Search Count: 58
All
Selected
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-04-08 Classification: PROTEIN TRANSPORT |
 |
Organism: Shewanella baltica
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-02-25 Classification: LIPID BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: A1JLE, SO4, PG4, PGE, PEG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: PEG, A1JLD, DMS |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: A1JLB, PEG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-12-10 Classification: OXIDOREDUCTASE Ligands: PEG, A1JLA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-01 Classification: OXIDOREDUCTASE Ligands: PG4, PGE, DMS, A1IEE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-10-01 Classification: OXIDOREDUCTASE Ligands: PGE, A1JGL |
 |
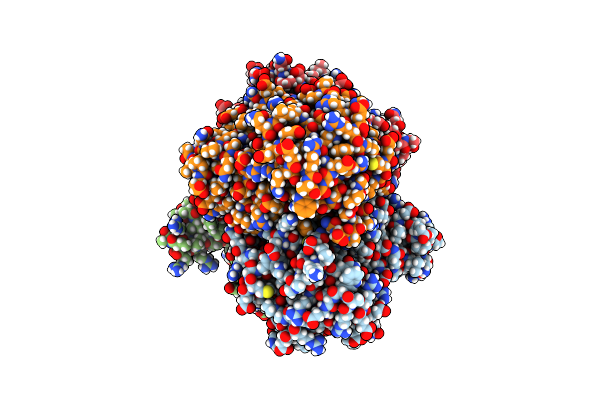
Heme-Dependent L-Tyrosine Hydroxylase (Tyrh) From Streptomyces Sclerotialus: Fourfold Mutant
Organism: Streptomyces sclerotialus
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-12-18 Classification: OXIDOREDUCTASE Ligands: HEM, IND, PEG, ACT |
 |
Organism: Thermus caliditerrae
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2024-06-26 Classification: OXIDOREDUCTASE Ligands: MG |
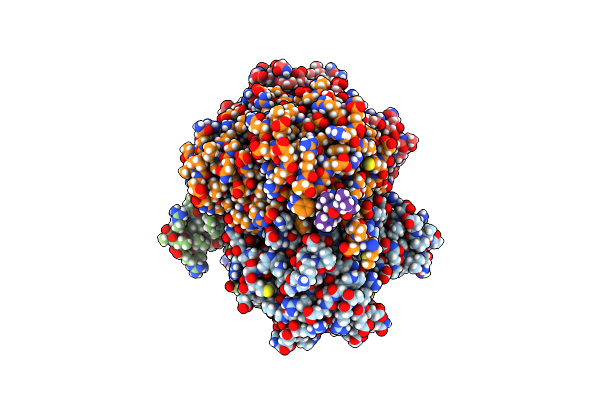 |
Short-Chain Dehydrogenase/Reductase (Sdr) From Thermus Caliditerrae In Complex With Nadp
Organism: Thermus caliditerrae
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-06-26 Classification: OXIDOREDUCTASE Ligands: NAP, MG |
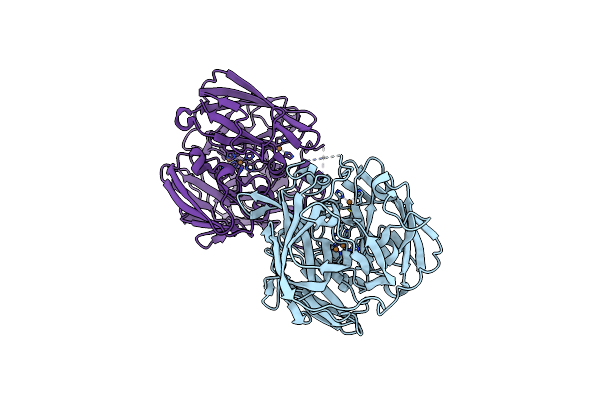 |
Crystal Structure Of A Multicopper Oxidase 3F3 Variant From Pyrobaculum Aerophilum
Organism: Pyrobaculum aerophilum str. im2
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2024-02-21 Classification: OXIDOREDUCTASE Ligands: CU, CL |
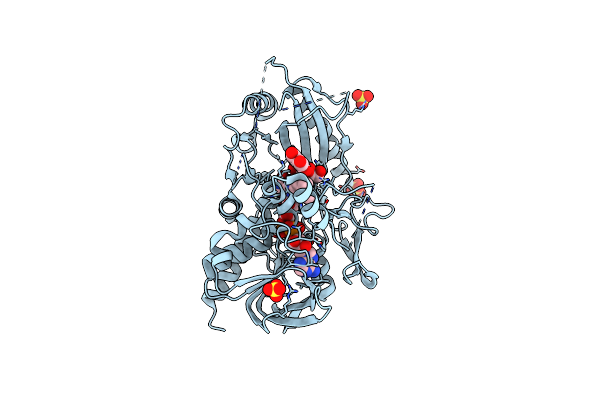 |
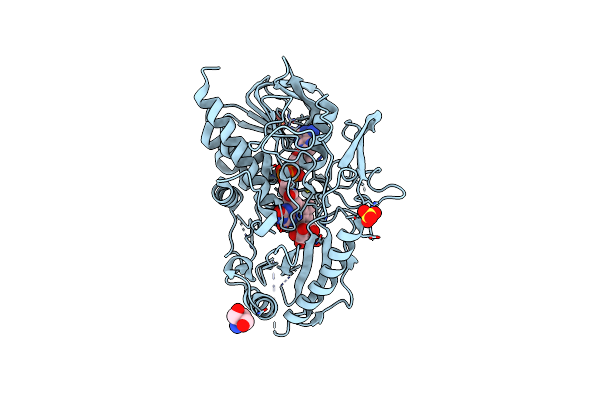
Crystal Structure Of A Bacterial Pyranose 2-Oxidase In Complex With Mangiferin
Organism: Pseudarthrobacter siccitolerans
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2023-08-16 Classification: OXIDOREDUCTASE Ligands: FAD, HZI, SO4 |
 |
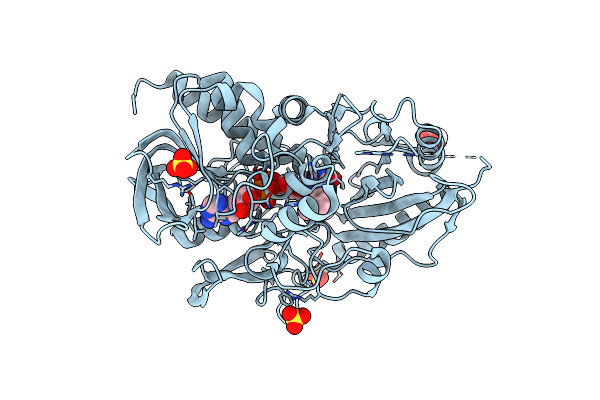
Organism: Pseudarthrobacter siccitolerans
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-07-05 Classification: OXIDOREDUCTASE Ligands: FAD, GLC, SO4, TRS |
 |
Crystal Structure Of A Bacterial Pyranose 2-Oxidase From Pseudoarthrobacter Siccitolerans
Organism: Pseudarthrobacter siccitolerans
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-06-14 Classification: OXIDOREDUCTASE Ligands: SO4, TRS, FAD |
 |
Stilbene Dioxygenase (Nov1) From Novosphingobium Aromaticivorans: Ser283Phe Mutant
Organism: Novosphingobium aromaticivorans
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2022-08-10 Classification: OXIDOREDUCTASE Ligands: FE, OXY |
 |
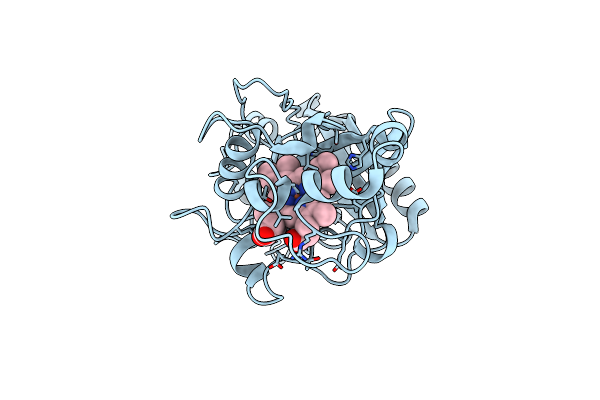
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2022-08-03 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
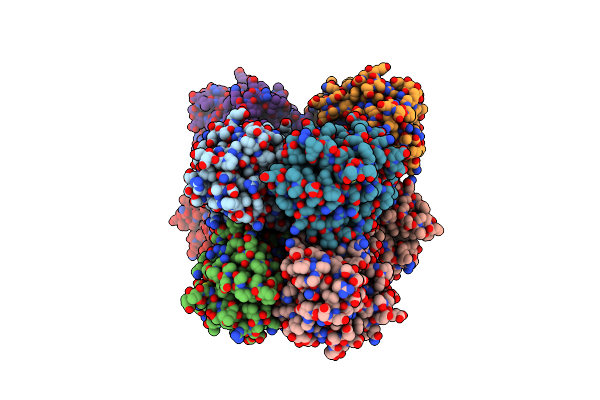
Crystal Structure Of A Dyp-Type Peroxidase 6E10 Variant From Pseudomonas Putida
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2022-08-03 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Crystal Structure Of A Dyp-Type Peroxidase 29E4 Variant From Pseudomonas Putida
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2022-08-03 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Crystal Structure Of Mcoa Multicopper Oxidase 2F4 Variant From The Hyperthermophile Aquifex Aeolicus
Organism: Aquifex aeolicus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2021-04-07 Classification: OXIDOREDUCTASE Ligands: CU, OXY, SO4, GOL |

