Search Count: 626
All
Selected
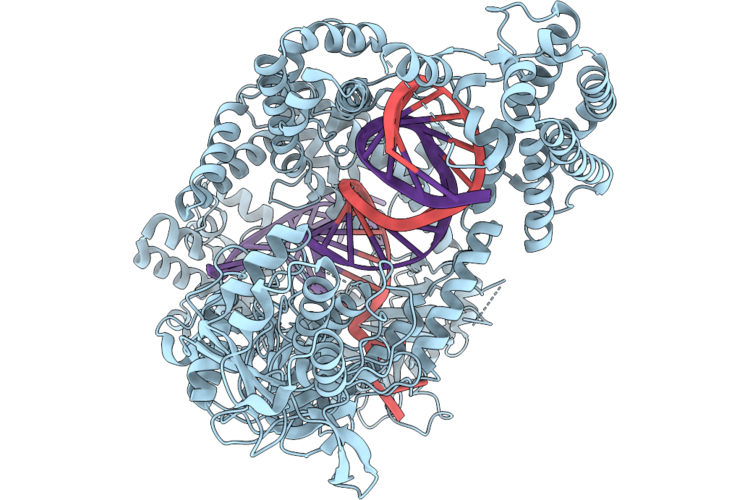 |
Organism: Acidaminococcus sp. bv3l6, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
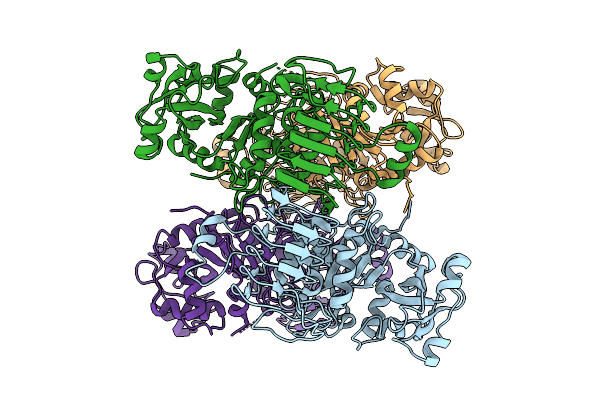 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.95 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
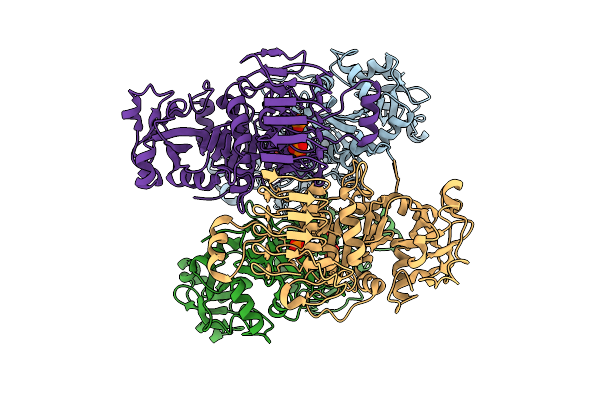 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: PO4 |
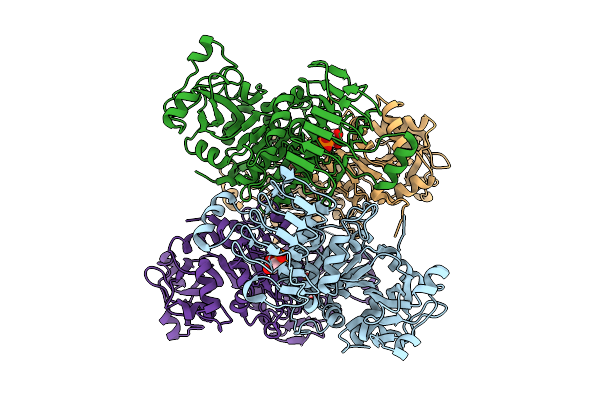 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: 3PG |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: 3PG, ADQ, MG |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN Ligands: 3PG, ATP, MG |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.36 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.65 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.38 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.49 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
Organism: Enterococcus faecalis
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN Ligands: CA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA/RNA |
 |
Organism: Enterococcus faecalis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.76 Å Release Date: 2026-04-01 Classification: DNA BINDING PROTEIN/DNA Ligands: CA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-18 Classification: LIGASE Ligands: CL |
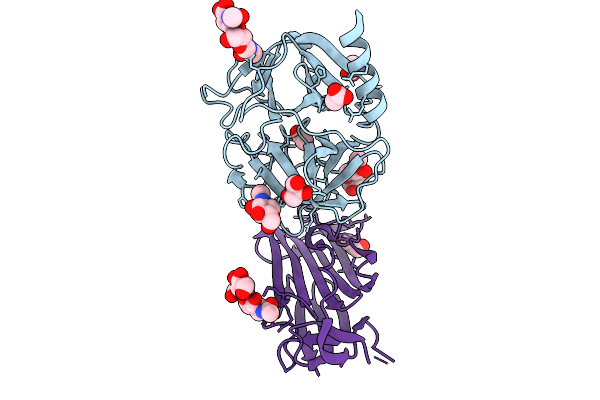 |
Pr3 S203A I221N W222N G223T Mutant In Complex With The Extracellular Domain Of Cd177
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-25 Classification: IMMUNE SYSTEM Ligands: NAG, CL, GOL, ACT |
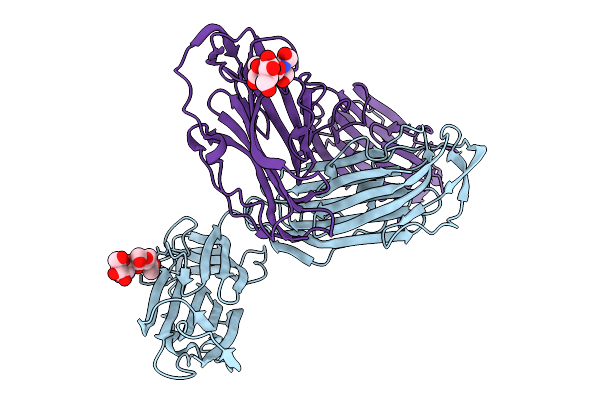 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-25 Classification: IMMUNE SYSTEM Ligands: CL |
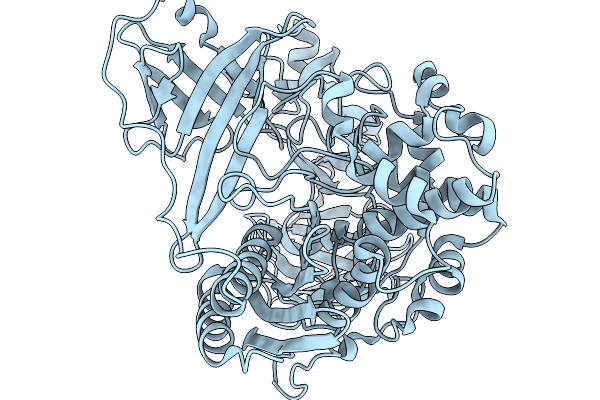 |
Organism: Paenibacillus
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-02-18 Classification: HYDROLASE |
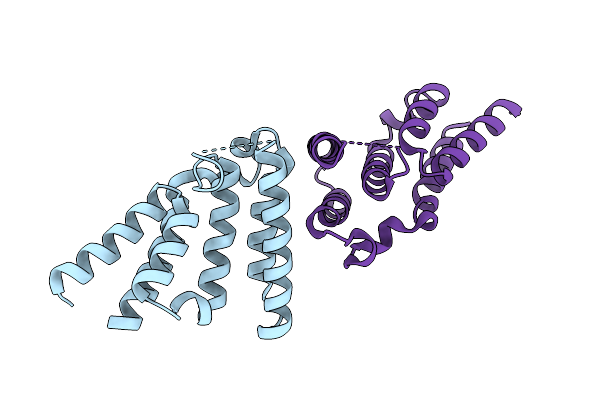 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-02-18 Classification: LIGASE |

