Search Count: 111
All
Selected
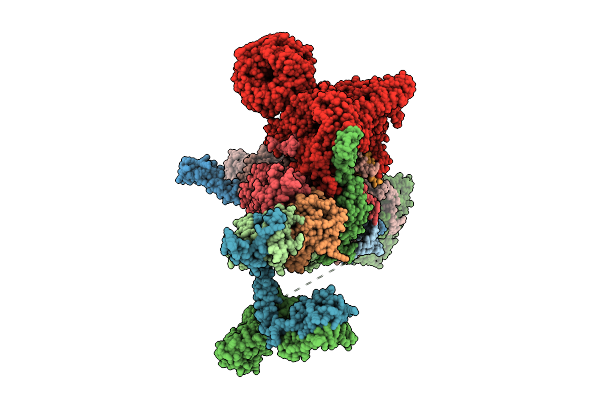 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Proximal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
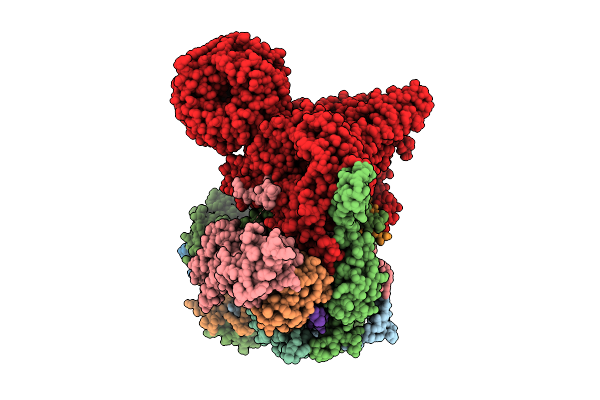 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Core)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
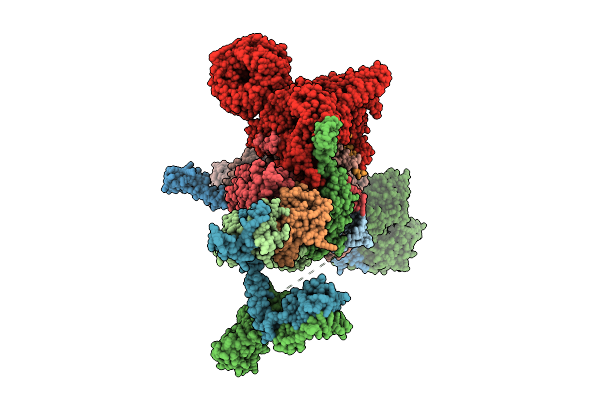 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Distal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
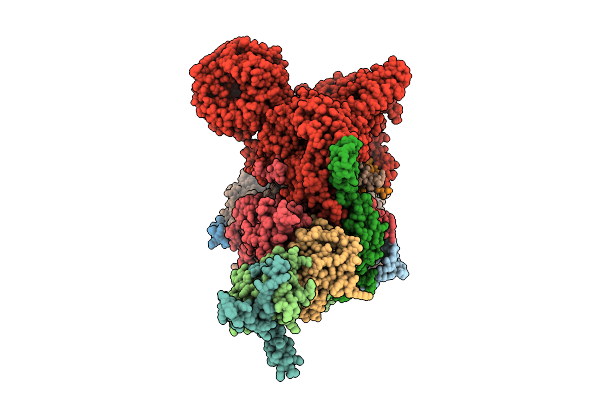 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
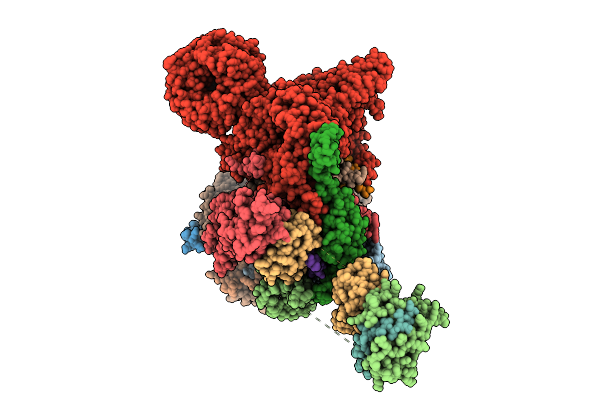 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
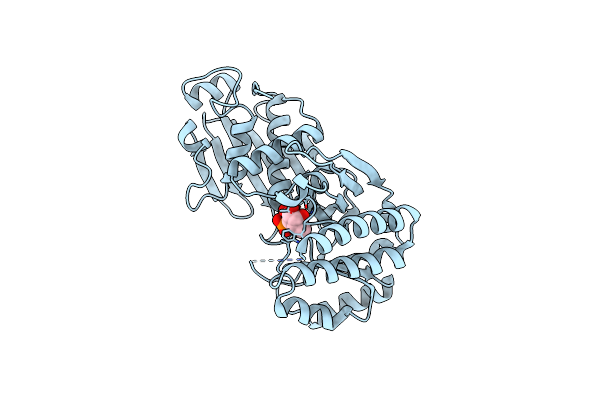 |
Organism: Bacteroides fragilis
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: ED6 |
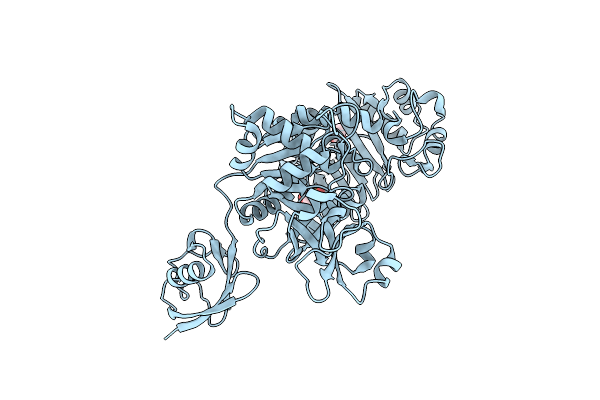 |
Organism: Saccharomyces cerevisiae s288c, Bacteroides fragilis
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: EPE, ACT |
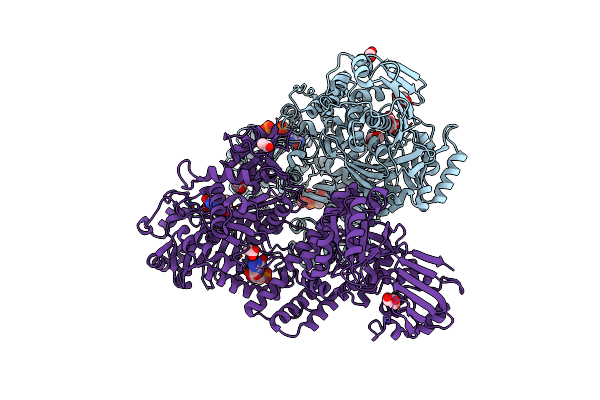 |
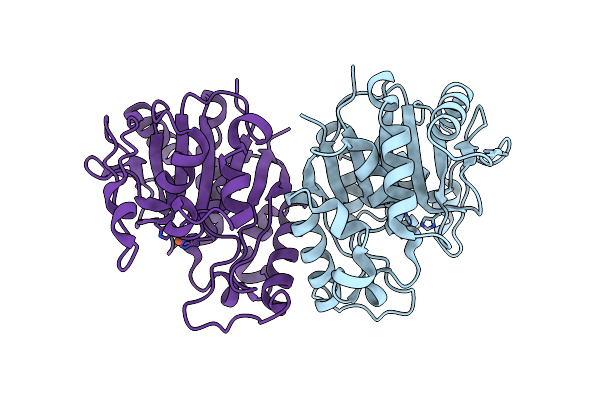
Organism: Bacteroides fragilis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: GTP, FMT, MLA, GOL, GDP, ACT |
 |
Organism: Beta vulgaris
Method: X-RAY DIFFRACTION Resolution:3.08 Å Release Date: 2024-04-24 Classification: OXIDOREDUCTASE Ligands: FE |
 |
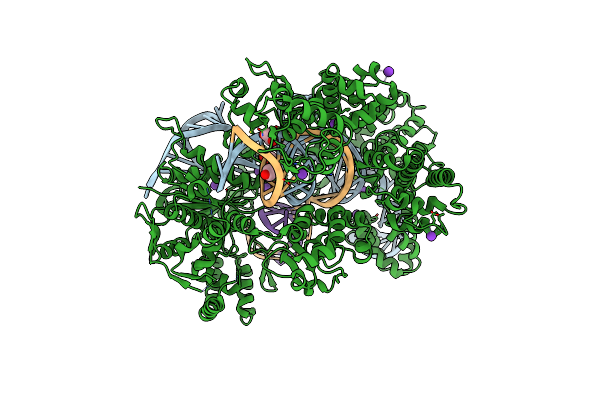
Organism: Synthetic construct, Streptococcus pyogenes
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K, EDO |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K, EDO |
 |
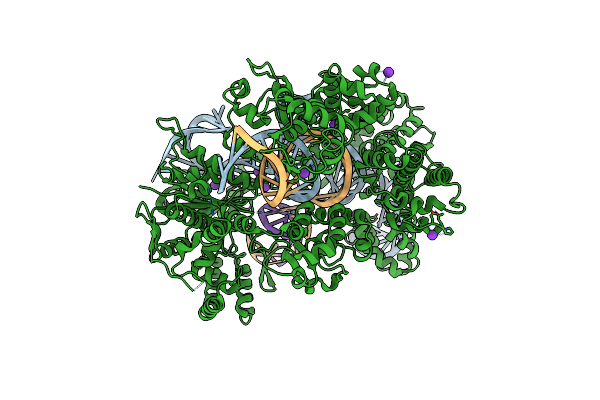
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K, EDO |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
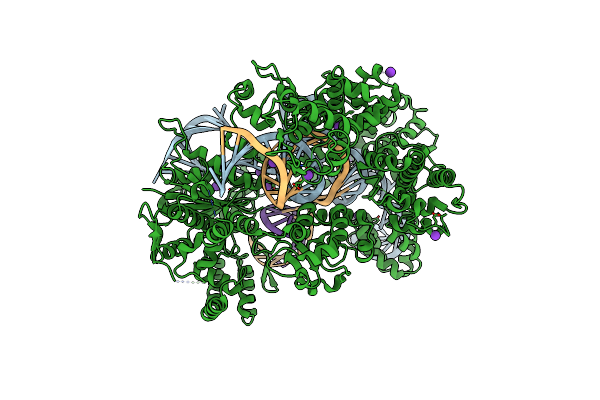
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
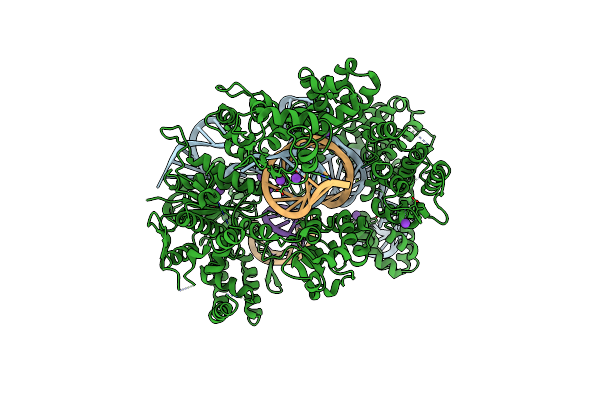
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
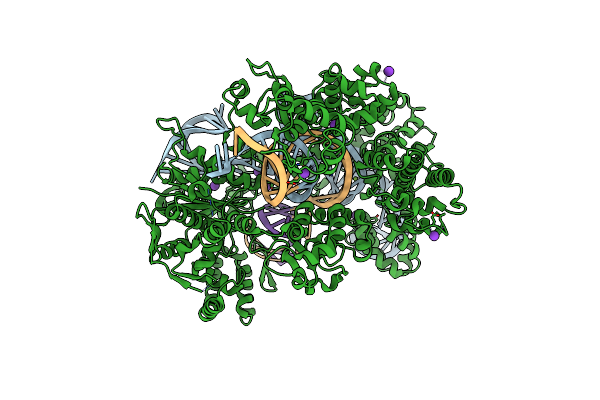
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |
 |
Organism: Streptococcus pyogenes, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2022-10-26 Classification: HYDROLASE Ligands: MG, K |

