Search Count: 259
All
Selected
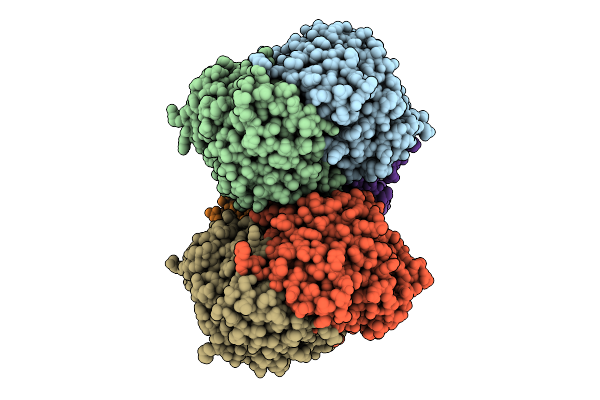 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
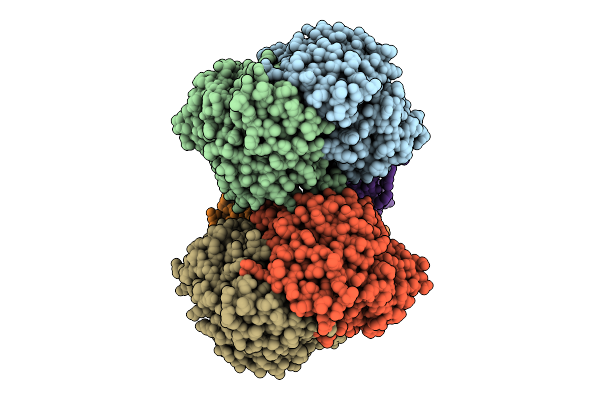 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
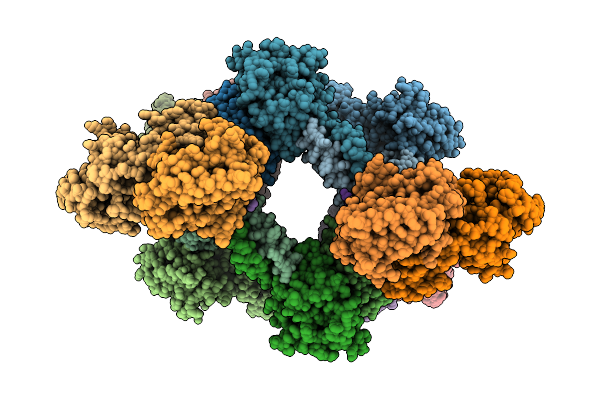 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
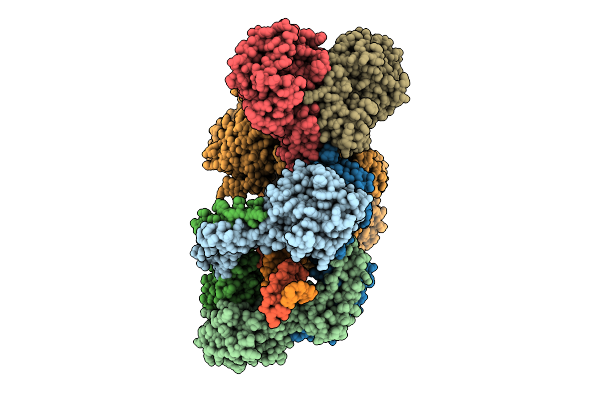 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: MG, ZN |
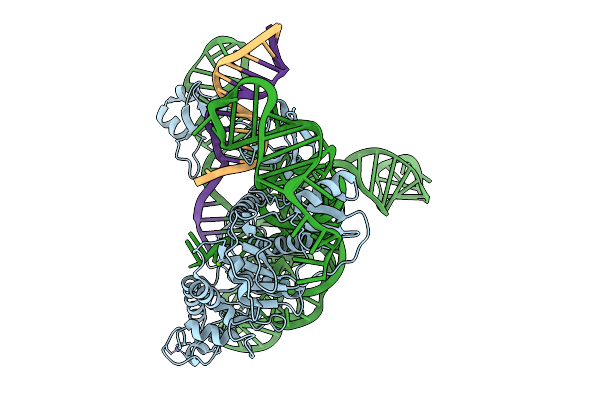 |
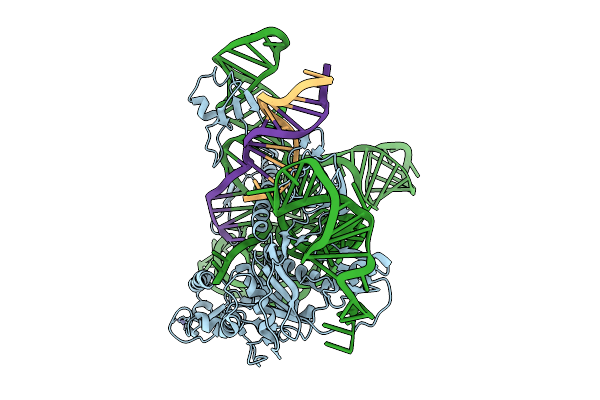
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
 |
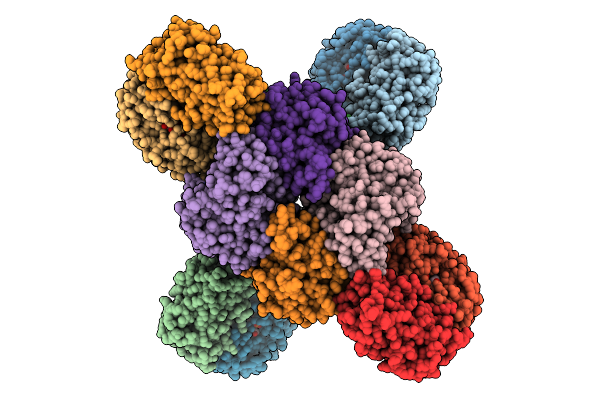
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.09 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: ZN, MG |
 |
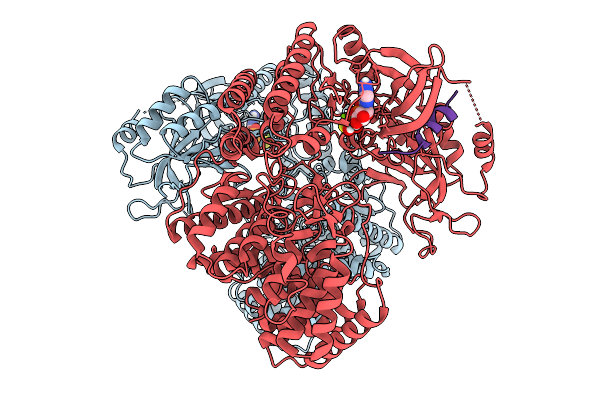
Cryo-Em Helical Structure Of The Ditp-Kombc(H146N) Complex With Nad Fragments
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-01-21 Classification: IMMUNE SYSTEM Ligands: Y43, AMP |
 |
Cryo-Em Structure Of Classiii Salivaricin Modification Enzyme Salkc In The Presence Of Sala
Organism: Streptococcus salivarius
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2025-09-03 Classification: ANTIMICROBIAL PROTEIN Ligands: AGS, MG |
 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-07-23 Classification: CELL CYCLE Ligands: AGS |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-06-18 Classification: HYDROLASE Ligands: ZN |
 |
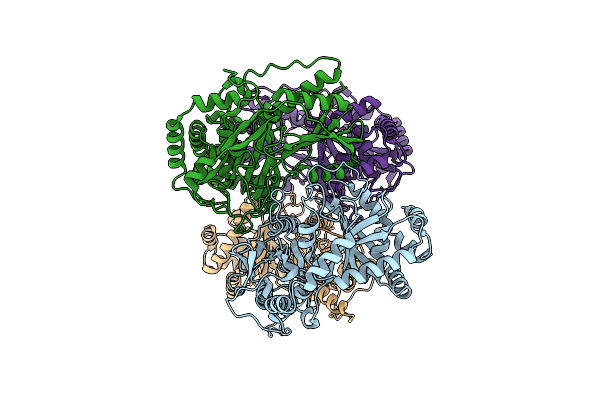
Organism: Streptomyces
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Streptomyces
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: BIOSYNTHETIC PROTEIN |
 |
Organism: Streptomyces
Method: ELECTRON MICROSCOPY Release Date: 2025-03-19 Classification: BIOSYNTHETIC PROTEIN |
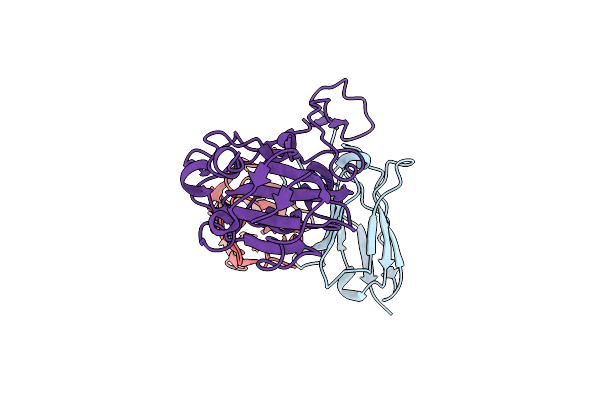 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: VIRAL PROTEIN |
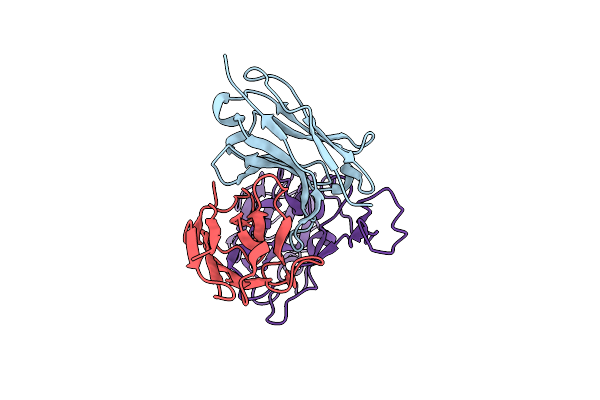 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2024-10-09 Classification: VIRAL PROTEIN |
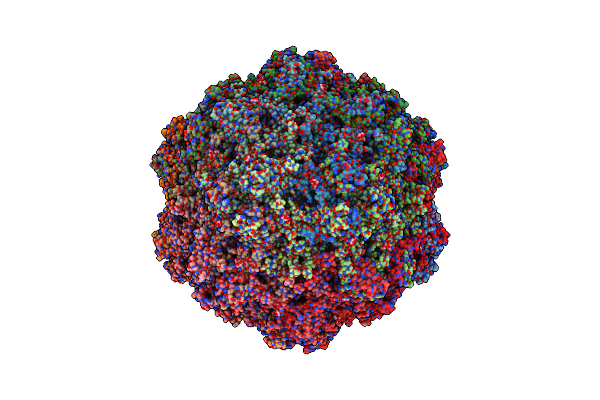 |
Organism: Bufavirus-1
Method: ELECTRON MICROSCOPY Release Date: 2024-09-18 Classification: VIRUS LIKE PARTICLE Ligands: SIA |

