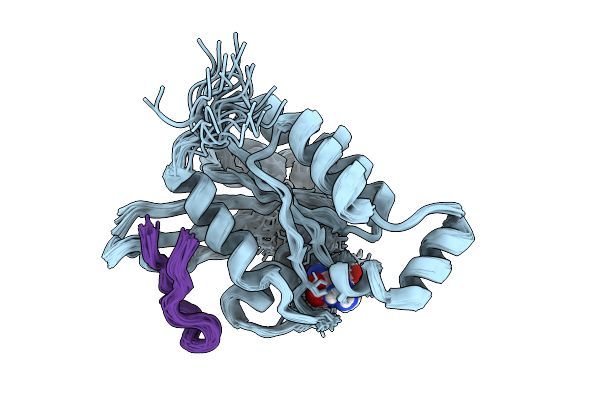 |
Organism: Homo sapiens, Synthetic construct
Method: SOLUTION NMR Release Date: 2026-01-21 Classification: CELL CYCLE Ligands: GNP, MG |
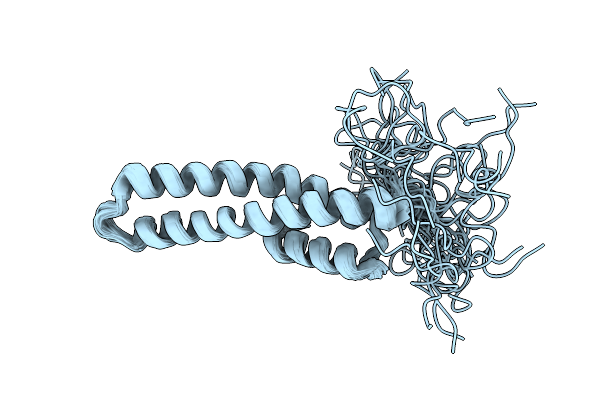 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-12-31 Classification: SIGNALING PROTEIN |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2025-10-22 Classification: STRUCTURAL PROTEIN |
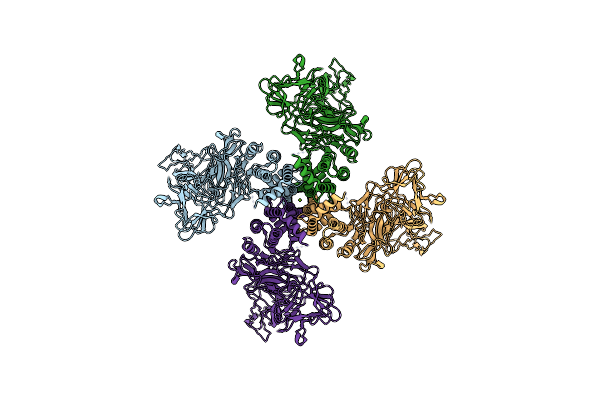 |
Organism: Paenibacillus popilliae
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: TOXIN Ligands: MG |
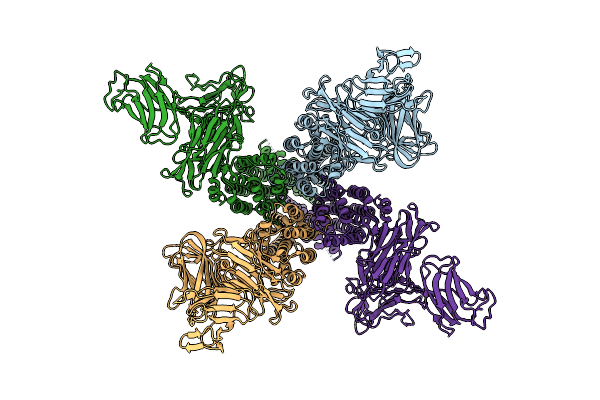 |
Organism: Paenibacillus popilliae
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: TOXIN |
 |
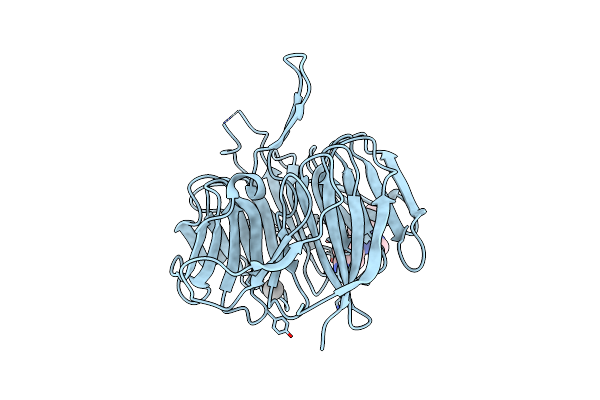
Organism: Bos taurus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-05-21 Classification: ENDOCYTOSIS |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.31 Å Release Date: 2025-02-26 Classification: TRANSPORT PROTEIN Ligands: A1BLV, UNX |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2025-02-05 Classification: TRANSFERASE Ligands: A1BLX |
 |
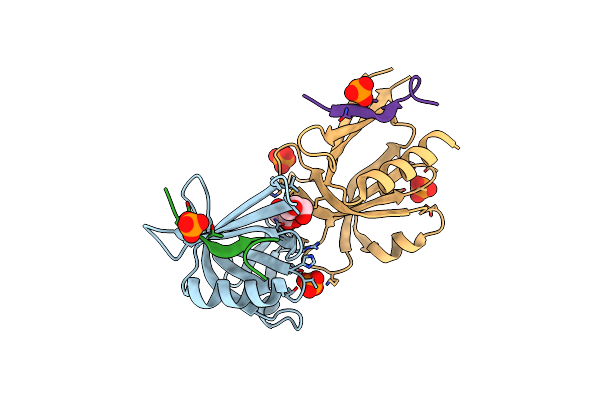
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2024-05-15 Classification: VIRAL PROTEIN Ligands: YDL |
 |
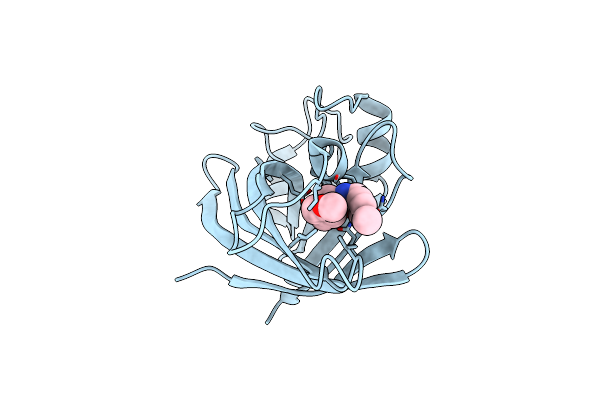
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2024-04-10 Classification: ENDOCYTOSIS Ligands: PO4, GOL |
 |
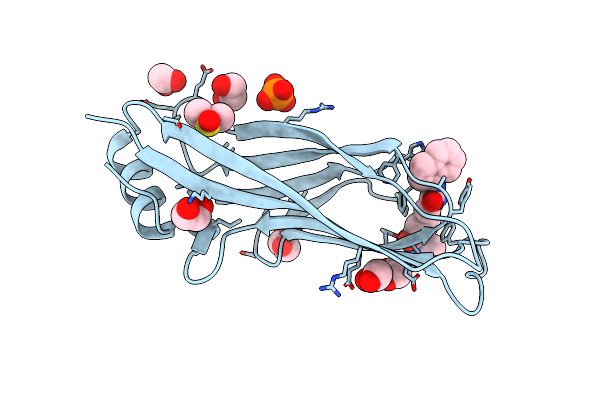
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2023-12-06 Classification: PEPTIDE BINDING PROTEIN Ligands: 98C |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.26 Å Release Date: 2023-11-22 Classification: PEPTIDE BINDING PROTEIN Ligands: ZJ9, EDO, DMS, PO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-11-22 Classification: PEPTIDE BINDING PROTEIN Ligands: EDO, DMS, ZJF, SO4 |
 |
Crystal Structure Of Human Yeats4 In Complex With Pfizer Small Molecule Compound 3B
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2023-01-11 Classification: PROTEIN BINDING Ligands: SJI |
 |
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2022-06-01 Classification: ENDOCYTOSIS Ligands: GOL, CIT |
 |
Organism: Rattus norvegicus, Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2022-06-01 Classification: ENDOCYTOSIS |
 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2022-06-01 Classification: ENDOCYTOSIS Ligands: SO4, MES |
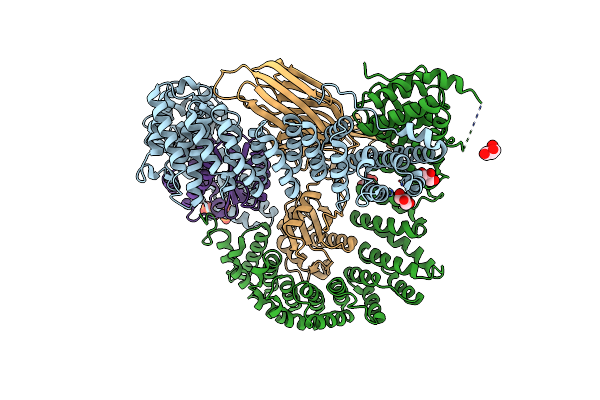 |
Organism: Rattus norvegicus, Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2022-06-01 Classification: ENDOCYTOSIS Ligands: IHP, GOL |

