Search Count: 17
All
Selected
 |
Structure Of The Arabidopsis Thaliana 80S Ribosome In Complex With P- And E-Site Trnas And Mrna
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:1.82 Å Release Date: 2026-02-04 Classification: RIBOSOME Ligands: MG, K, TER, SPD, EPE, ZN |
 |
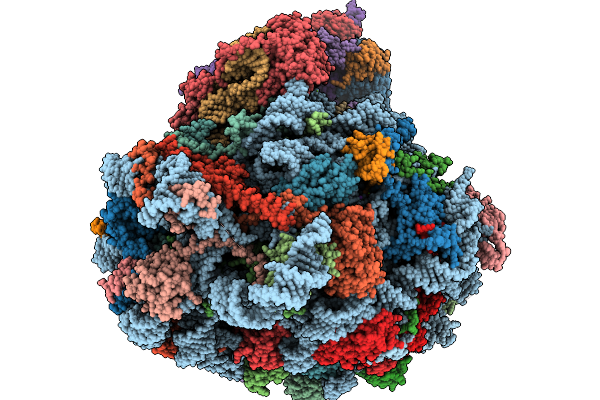
Structure Of The Arabidopsis Thaliana 80S Ribosome In Complex With P- And E-Site Trnas, Mrna, And Thermospermine
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.20 Å Release Date: 2026-02-04 Classification: RIBOSOME Ligands: TER, MG, K, SPD, EPE, ZN |
 |
Structure Of The Arabidopsis Thaliana 80S Ribosome Ovac Mutant In Complex With P- And E-Site Trnas And Mrna
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.25 Å Release Date: 2026-02-04 Classification: RIBOSOME Ligands: MG, K, TER, SPD, EPE, ZN |
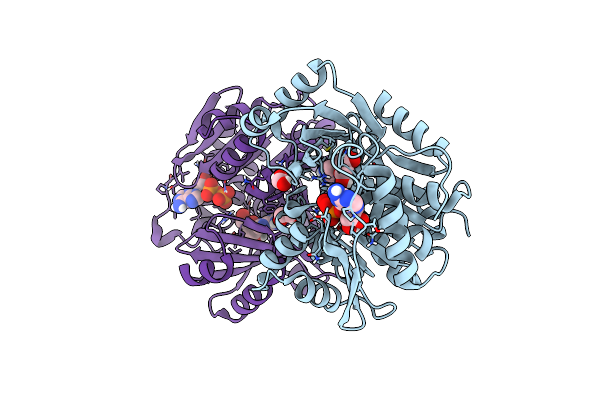 |
A Ternary Complex Of Plant Adenosine Kinase 1 From Moss Physcomitrella Patens (Ppadk1) With Adenosine And Adp
Organism: Physcomitrium patens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-04-09 Classification: TRANSFERASE Ligands: ADP, ADN, NA, EDO |
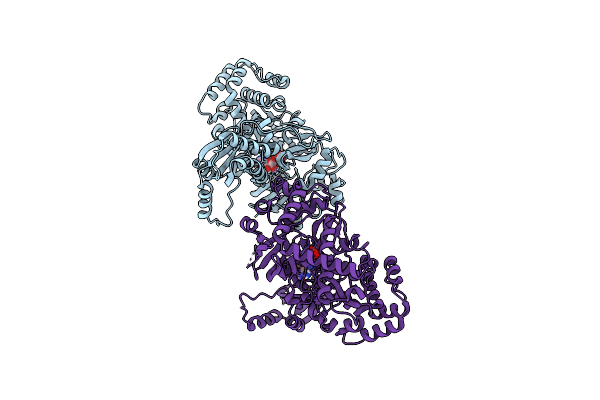 |
Structure Of Indole-3-Acetic Acid-Amido Synthetase Gh3.6 From A.Thaliana In Complex With Amp
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-04-09 Classification: LIGASE Ligands: AMP |
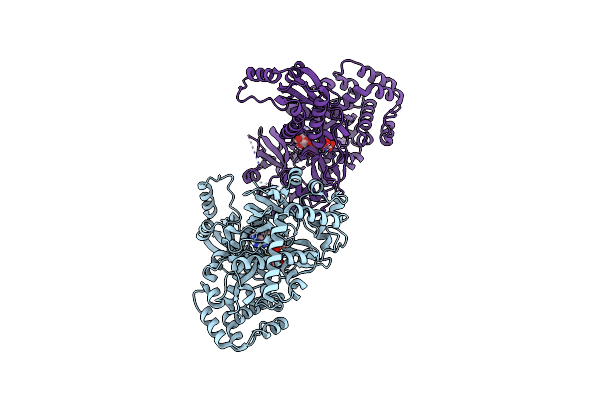 |
Structure Of Indole-3-Acetic Acid-Amido Synthetase Gh3.6 From A.Thaliana In Complex With Amp And Aspartate
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-04-09 Classification: LIGASE Ligands: AMP, ASP |
 |
Crystal Structure Of Zea Mays Adenosine Kinase 3 (Zmadk3) In Complex With Ap5A
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2025-01-29 Classification: TRANSFERASE Ligands: AP5, EDO, PEG, CL |
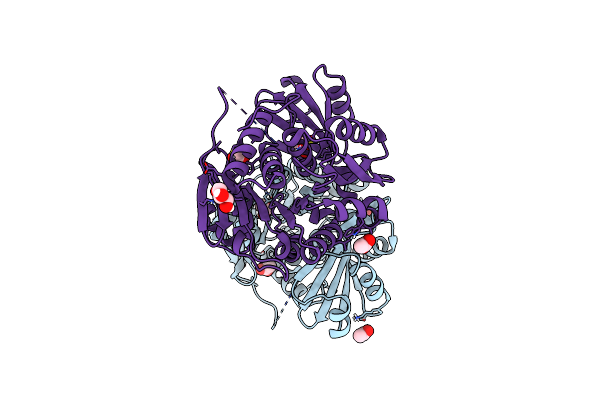 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2025-01-01 Classification: TRANSFERASE Ligands: SO4, ACT, GOL |
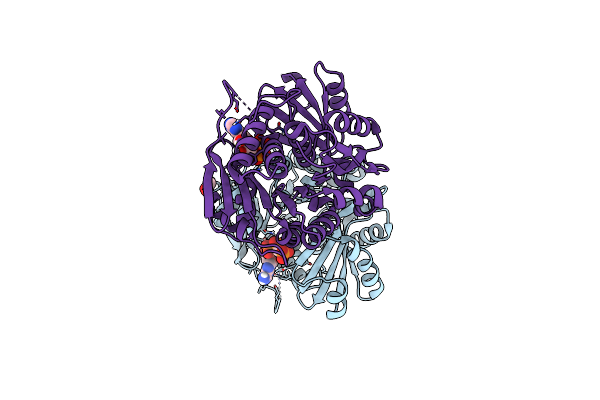 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2025-01-01 Classification: TRANSFERASE Ligands: ACP, ACT, GOL |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2022-01-12 Classification: HYDROLASE Ligands: CA, GOL, EDO, CL, PEG |
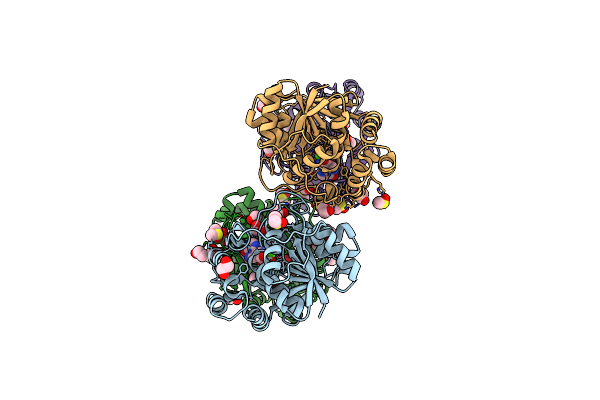 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2022-01-12 Classification: HYDROLASE Ligands: CA, IMH, DMS, EDO, PEG, PGE |
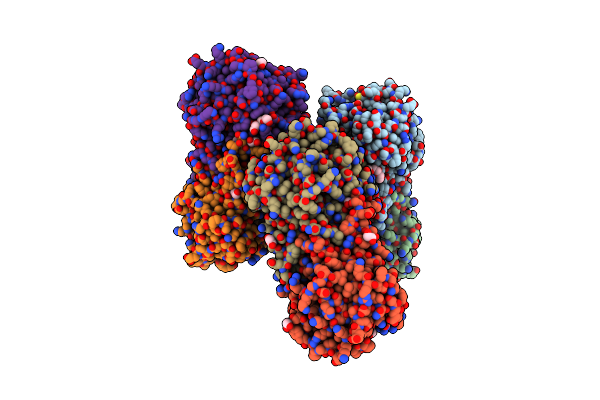 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2022-01-12 Classification: HYDROLASE Ligands: CA, RIB, EDO, PGE, PEG |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2022-01-12 Classification: HYDROLASE Ligands: CA, ADE, PEG, EDO |
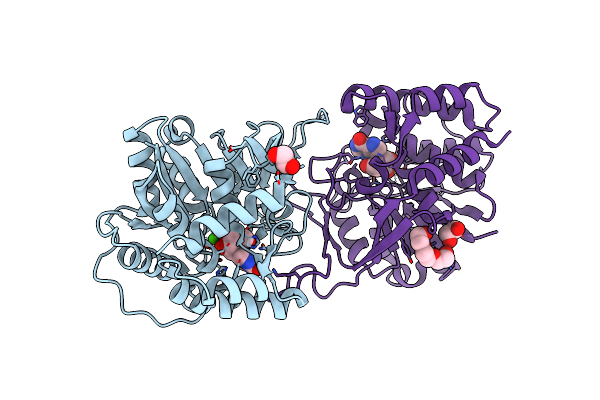 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-01-12 Classification: HYDROLASE Ligands: CA, IMH, EDO, PGE |
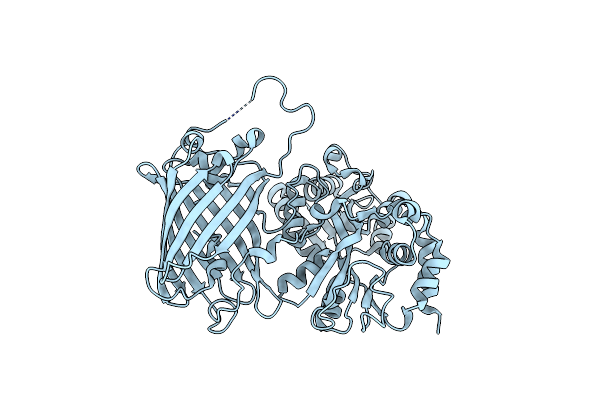 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2019-04-03 Classification: FLUORESCENT PROTEIN |
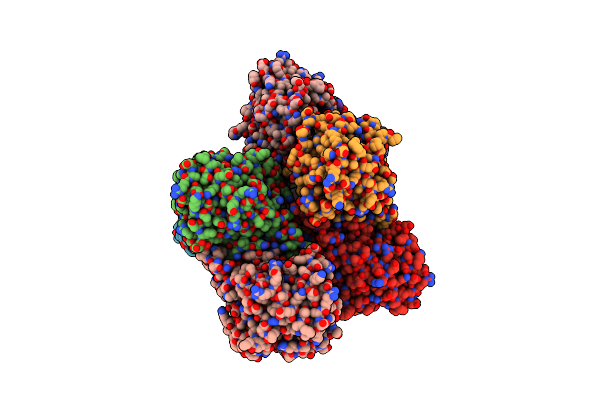 |
Organism: Physcomitrella patens
Method: X-RAY DIFFRACTION Resolution:3.35 Å Release Date: 2013-11-27 Classification: HYDROLASE Ligands: CA |
 |
Organism: Zea mays
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2013-11-27 Classification: HYDROLASE Ligands: CA |

