Search Count: 45
All
Selected
 |
Cryo-Em Structure Of Ni06063_D30_103 Fab In Complex With Influenza Virus Hemagglutinin From A/Hong Kong/485197/2014 (H3N2)
Organism: Homo sapiens, Influenza a virus (a/hong kong/485197/2014(h3n2))
Method: ELECTRON MICROSCOPY Resolution:2.62 Å Release Date: 2026-02-04 Classification: Viral protein/Immune System Ligands: NAG |
 |
Cryo-Em Structure Of Ni06063_D30_103 Fab In Complex With Influenza Virus Hemagglutinin From A/Michigan/45/2015 (H1N1)
Organism: Influenza a virus (a/michigan/45/2015(h1n1)), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-02-04 Classification: Viral protein/Immune System Ligands: NAG |
 |
Cryo-Em Structure Of Ni04359_D30_240 Fab In Complex With Influenza Virus Hemagglutinin From A/Hong Kong/485197/2014 (H3N2)
Organism: Influenza a virus (a/hong kong/485197/2014(h3n2)), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-02-04 Classification: Viral protein/Immune System Ligands: NAG |
 |
Cryo-Em Structure Of Ni04359_D30_240 Fab In Complex With Influenza Virus Hemagglutinin From A/Michigan/45/2015 (H1N1)
Organism: Influenza a virus (a/michigan/45/2015(h1n1)), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-02-04 Classification: Viral protein/Immune System Ligands: NAG |
 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: VIRUS Ligands: NAG |
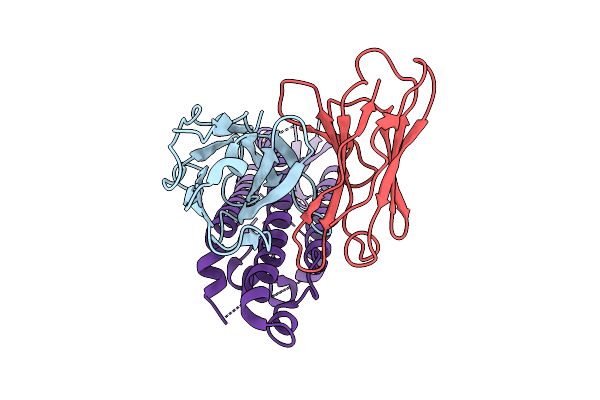 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-11-26 Classification: VIRUS |
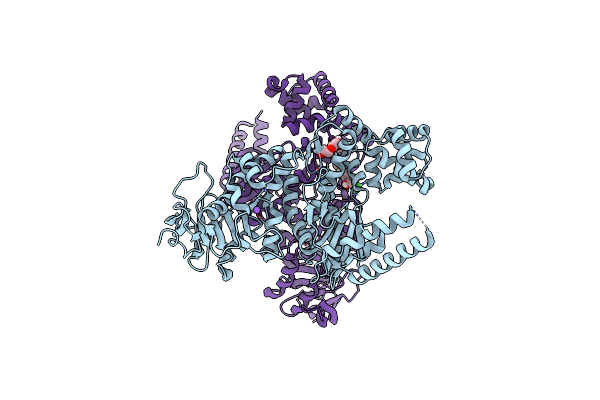 |
Organism: Plasmodium falciparum 3d7
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-05-14 Classification: REPLICATION |
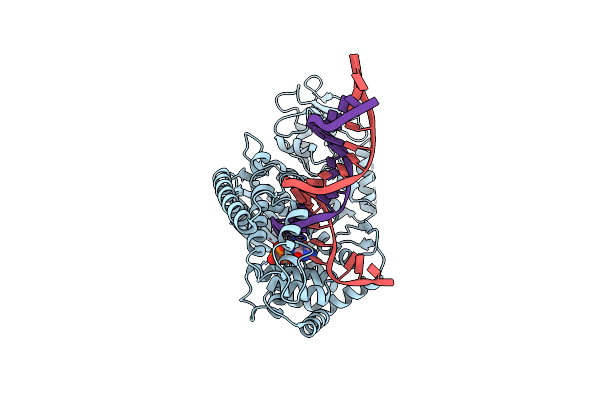 |
Ternary Structure Of Plasmodium Falciparum Apicoplast Dna Polymerase (Exo-Minus)
Organism: Homo sapiens, Plasmodium falciparum (isolate 3d7)
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: DNA BINDING PROTEIN/DNA Ligands: MG, DGT |
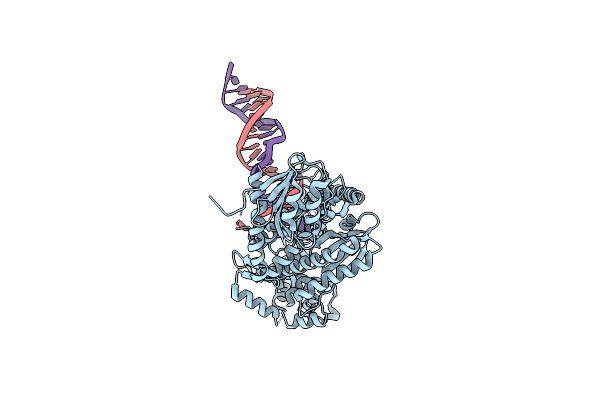 |
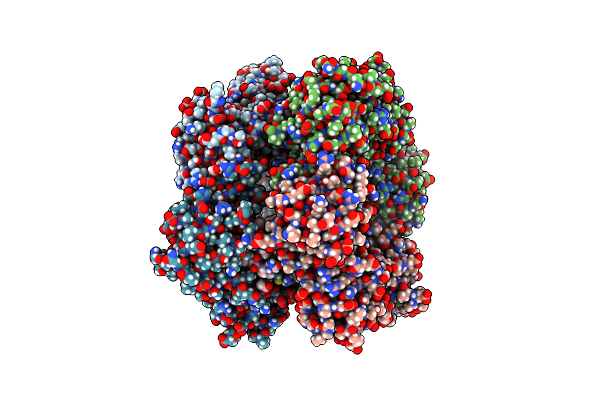
Structure Of Plasmodium Falciparum Apicoplast Dna Polymerase In Complex With Dna (Exo-Minus)
Organism: Plasmodium falciparum 3d7, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: DNA BINDING PROTEIN/DNA |
 |
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-07-26 Classification: DNA BINDING PROTEIN Ligands: FMT, ACT |
 |
Organism: Mycolicibacterium thermoresistibile atcc 19527, Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-01-18 Classification: DNA BINDING PROTEIN Ligands: ACE, NH2 |
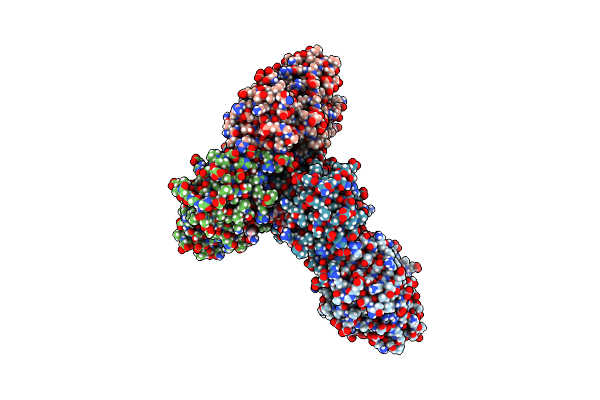 |
Organism: Mycolicibacterium thermoresistibile, Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-01-18 Classification: DNA BINDING PROTEIN Ligands: SO4, CL, ACE, NH2 |
 |
Organism: Mycolicibacterium thermoresistibile atcc 19527
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2023-01-18 Classification: DNA BINDING PROTEIN |
 |
Plasmodium Falciparum Apicoplast Dna Polymerase (Exo-Minus) Without Affinity Tag
Organism: Plasmodium falciparum (isolate 3d7)
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2022-10-26 Classification: REPLICATION, Transferase Ligands: GOL, NA, CL, PEG, EDO |
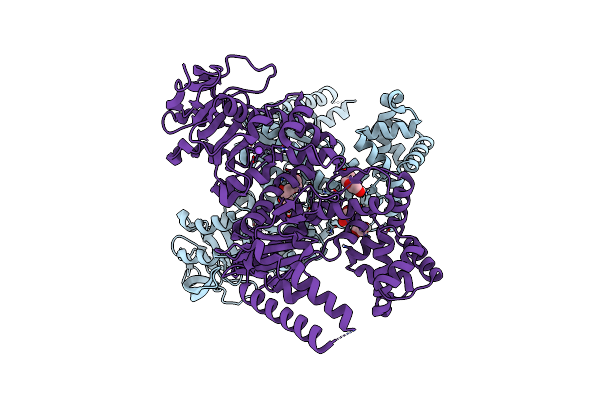 |
Plasmodium Falciparum Apicoplast Dna Polymerase (Exo-Minus) Without Affinity Tag
Organism: Plasmodium falciparum (isolate 3d7)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2022-10-19 Classification: REPLICATION, TRANSFERASE Ligands: PEG, CL, NA, EDO |
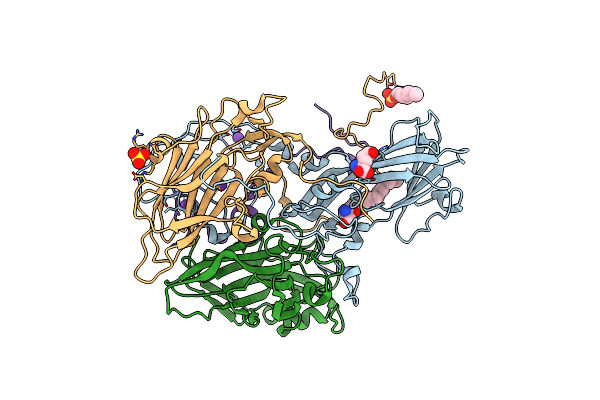 |
Crystal Structure Of Bovine Enterovirus 2 Determined With Serial Femtosecond X-Ray Crystallography
Organism: Enterovirus e
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2017-08-30 Classification: VIRUS Ligands: SPH, K, GLU, OSF, SO4 |
 |
Organism: Operophtera brumata cypovirus 18
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2017-06-21 Classification: VIRAL PROTEIN Ligands: CL, GTP, ATP, MG |
 |
Structure Of Pas-Gaf Fragment Of Deinococcus Phytochrome By Serial Femtosecond Crystallography
Organism: Deinococcus radiodurans (strain atcc 13939 / dsm 20539 / jcm 16871 / lmg 4051 / nbrc 15346 / ncimb 9279 / r1 / vkm b-1422)
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2017-02-22 Classification: TRANSFERASE Ligands: LBV, NI, CL, EDO |
 |
Structure Of The Photosensory Module Of Deinococcus Phytochrome By Serial Femtosecond X-Ray Crystallography
Organism: Deinococcus radiodurans (strain atcc 13939 / dsm 20539 / jcm 16871 / lmg 4051 / nbrc 15346 / ncimb 9279 / r1 / vkm b-1422)
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2017-02-22 Classification: TRANSFERASE Ligands: LBV |
 |
Organism: Thermosynechococcus elongatus (strain bp-1)
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2016-11-23 Classification: ELECTRON TRANSPORT Ligands: OEX, FE2, SQD, CL, CLA, PHO, BCR, PL9, LMG, UNL, LHG, BCT, DGD, HEM, HEC |

