Search Count: 489
All
Selected
 |
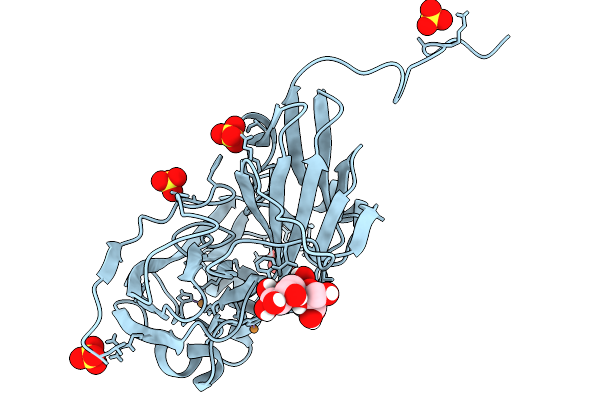
Atomic Resolution (1.02 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, NO2, GLC, FRU, SO4 |
 |
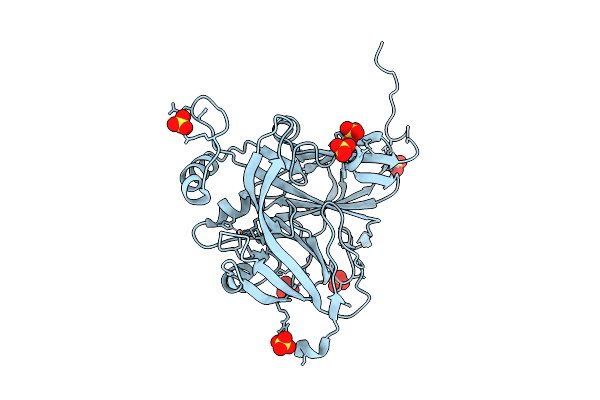
Atomic Resolution (1.00 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At High Ph (7.3) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium diazoefficiens usda 110
Method: X-RAY DIFFRACTION Resolution:1.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4 |
 |
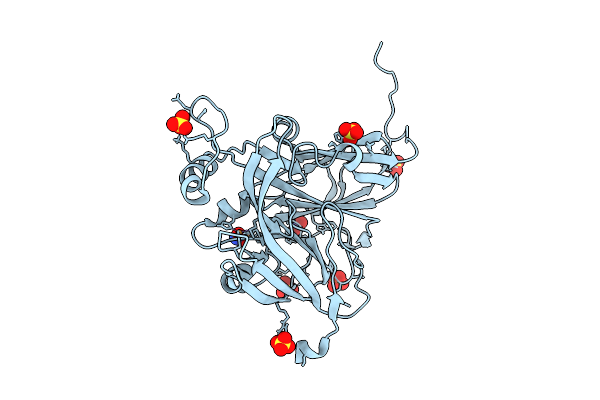
Sub-Atomic Resolution (0.95 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Achromobacter Cycloclastes Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox)
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:0.95 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
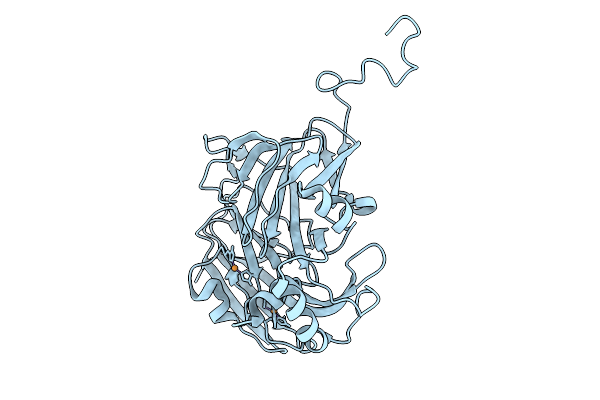
Sub-Atomic Resolution (0.95 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Achromobacter Cycloclastes Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:0.95 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, SO4, NO2 |
 |
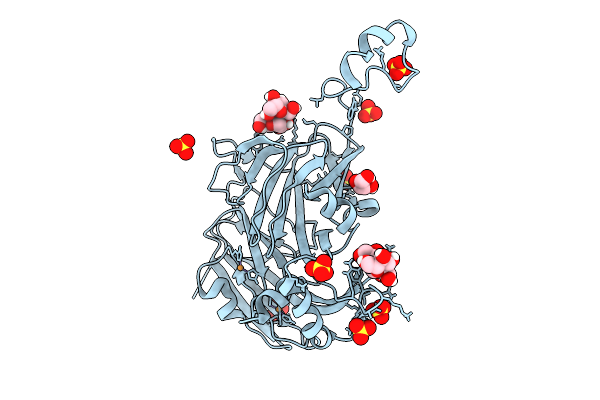
Atomic Resolution (1.05 A) Xfel Structure Of Chemically-Reduced Copper Nitrite Reductase From Bradyrhizobium Sp. Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.05 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Atomic Resolution (1.15 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Bradyrhizobium Sp. At Low Ph (5.5) Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Bradyrhizobium sp. ors 375
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, GLC, FRU, SO4, GOL |
 |
Low-Dose (14.9 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Higher-Dose (283.1 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
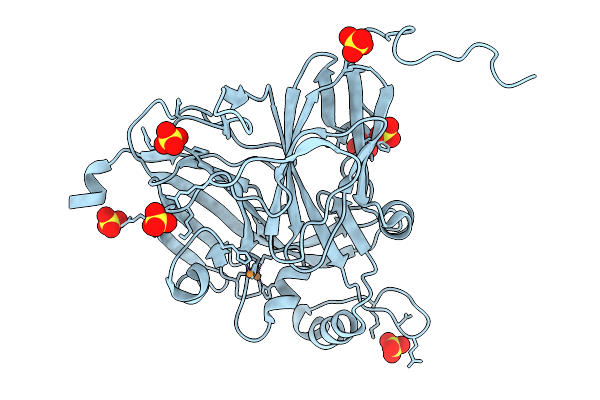
Low-Dose (33.3 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
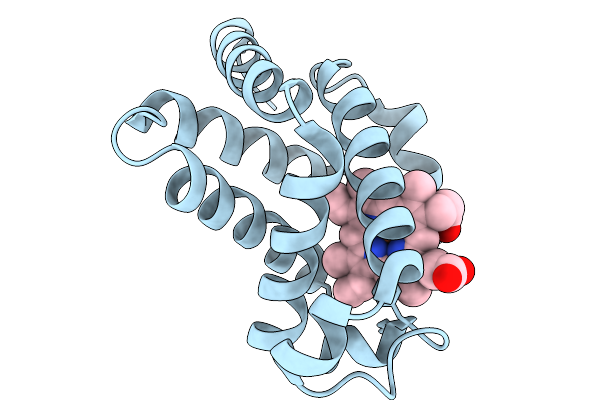
Higher-Dose (1332 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
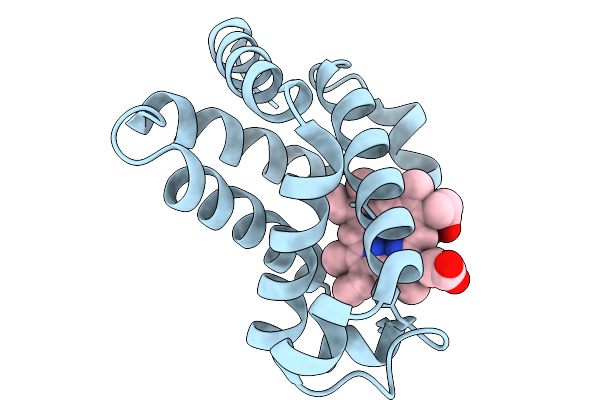
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM |
 |
Higher-Dose (260.0 Kgy) Structure Of Horse-Heart Myoglobin At Room Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Higher-Dose (676.8 Kgy) Structure Of Horse-Heart Myoglobin At Cryogenic Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM, MLA, NA |
 |
Low-Dose (14.4 Kgy) Structure Of Horse-Heart Myoglobin At Cryogenic Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM, MLI, NA |
 |
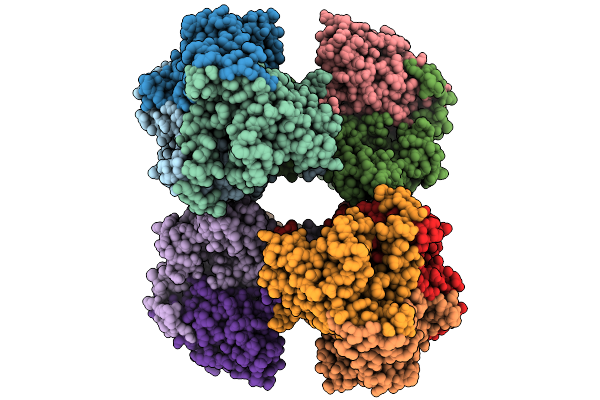
Structure Of Mycobacterium Tuberculosis Pyruvate Dehydrogenase Complex E2P Core Subunit Dlat In A Hexamer State
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-02-25 Classification: TRANSFERASE |
 |
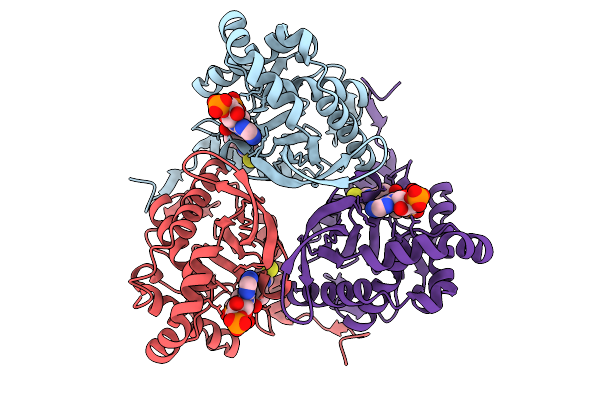
Structure Of Mycobacterium Tuberculosis Pyruvate Dehydrogenase Complex E2P Core Subunit Dlat In A Two-Hexamer State
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:2.51 Å Release Date: 2026-02-25 Classification: TRANSFERASE |
 |
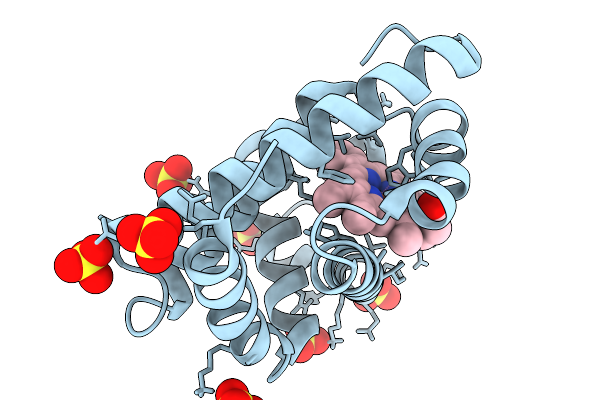
Structure Of Mycobacterium Tuberculosis Pyruvate Dehydrogenase Complex E2P Core Subunit Dlat Bound To Coenzyme A In A Hexamer State
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:4.20 Å Release Date: 2026-02-25 Classification: TRANSFERASE Ligands: COA |
 |
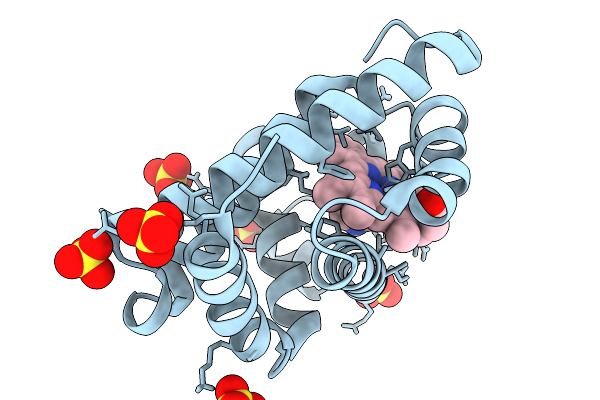
Organism: Physeter macrocephalus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-02-04 Classification: OXYGEN STORAGE Ligands: SO4, HEM |
 |
Organism: Physeter macrocephalus
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2026-02-04 Classification: OXYGEN STORAGE Ligands: IMD, SO4, HEM |
 |
Organism: Physeter macrocephalus
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-01-21 Classification: METAL BINDING PROTEIN Ligands: HEM |

