Search Count: 318
All
Selected
 |
Organism: Cypridina noctiluca
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-04-08 Classification: LUMINESCENT PROTEIN Ligands: NAG |
 |
Organism: Cypridina noctiluca
Method: X-RAY DIFFRACTION Resolution:3.14 Å Release Date: 2026-04-08 Classification: LUMINESCENT PROTEIN Ligands: NAG, A1L7I |
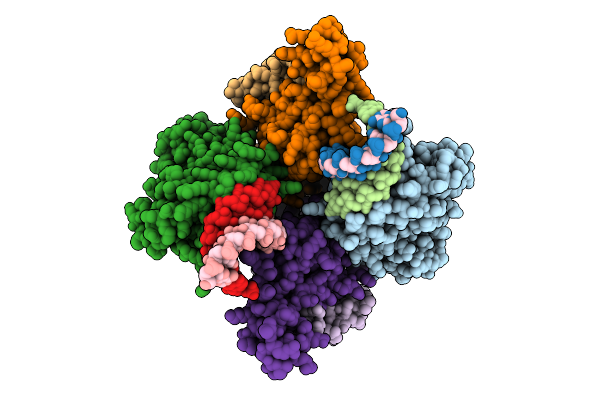 |
Structural Insights Into Hiv-1 Vif-Mediated Ubiquitination And Degradation Of Apobec3H
Organism: Pan troglodytes, Homo sapiens, Human immunodeficiency virus 1, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: VIRAL PROTEIN/RNA Ligands: ZN |
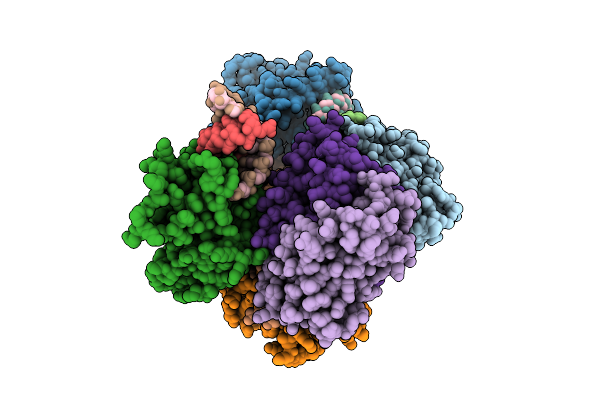 |
Structural Insights Into Hiv-1 Vif-Mediated Ubiquitination And Degradation Of Apobec3H
Organism: Pan troglodytes, Homo sapiens, Human immunodeficiency virus 1, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: VIRAL PROTEIN/RNA Ligands: ZN |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: POV, AV0, ERG |
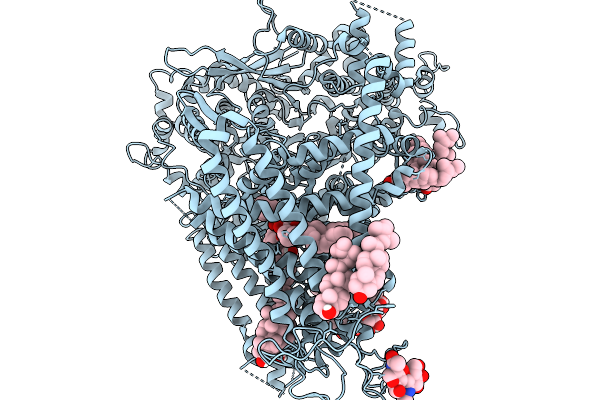 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
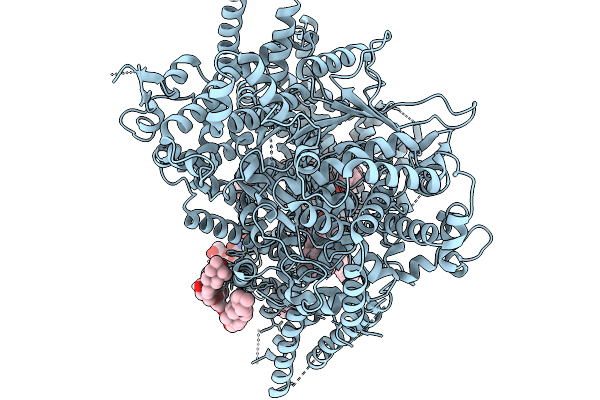
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: AV0, ERG, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
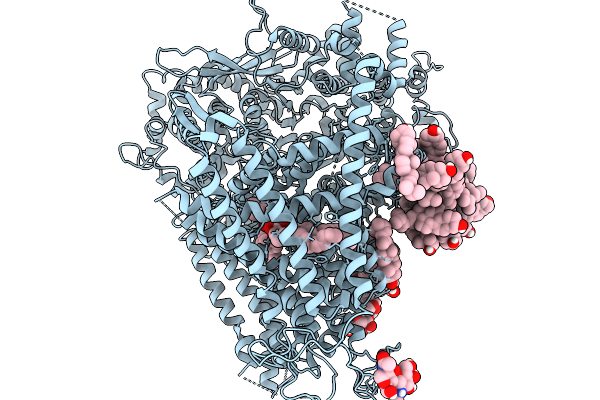
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN |
 |
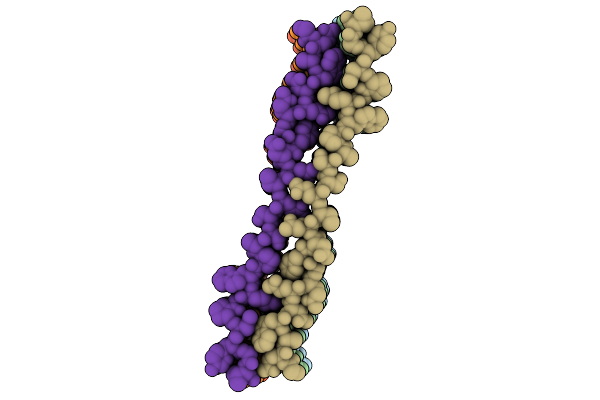
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: ERG, A1E06, POV |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: PROTEIN FIBRIL |
 |
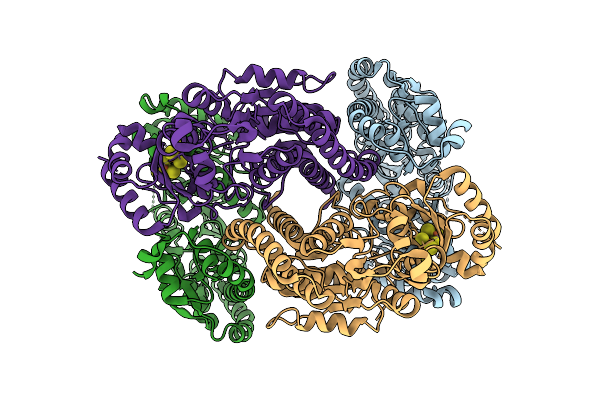
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: PROTEIN FIBRIL |
 |
Rhodospirillum Rubrum Nitrogenase-Like Methylthio-Alkane Reductase Complex With An Oxidized P-Cluster
Organism: Rhodospirillum rubrum atcc 11170
Method: ELECTRON MICROSCOPY Resolution:2.35 Å Release Date: 2025-11-05 Classification: OXIDOREDUCTASE Ligands: CLF |
 |
3-Hydroxypropionyl-Coa Synthetase (Adp-Forming) From Nitrosopumilus Maritimus.
Organism: Nitrosopumilus maritimus scm1
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-10-01 Classification: LIGASE Ligands: ANP, 3OH, PO4, MG |
 |
Crystal Structure Of Theophylline Aptamer Obtained In The Presence Of Caffeine
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2025-09-03 Classification: RNA Ligands: NCO, K |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2025-09-03 Classification: RNA Ligands: NCO, K |
 |
Cryo-Em Structure Of The Pma1 With Ordered N-Terminal Extension In The Activated State
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Release Date: 2025-06-04 Classification: PROTON TRANSPORT |
 |
Cryo-Em Structure Of The Pma1 With Ordered N-Terminal Extension In The Autoinhibited State
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: ELECTRON MICROSCOPY Resolution:3.52 Å Release Date: 2025-06-04 Classification: PROTON TRANSPORT |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:0.80 Å Release Date: 2025-05-28 Classification: HYDROLASE Ligands: GD, DO3, NO3, CL, NA |

