Search Count: 2,395
All
Selected
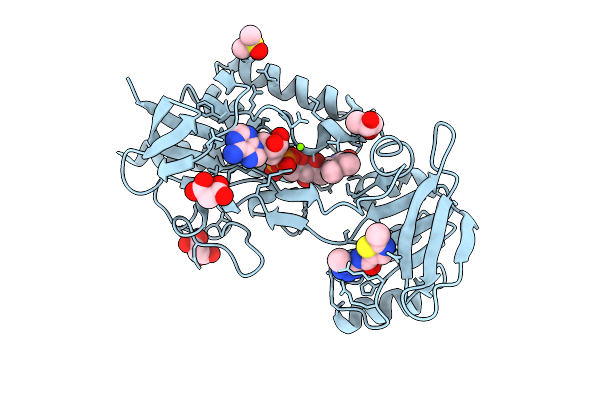 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2997505083
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1JAC, DMS |
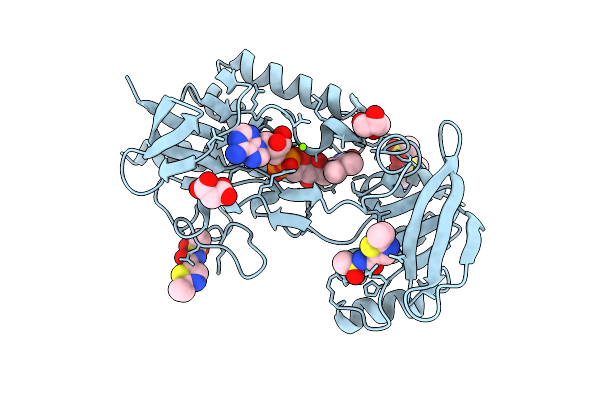 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z854627136
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I94 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z30008604
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I96 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z2858787682
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I97 |
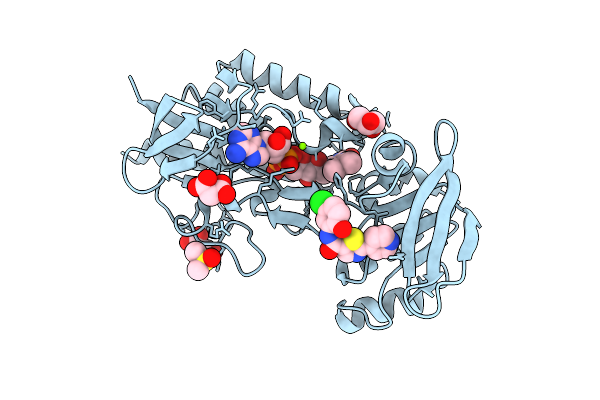 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z1280094148
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I95, DMS |
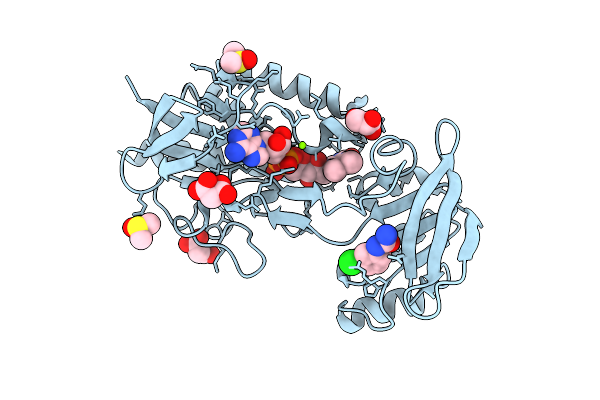 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z166687084
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, DMS, A1I98 |
 |
Crystal Structure Of Mycobacterium Smegmatis Thioredoxin Reductase In Complex With Z741560256
Organism: Mycolicibacterium smegmatis mc2 155
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, MLI, MG, A1I99 |
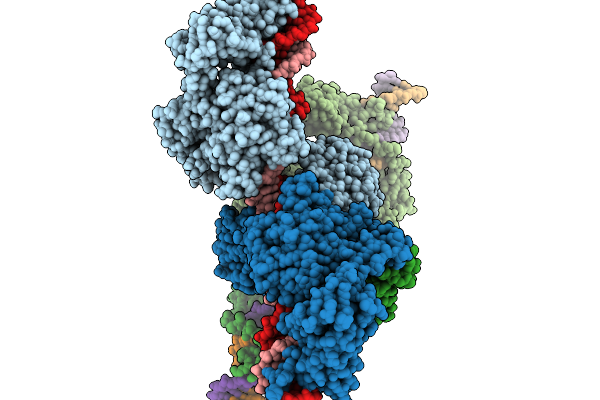 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Gt/Tt Cdn) In The Pre-Strand Exchange State
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
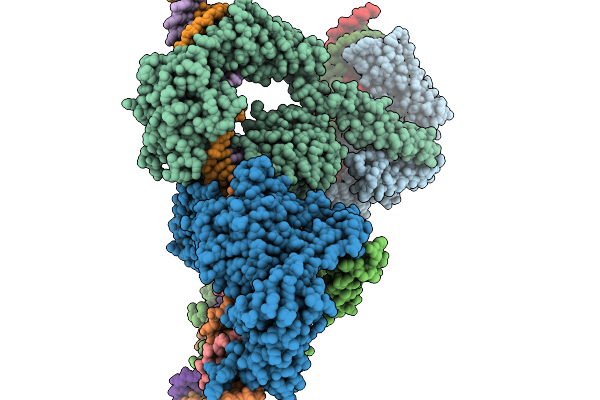 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Gt/Tt Cdn) In The Post-Strand Exchange State
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:3.97 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
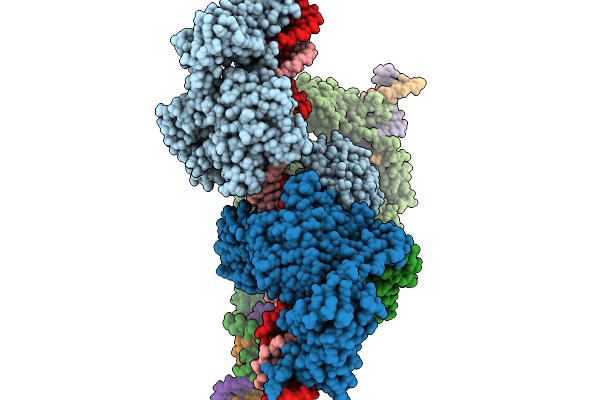 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Ca/Ca Cdn) In The Pre-Strand Exchange State
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:3.58 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
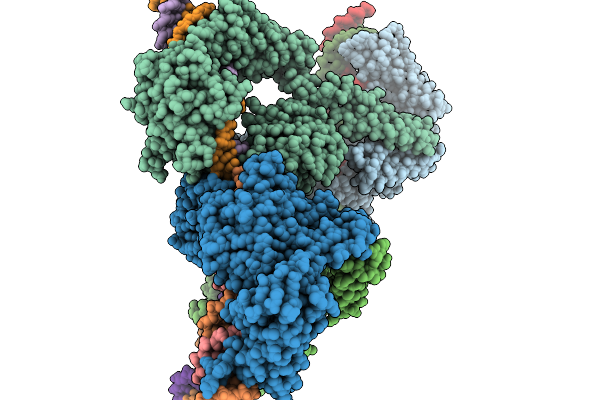 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Ca/Ca Cdn) In The Post-Strand Exchange State
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:3.63 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
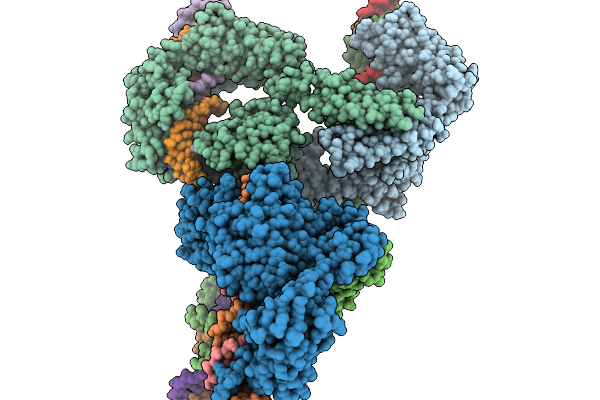 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Ca/Ca Cdn) In The Intermediate-Strand Exchange State 1
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:3.97 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
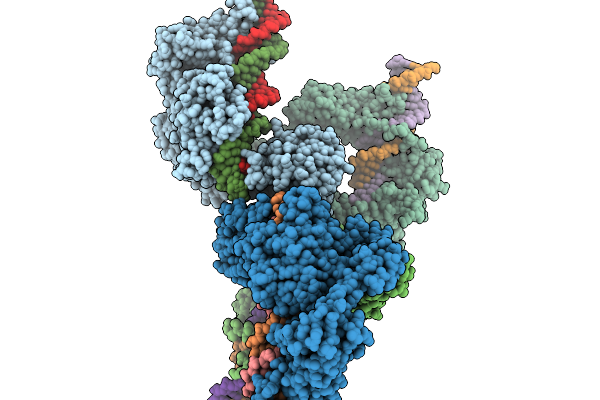 |
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Ca/Ca Cdn) In The Intermediate-Strand Exchange State 2
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Resolution:4.08 Å Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
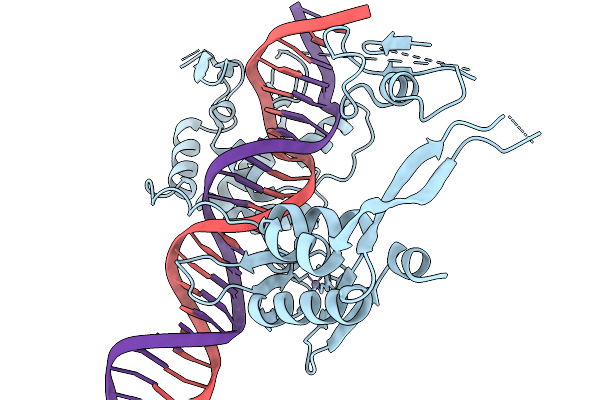
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Gt/Tt Cdn) In The Pre-Strand Exchange State (Attb-L)
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
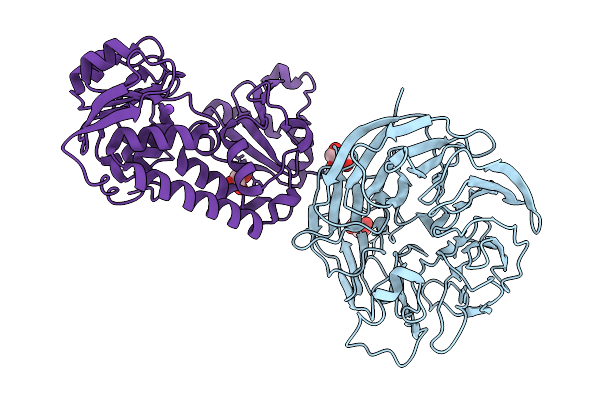
Cryo-Em Structure Of The Large Serine Recombinase Bxb1 In Complex With Attp And Attb (Gt/Tt Cdn) In The Pre-Strand Exchange State (Attb-R)
Organism: Mycobacterium phage bxb1, Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Structure Of M. Thermoresistible Rv3035 In Complex With M. Tuberculosis Fecb
Organism: Mycolicibacterium thermoresistibile, Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:3.35 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: GOL, NA |
 |
Organism: Mycolicibacterium thermoresistibile
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: GOL, SO4 |
 |
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: ELECTRON TRANSPORT Ligands: CDL, HDD, HEB, MQ9 |
 |
Structure Of The Lactate Monooxygenase Mutant Y44N, Y152F, H290S, T181A, F184P
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: FMN, GOL, CIT, SO4 |
 |
Organism: Mycolicibacterium fortuitum
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-02-18 Classification: FLAVOPROTEIN |

