Search Count: 480
All
Selected
 |
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2025-12-24 Classification: METAL BINDING PROTEIN Ligands: CU, SO4, NA |
 |
Crystal Structure Of Gnat Superfamily Acetyltransferase Pa2271 From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2025-05-28 Classification: TRANSFERASE Ligands: ACO, NA, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-11-09 Classification: TRANSPORT PROTEIN Ligands: CO, MYR |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2022-11-09 Classification: TRANSPORT PROTEIN Ligands: CO, MYR |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2022-11-09 Classification: TRANSPORT PROTEIN Ligands: CO, MYR |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: EDO |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: SO4 |
 |
Beta-Lactoglobulin Mutant Faw (I56F/L39A/M107W) In Complex With Desipramine (Faw-Dsm#3)
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: DSM, EDO, SO4, CL |
 |
Beta-Lactoglobulin Mutant Faf (I56F/L39A/M107F) In Complex With Desipramine (Faf-Dsm)
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: DSM, GOL, CL, EDO, SO4 |
 |
Beta-Lactoglobulin Mutant Faw (I56F/L39A/M107W) In Complex With Desipramine (Faw-Dsm#1)
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: SO4, DSM, CL, EDO |
 |
Beta-Lactoglobulin Mutant Faw (I56F/L39A/M107W) In Complex With Desipramine (Faw-Dsm#2)
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2022-05-11 Classification: TRANSPORT PROTEIN Ligands: EDO, DSM, SO4, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2022-01-12 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2022-01-12 Classification: CELL CYCLE Ligands: ZN |
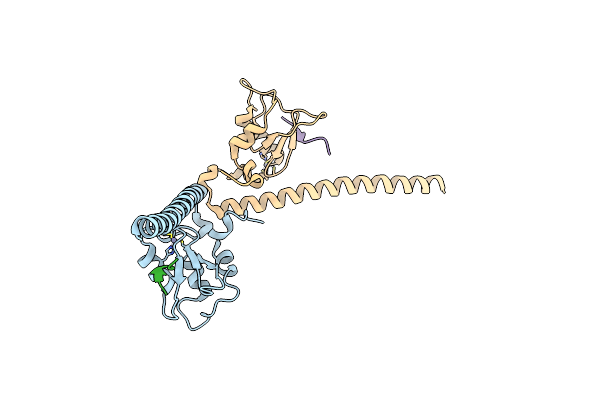 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2022-01-12 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2022-01-12 Classification: CELL CYCLE Ligands: ZN |
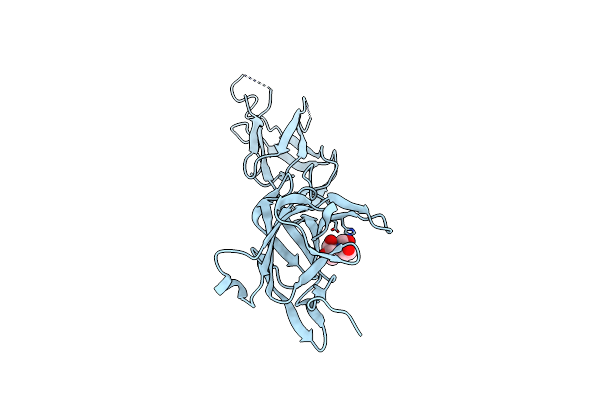 |
Crystal Structure Of Srtb-Anchored Collagen-Binding Adhesin Fragment (Residues 206-565) From Clostridioides Difficile Strain 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-10-20 Classification: CELL ADHESION Ligands: BTB |
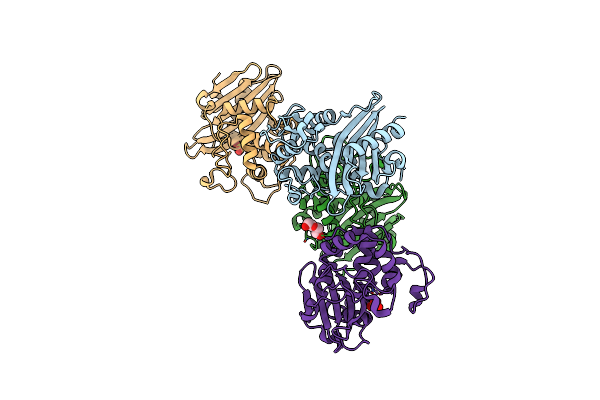 |
Crystal Structure Of C79A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: PEG, SO4 |
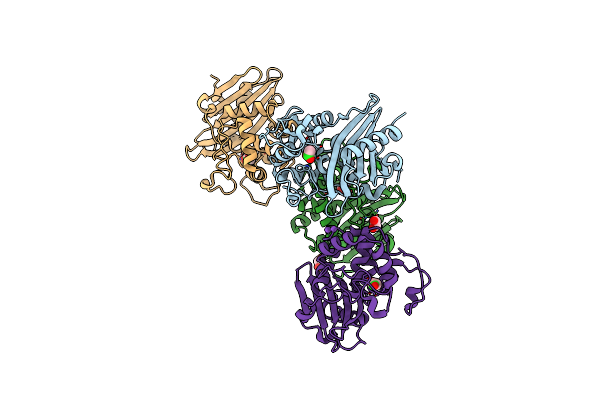 |
Crystal Structure Of K83A Mutant Of Class D Beta-Lactamase From Clostridium Difficile 630
Organism: Clostridioides difficile
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-08-11 Classification: HYDROLASE Ligands: CL, ACT, EDO, NA |
 |
Crystal Structure Of 3-Ketosteroid Delta1-Dehydrogenase From Sterolibacterium Denitrificans In Complex With 1,4-Androstadiene-3,17-Dione
Organism: Sterolibacterium denitrificans
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2021-07-21 Classification: OXIDOREDUCTASE Ligands: FAD, ANB, PEG, GOL, NA |
 |
Crystal Structure Of Truncated (Act Domain Removed) Prephenate Dehydrogenase Tyra From Bacillus Anthracis In Complex With Nad
Organism: Bacillus anthracis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-05-19 Classification: HYDROLASE Ligands: NAD, GOL, CL |

