Search Count: 64
All
Selected
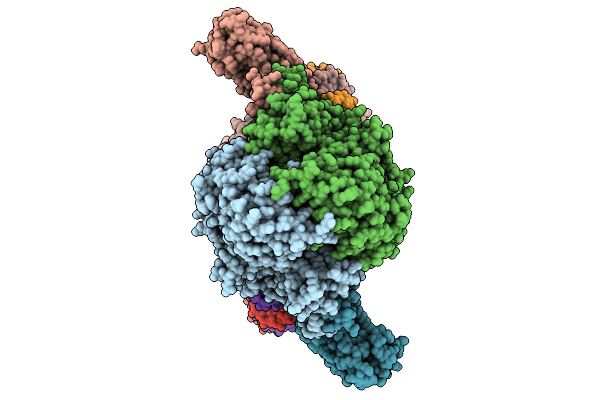 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Cleavage State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: CA, ANP, K |
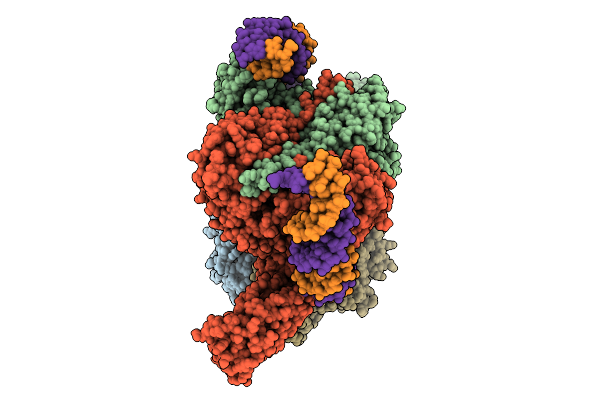 |
Cryoem Structure Of M. Mazei Topoisomerase Vi(A-E342Q)-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, CA, K |
 |
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
 |
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Partially Unfolded Transducer State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: MG, ANP, K |
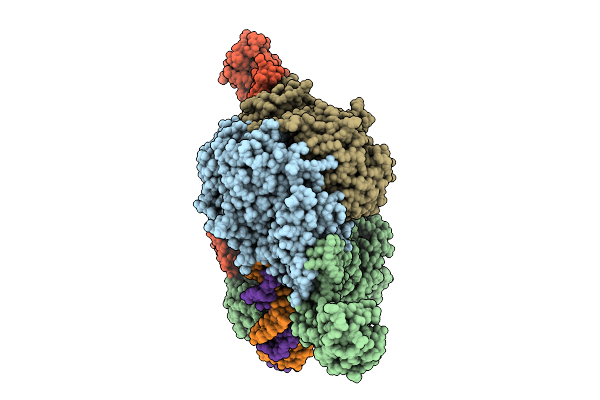 |
Cryoem Structure Of M. Mazei Topoisomerase Vi-Minicircle Dna Complex In Asymmetric State
Organism: Methanosarcina mazei go1, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: ISOMERASE/DNA Ligands: ANP, MG, K |
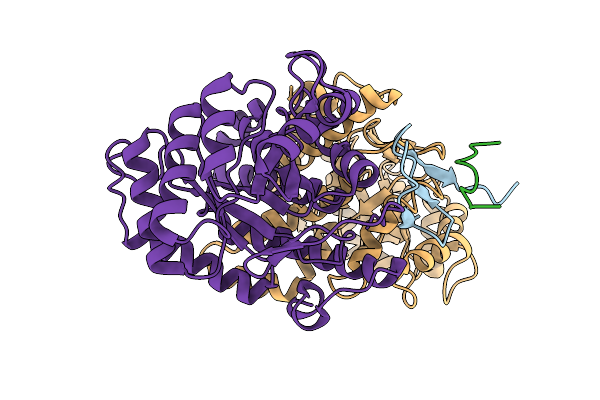 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN |
 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: A1JAP, L1P, A1JAO, NA, A1JAN |
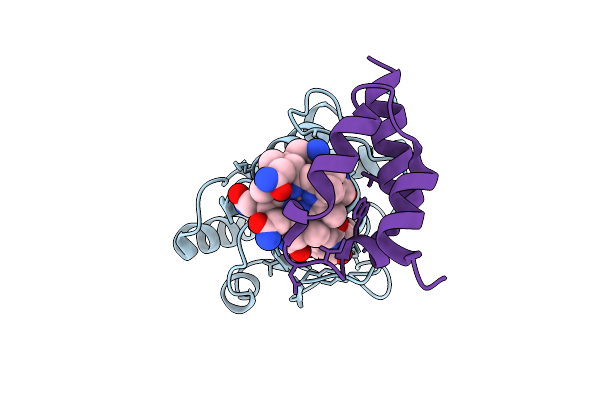 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13 |
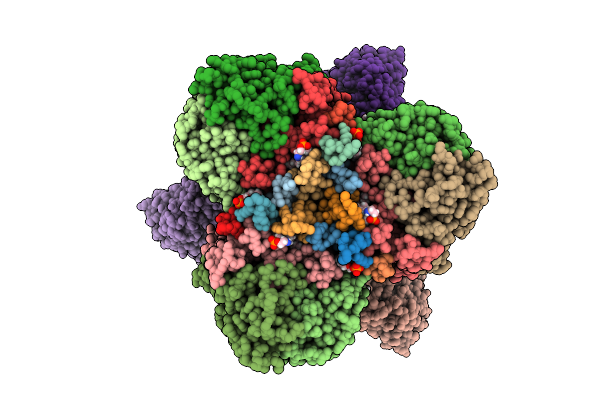 |
Structure Of The Energy Converting Methyltransferase (Mtr) Of Methanosarcina Mazei In Complex With A Novel Protein Binder
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: MEMBRANE PROTEIN Ligands: B13, A1JAP, L1P, A1JAO, NA, A1JAN |
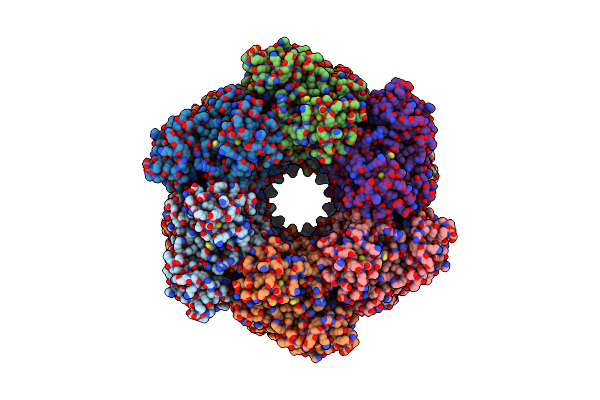 |
Cryo-Em Structure Of The Active Dodecameric Methanosarcina Mazei Glutamine Synthetase.
Organism: Methanosarcina mazei go1
Method: ELECTRON MICROSCOPY Release Date: 2025-01-08 Classification: CYTOSOLIC PROTEIN Ligands: AKG |
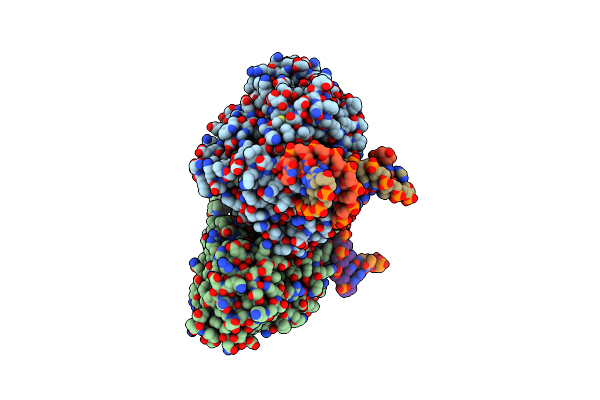 |
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (3 Picosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (300 Ps Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (1 Nanosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (3 Nanosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-11-22 Classification: LYASE Ligands: FDA, SO4 |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (10 Nanosecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2023-11-22 Classification: LYASE Ligands: FDA, SO4 |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (30 Nanosecond Timepoint)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (1 Microsecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (10 Microsecond Pump Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |
 |
Time-Resolved Sfx Structure Of The Class Ii Photolyase Complexed With A Thymine Dimer (30 Microsecond Pump-Probe Delay)
Organism: Methanosarcina mazei go1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-11-22 Classification: LYASE Ligands: SO4, FDA |

