Search Count: 25
All
Selected
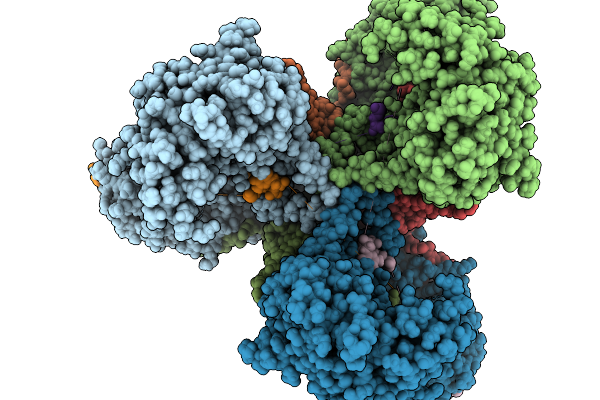 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-02-18 Classification: ANTIVIRAL PROTEIN Ligands: MG |
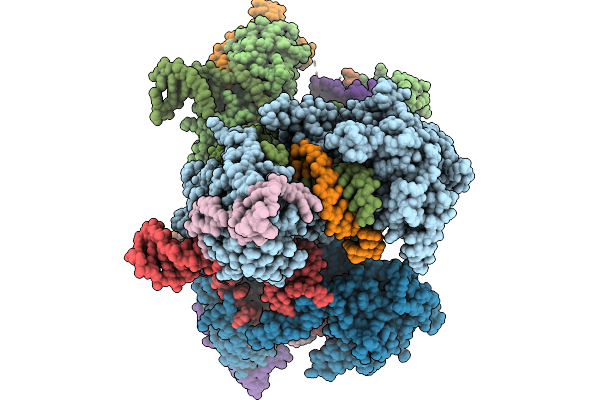 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: ANTIVIRAL PROTEIN |
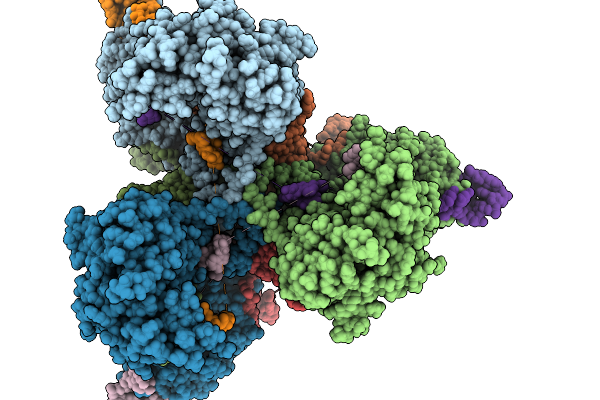 |
Organism: Escherichia coli nctc 86
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-04 Classification: ANTIVIRAL PROTEIN Ligands: MG |
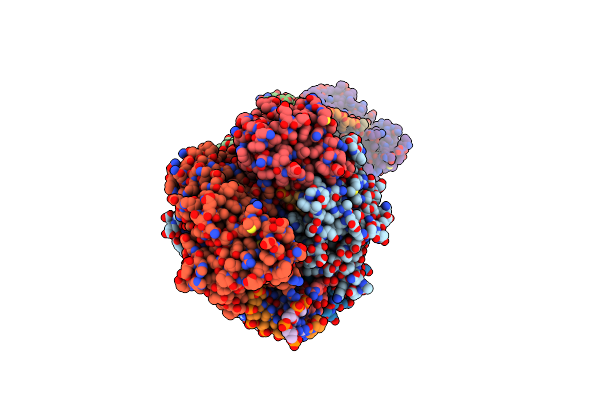 |
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: GENE REGULATION Ligands: ZN |
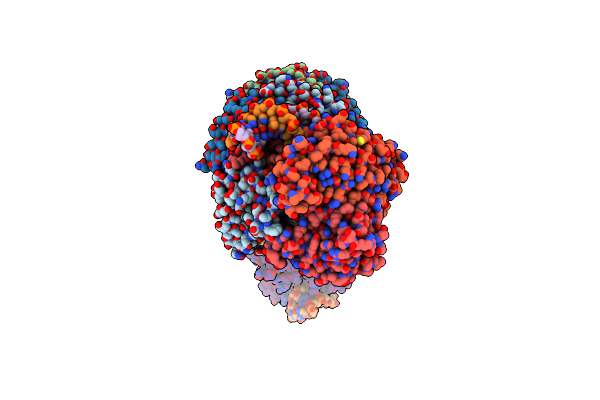 |
Organism: Pseudomonas oleovorans
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: GENE REGULATION Ligands: ZN |
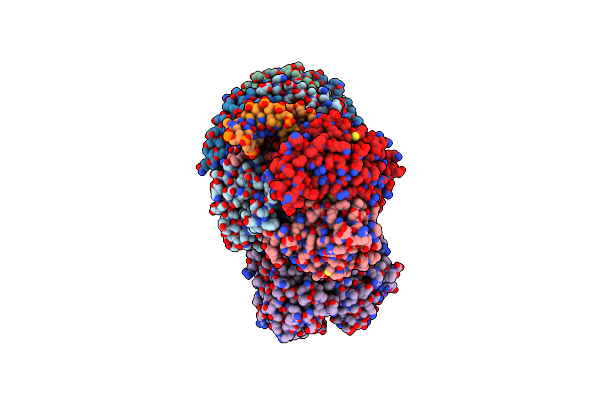 |
Dna Bound Type Iv-A1 Crispr Effector Complex With The Ding Helicase From P. Oleovorans
Organism: Pseudomonas oleovorans
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: GENE REGULATION Ligands: ZN |
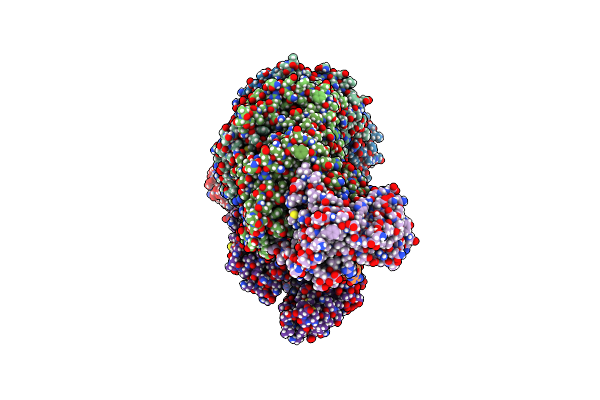 |
Dna-Bound Type Iv-A3 Crispr Effector In Complex With Ding Helicase From K. Pneumoniae (State I)
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ANTIVIRAL PROTEIN Ligands: ZN |
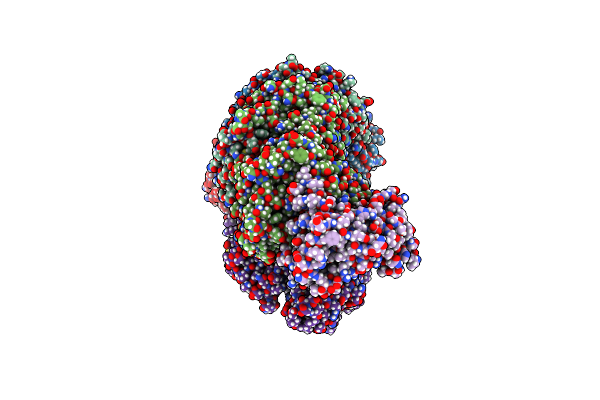 |
Dna-Bound Type Iv-A3 Crispr Effector In Complex With Ding Helicase From K. Pneumoniae (State Ii)
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ANTIVIRAL PROTEIN Ligands: ZN |
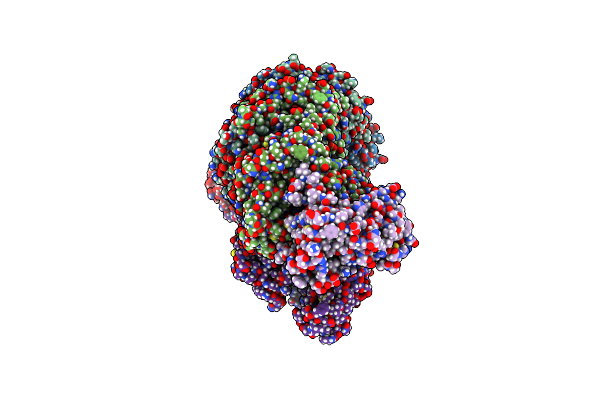 |
Dna-Bound Type Iv-A3 Crispr Effector In Complex With Ding Helicase From K. Pneumoniae (State Iii)
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Release Date: 2024-11-06 Classification: ANTIVIRAL PROTEIN Ligands: ZN |
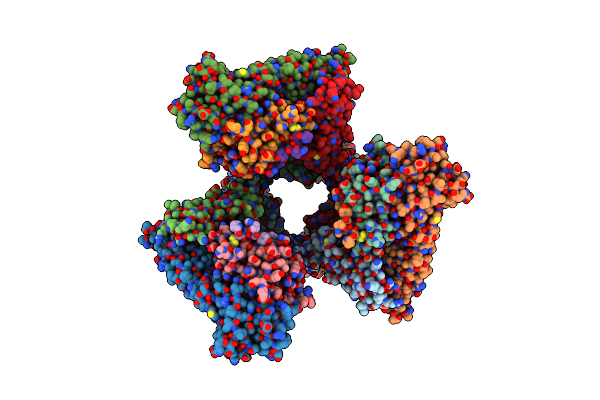 |
Cryo-Em Structure Of The Ternary Complex Between Netrin-1, Neogenin And Repulsive Guidance Molecule B
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2021-03-31 Classification: SIGNALING PROTEIN Ligands: CA, NAG |
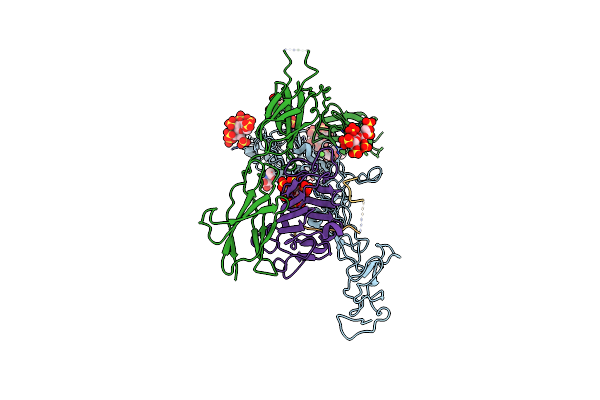 |
Structure Of The Ternary Complex Between Netrin-1, Repulsive-Guidance Molecule-B (Rgmb) And Neogenin
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.25 Å Release Date: 2021-03-31 Classification: SIGNALING PROTEIN Ligands: NAG, CA, SO4 |
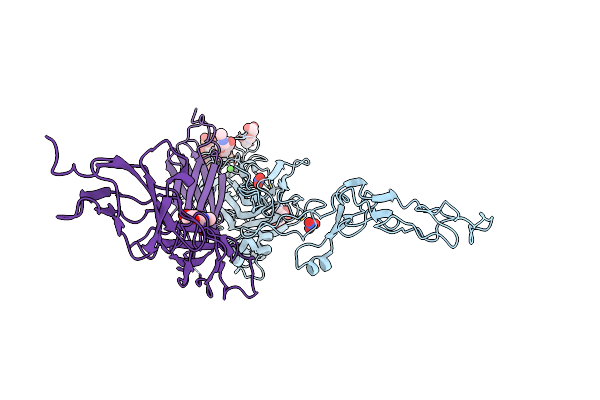 |
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2021-03-31 Classification: SIGNALING PROTEIN Ligands: NAG, CA, NO3 |
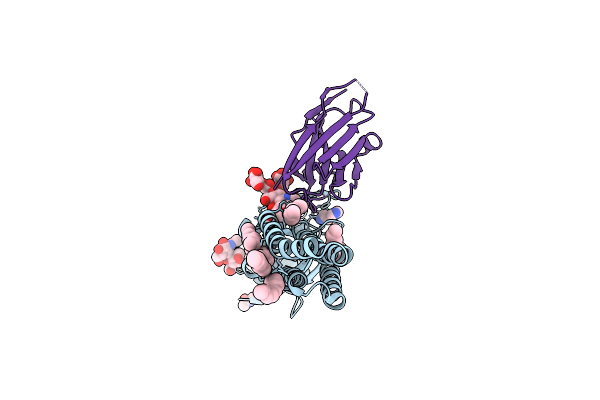 |
Cryoem Structure Of A Human Gamma-Aminobutyric Acid Receptor, The Gaba(A)R-Beta3 Homopentamer, In Complex With Histamine And Megabody Mb25 In Lipid Nanodisc
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Resolution:1.70 Å Release Date: 2020-11-18 Classification: MEMBRANE PROTEIN Ligands: R16, D10, NAG, OCT, HSM, CL |
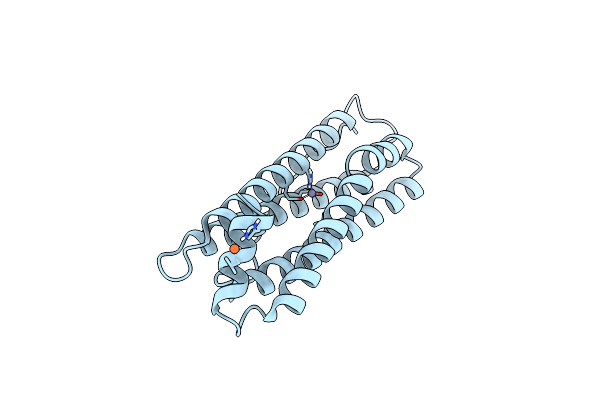 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:1.22 Å Release Date: 2020-10-28 Classification: METAL BINDING PROTEIN Ligands: FE, ZN |
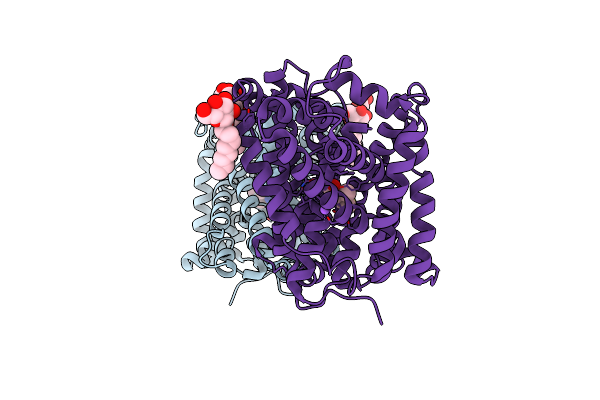 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN Ligands: BOG, NA, ILE |
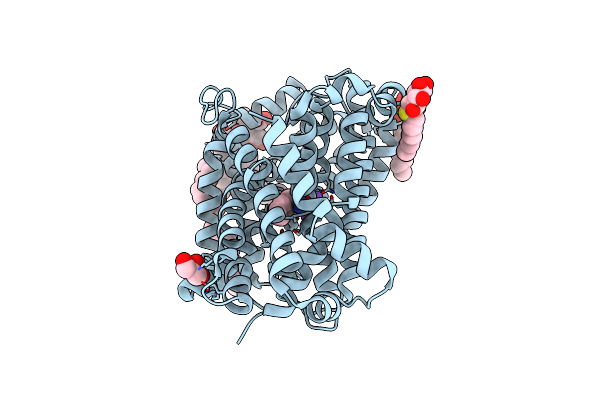 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN |
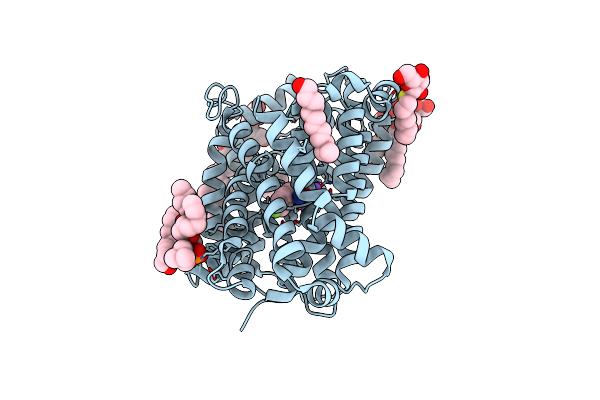 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN |
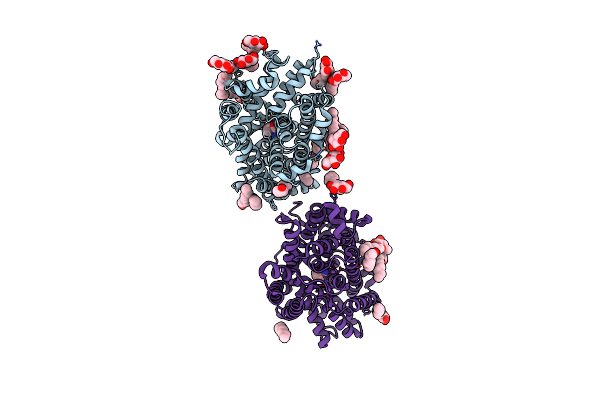 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN Ligands: VAL, NA, BNG, LMT |
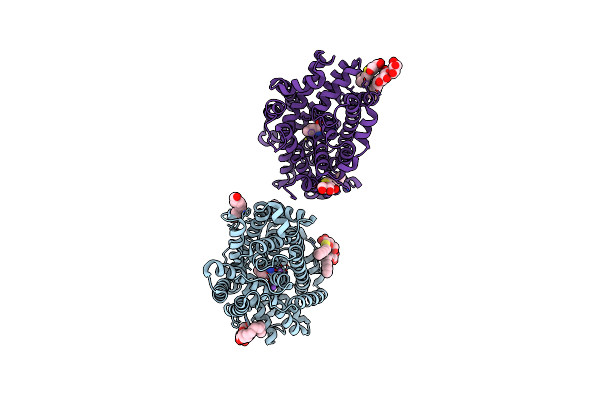 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN |
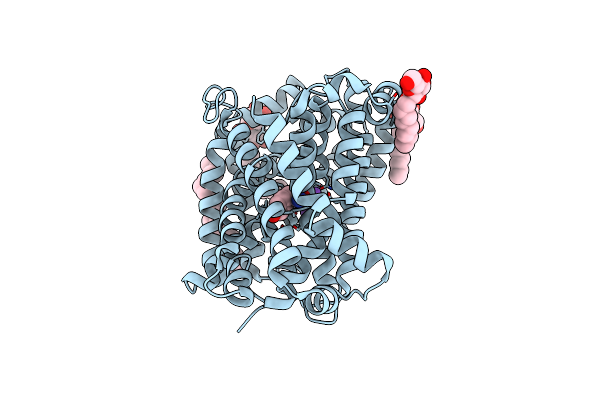 |
Organism: Bacillus halodurans
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2020-07-15 Classification: TRANSPORT PROTEIN Ligands: LMT, NA, TYR, BNG |

