Search Count: 33
All
Selected
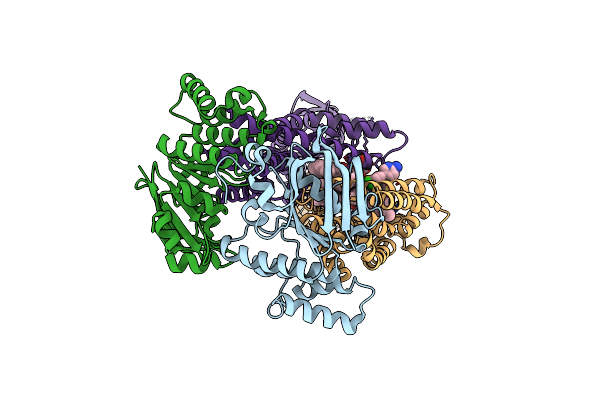 |
Acinetobacter Baylyi Lptb2Fg Bound To Lipopolysaccharide And A Macrocyclic Peptide
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: Y75, JSG, LMT |
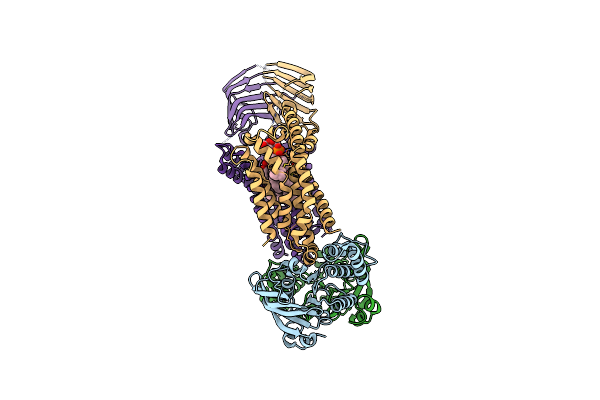 |
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: JSG |
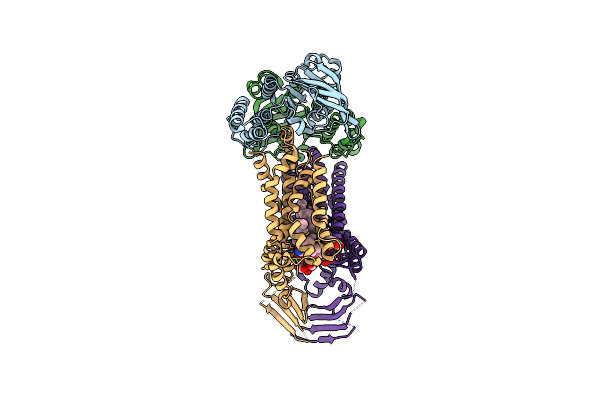 |
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: JSG, VB6, LMT |
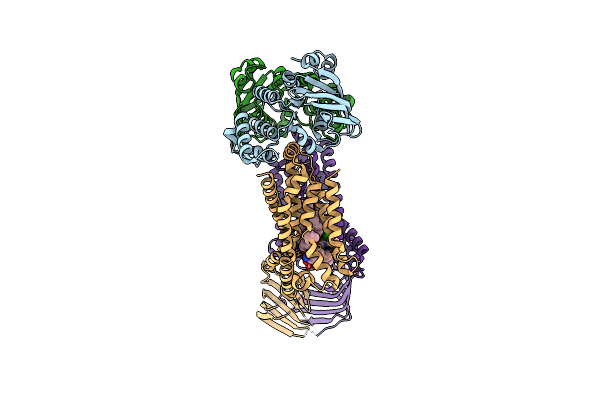 |
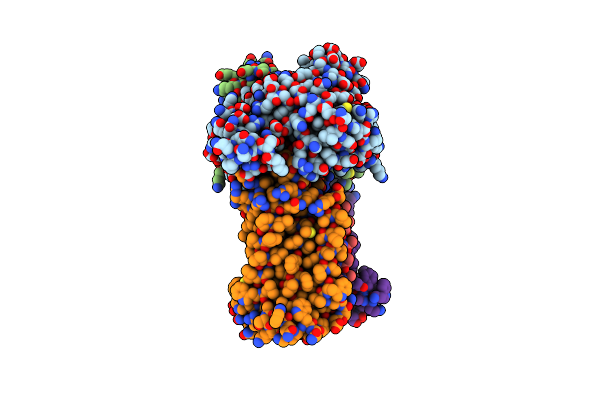
Acinetobacter Baylyi Lptb2Fg Bound To Lipopolysaccharide And A Macrocyclic Peptide
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: JSG, MG3 |
 |
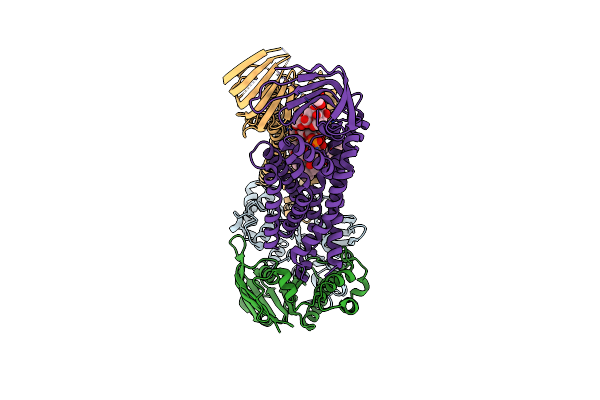
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT |
 |
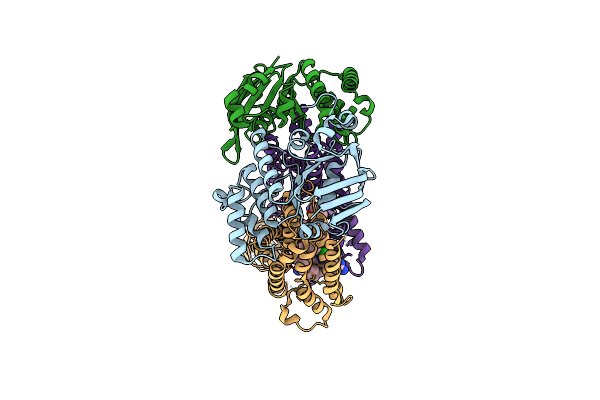
Acinetobacter Baylyi Lptb2Fg Bound To Acinetobacter Baylyi Lipopolysaccharide
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: LMT, WJR |
 |
Acinetobacter Baylyi Lptb2Fg Bound To Acinetobacter Baylyi Lipopolysaccharide And A Macrocyclic Peptide
Organism: Acinetobacter baylyi adp1
Method: ELECTRON MICROSCOPY Release Date: 2024-01-03 Classification: LIPID TRANSPORT Ligands: WJW, Y75 |
 |
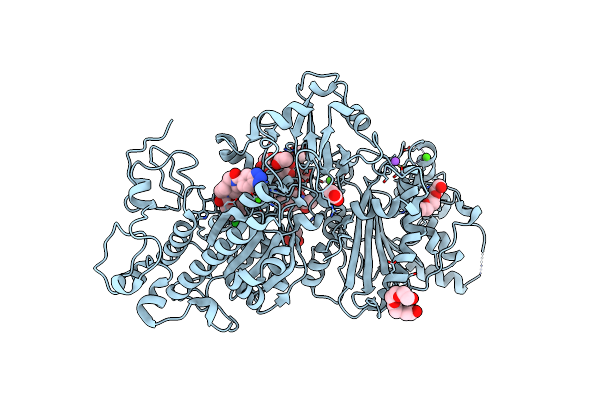
Autotaxin, 4-[3-Oxo-3-(2-Oxo-2,3-Dihydro-Benzooxazol-6-Yl)-Propyl]-Piperazine-1-Carboxylic Acid 3,5-Dichloro-Benzyl Ester, 1.90A, P212121, Rfree=19.1%
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2022-04-13 Classification: HYDROLASE Ligands: CA, K, ACT, 6ZO, ZN, NA |
 |
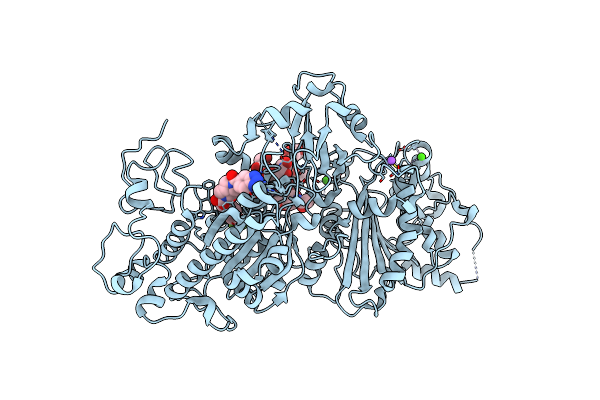
Autotaxin, (3,5-Dichlorophenyl)Methyl (3As,8Ar)-2-(1H-Benzotriazole-5-Carbonyl)-1,3,3A,4,5,7,8,8A-Octahydropyrrolo[3,4-D]Azepine-6-Carboxylate, 1.80A, P212121, Rfree=21.1%
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-04-13 Classification: HYDROLASE Ligands: YV1, ACT, PG4, ZN, CA, CL, NA |
 |
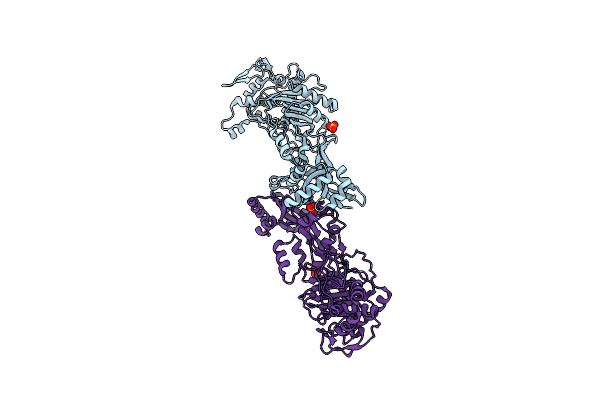
Autotaxin, [4-(Trifluoromethoxy)Phenyl]Methyl (3As,6As)-2-(1H-Benzotriazole-5-Carbonyl)-1,3,3A,4,6,6A-Hexahydropyrrolo[3,4-C]Pyrrole-5-Carboxylate, 1.80A, P212121, Rfree=23.3%
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2022-04-13 Classification: HYDROLASE Ligands: YVS, ZN, CA, CL, NA |
 |
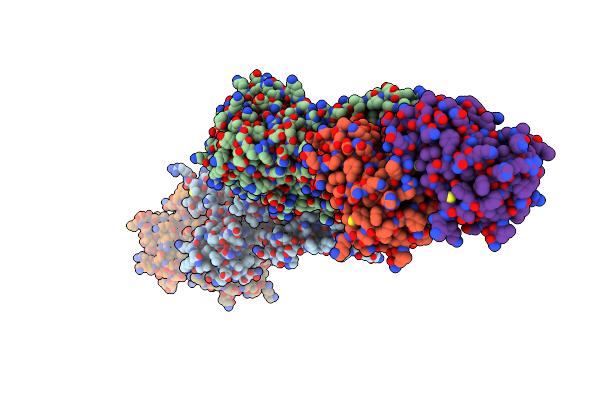
Organism: Helicobacter pylori (strain atcc 700392 / 26695)
Method: X-RAY DIFFRACTION Resolution:3.03 Å Release Date: 2017-08-23 Classification: HYDROLASE/ANTIBIOTIC Ligands: SO4 |
 |
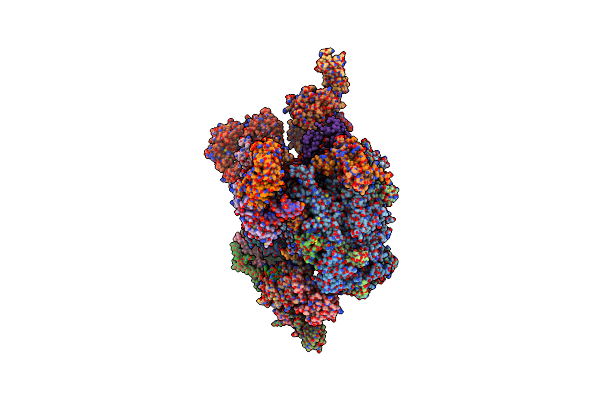
Complex Between Penicillin-Binding Protein (Pbp2) And Mrec From Helicobacter Pylori
Organism: Helicobacter pylori (strain atcc 700392 / 26695)
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2017-08-23 Classification: HYDROLASE/ANTIBIOTIC |
 |
Method: ELECTRON MICROSCOPY
Release Date: 2016-09-28 Classification: TRANSCRIPTION, TRANSFERASE/DNA Ligands: ZN, MG |
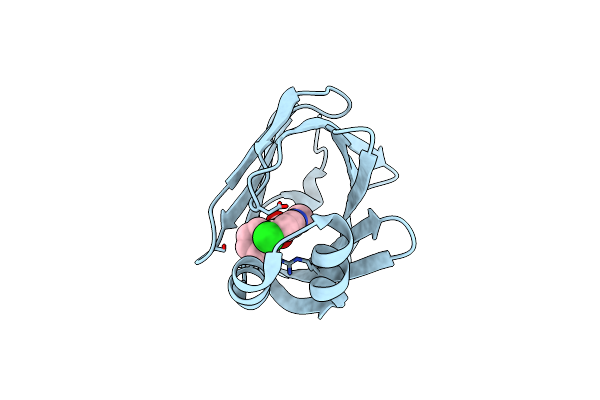 |
Human Fatty Acid Binding Protein 4 In Complex With 6-Chloro-2-Methyl-4-Phenyl-Quinoline-3-Carboxylic Acid At 1.18A
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.18 Å Release Date: 2016-03-09 Classification: LIPID BINDING PROTEIN Ligands: 5M8 |
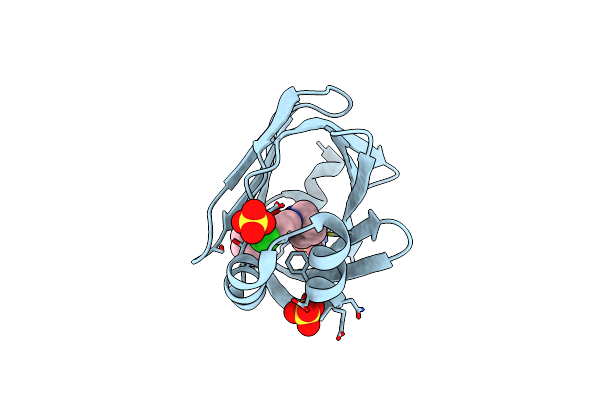 |
Human Fabp4 In Complex With 6-Chloro-4-Phenyl-2-Piperidin-1-Yl-Quinoline-3-Carboxylic Acid At 1.29A
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.29 Å Release Date: 2016-03-09 Classification: LIPID BINDING PROTEIN Ligands: 5M7, SO4 |
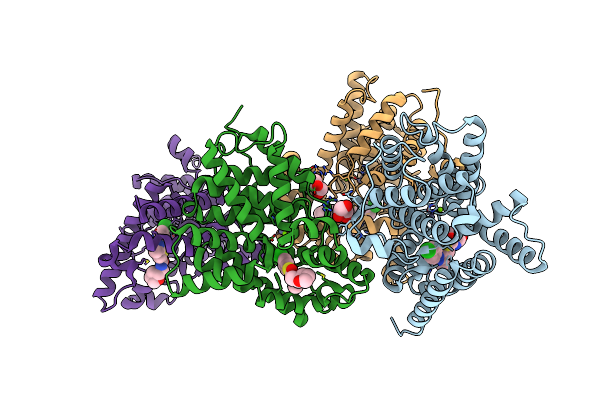 |
Crystal Structure Of Human Phosphodiesterase 10 In Complex With C13C(Cc(S1)C(Ncc2Occc2)=O)C(Nn3C4Ccc(Cc4)Cl)C, Micromolar Ic50=0.217
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2016-03-09 Classification: HYDROLASE Ligands: ZN, MG, GOL, 5M6 |
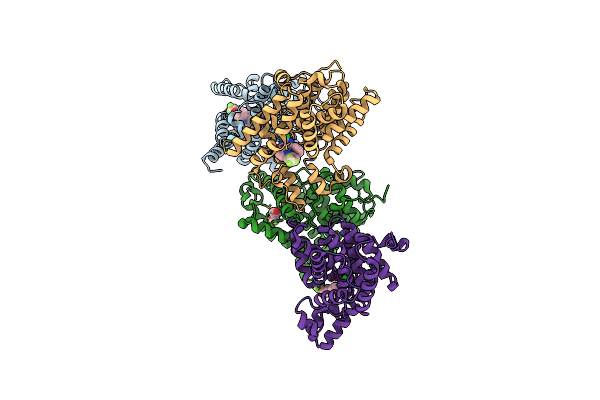 |
Crystal Structure Of Human Phosphodiesterase 10 In Complex With C1(C(Nc([Nh]1)Cl)C2Ccccc2)C4=Nn(C3Cccc(C3)Oc(F)(F)F)C=Cc4=O, Micromolar Ic50=0.029618
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2016-03-09 Classification: HYDROLASE Ligands: ZN, MG, 5MG |
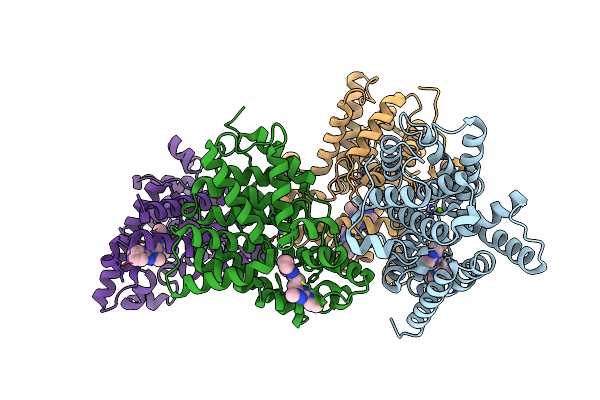 |
Crystal Structure Of Human Phosphodiesterase 10 In Complex With N4C(C)N1C(Nc(N1)Ccc2Nc(Nn2C)N3Cccc3)C(C4)Cc, Micromolar Ic50=0.0037753
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2016-03-09 Classification: HYDROLASE Ligands: ZN, MG, 5MF |
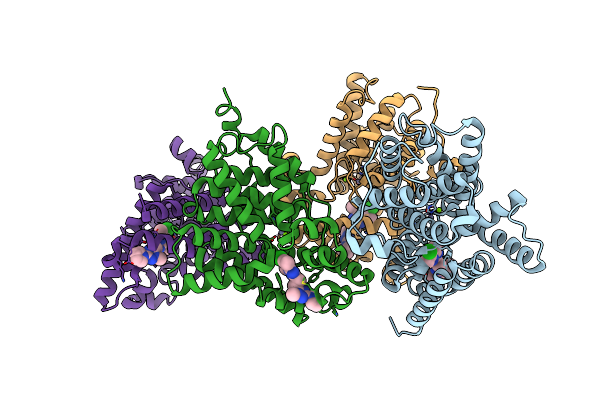 |
Crystal Structure Of Human Phosphodiesterase 10 In Complex With C2(C(N1Nc(Nc1C(C2)C)Ccc3Nc(Nn3C)N4Cccc4)C)Cl, Micromolar Ic50=0.000279
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2016-03-09 Classification: HYDROLASE Ligands: ZN, MG, 5M9 |
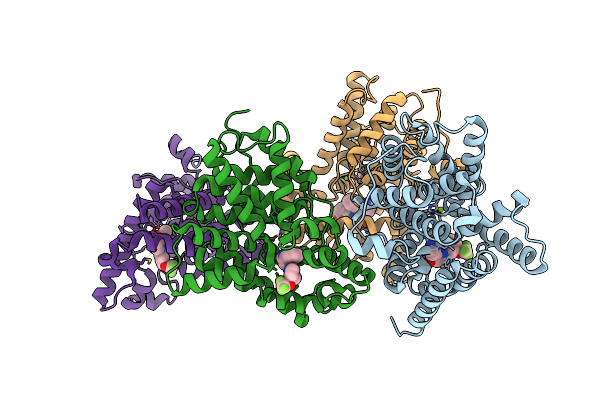 |
Crystal Structure Of Human Phosphodiesterase 10 In Complex With C2(=Nn(C1Cccc(C1)Oc(F)(F)F)C=Cc2=O)C3Ccnn3C4Ccccc4, Micromolar Ic50=0.019462
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2016-03-09 Classification: HYDROLASE Ligands: ZN, MG, 67A |

