Search Count: 28
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA |
 |
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
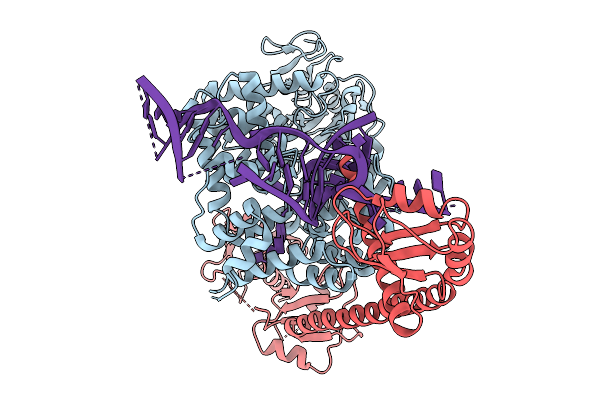 |
Cryo-Em Structure Of Ro60/La/Truncated Misfolded Human Pre-5S Rrna Complex With Fab, Composite Map
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA |
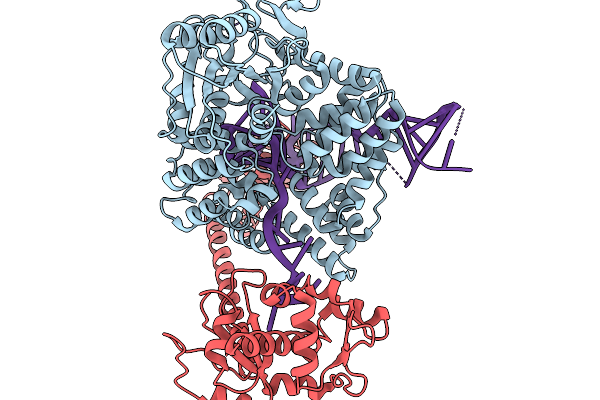 |
Cryo-Em Structure Of Ro60/La/Minimal Misfolded Pre-5S Rrna Complex With Fab, Composite Map
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-02-14 Classification: CELL CYCLE |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: STE, FAD |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: FAD, PJ8, CO2, STE |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: STE, FAD |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: FAD, PJ8, CO2, STE, 1PE |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: STE, FAD, PG4 |
 |
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2021-04-21 Classification: BIOSYNTHETIC PROTEIN Ligands: STE, FAD |
 |
Dark State Structure Of The C432S Mutant Of Fatty Acid Photodecarboxylase (Fap)
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2021-04-21 Classification: OXIDOREDUCTASE Ligands: STE, FAD, GOL |
 |
Crystal Structure Of Fatty Acid Photodecarboxylase In The Dark State Determined By Serial Femtosecond Crystallography At Room Temperature
Organism: Chlorella variabilis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-04-14 Classification: OXIDOREDUCTASE Ligands: FAD, STE |
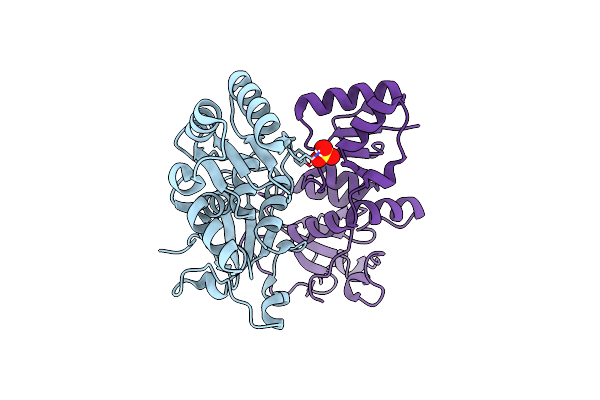 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2021-02-24 Classification: TRANSFERASE Ligands: SO4 |
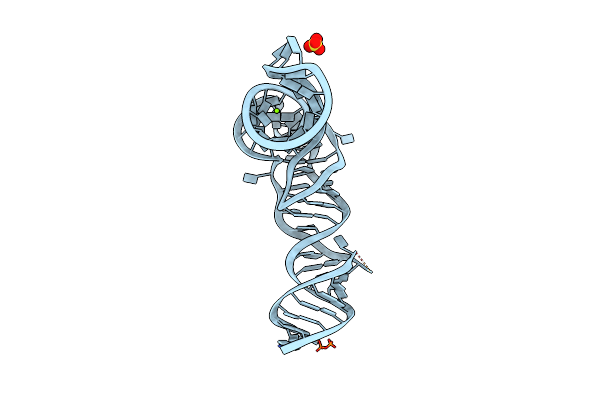 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2018-10-31 Classification: RNA Ligands: MG, SO4 |
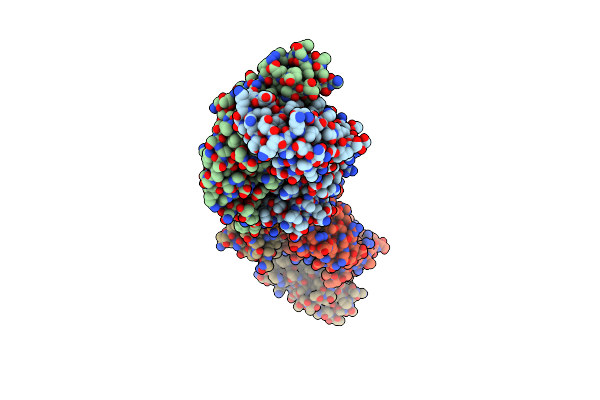 |
Organism: Homo sapiens, Human immunodeficiency virus 1
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2015-08-12 Classification: IMMUNE SYSTEM |
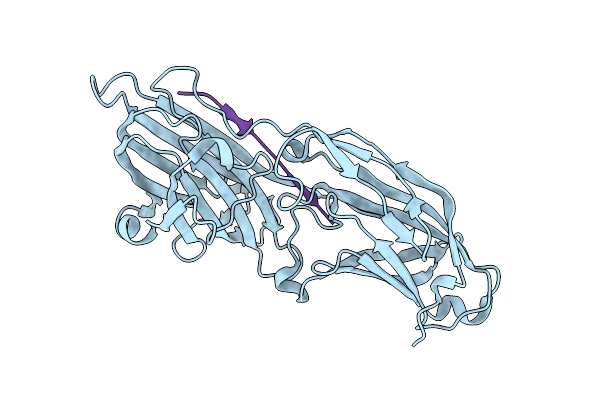 |
Organism: Staphylococcus aureus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2011-05-04 Classification: CELL ADHESION |
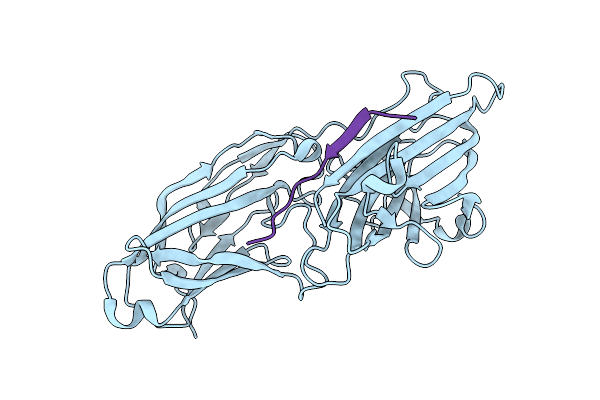 |
Organism: Staphylococcus aureus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2011-05-04 Classification: CELL ADHESION/BLOOD CLOTTING |
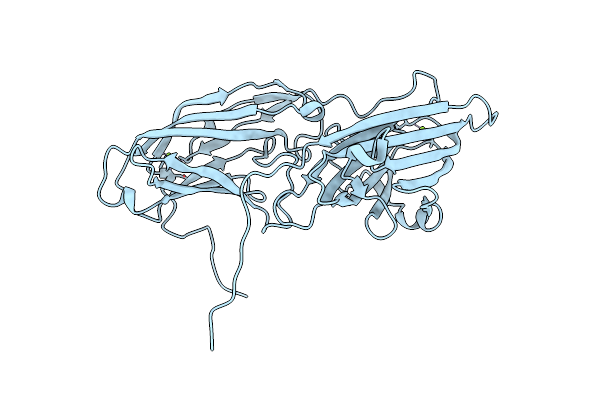 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2011-05-04 Classification: CELL ADHESION Ligands: MG |
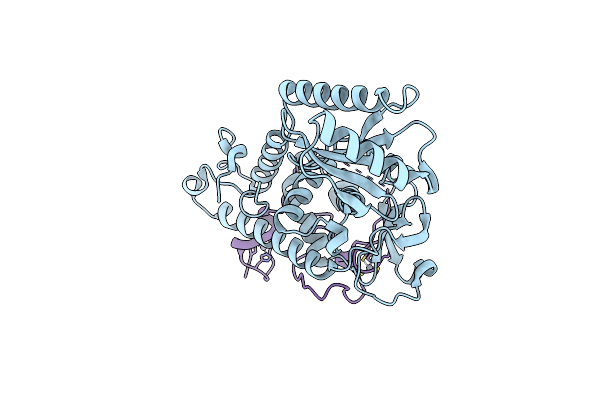 |
Structure And Function Of The Polymerase Core Of Tramp, A Rna Surveillance Complex
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2010-08-04 Classification: TRANSFERASE/RNA BINDING PROTEIN Ligands: ZN |

