Search Count: 157
All
Selected
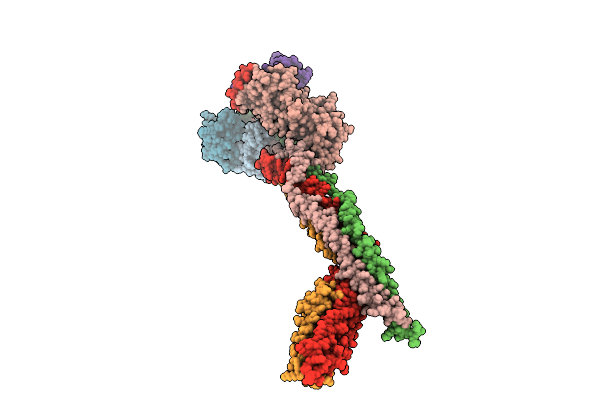 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Lacking The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
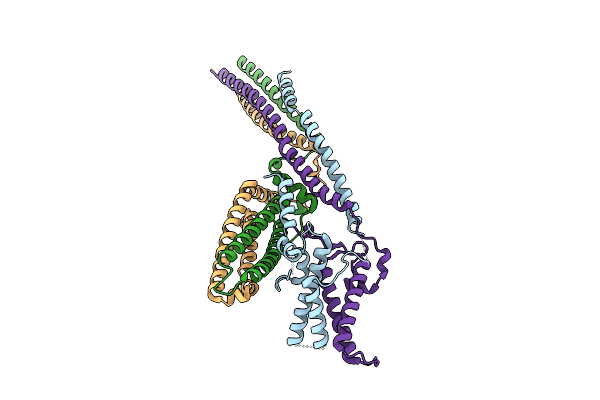 |
Cryo-Em Structure Of The Base Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
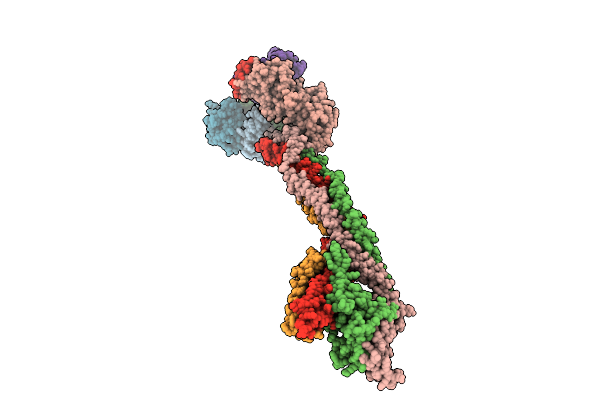 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae, Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.26 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: NAG |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.55 Å Release Date: 2025-12-31 Classification: SIGNALING PROTEIN Ligands: E2Q |
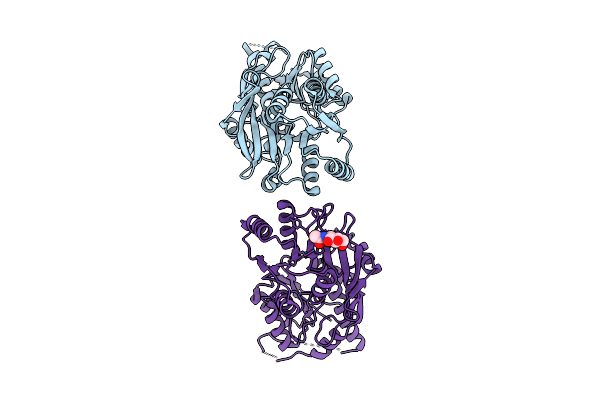 |
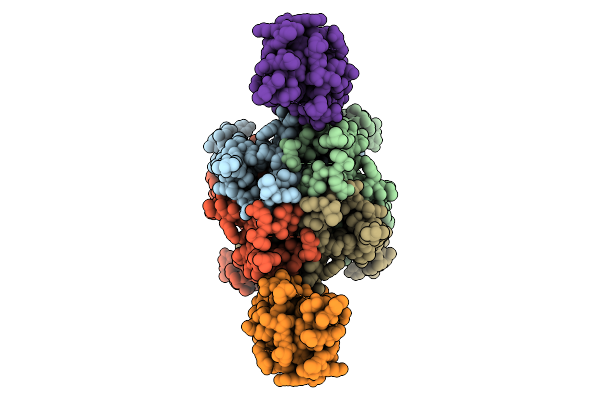
Cerebellar Glua2/4 Ntd Heterophilic Tetramer Interface (Focused Refinement)
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2025-12-24 Classification: SIGNALING PROTEIN Ligands: NAG |
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.66 Å Release Date: 2025-12-24 Classification: SIGNALING PROTEIN |
 |
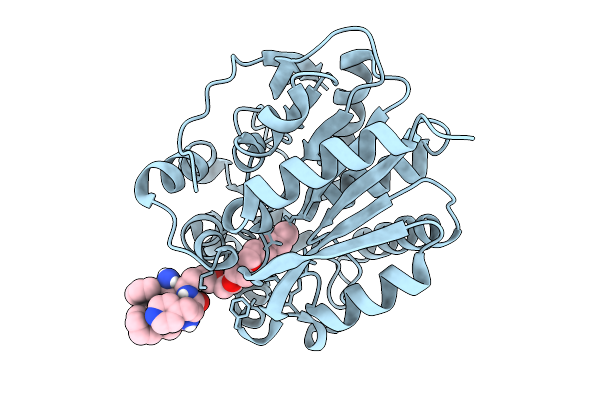
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: A1I4Z |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: A1I4X |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: A1I4Y |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.33 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: A1I4W |
 |
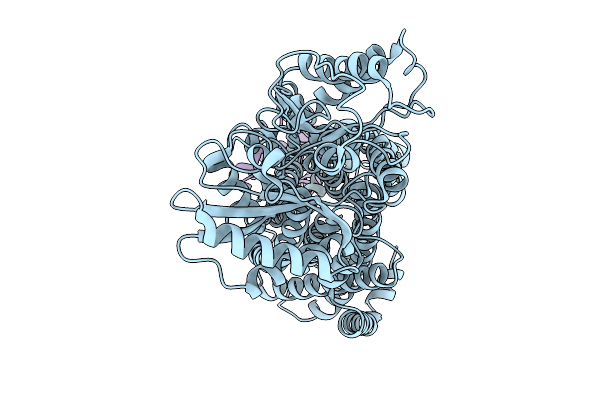
Organism: Rhodococcus sp. (in: high g+c gram-positive bacteria)
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2025-10-08 Classification: LIGASE Ligands: CL, A1JC1 |
 |
Organism: Mycobacterium tuberculosis, Aequorea victoria, Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-10-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Mycobacterium tuberculosis h37rv, Mycobacterium tuberculosis, Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-08-13 Classification: MEMBRANE PROTEIN Ligands: L9Q |
 |
Organism: Mycobacterium tuberculosis, Aequorea victoria, Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-08-13 Classification: MEMBRANE PROTEIN Ligands: L9Q |
 |
Solution Structure And Chemical Shift Assignments For Hmg-D Y12F Mutant Complexed To A 14:12 Da2 Bulge Dna
Organism: Drosophila melanogaster, Synthetic construct
Method: SOLUTION NMR Release Date: 2024-09-11 Classification: DNA BINDING PROTEIN |
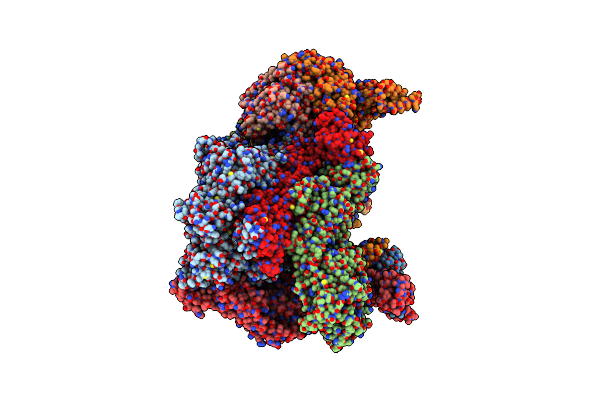 |
High-Resolution Structure Of The Anaphase-Promoting Complex/Cyclosome (Apc/C) Bound To Co-Activator Cdh1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2024-08-14 Classification: CELL CYCLE Ligands: ZN |
 |
Organism: Gallus gallus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-07-31 Classification: DNA BINDING PROTEIN |
 |
Organism: Gallus gallus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2024-07-31 Classification: DNA BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.35 Å Release Date: 2023-10-18 Classification: PROTEIN FIBRIL |

