Search Count: 1,215
All
Selected
 |
Crystal Structure Of Sars-Cov-2 Main Protease G15S Mutant In Complex With Leritrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: A1EN0 |
 |
Crystal Structure Of Sars-Cov-2 Main Protease G15S Mutant In Complex With Leritrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: 7ON |
 |
Crystal Structure Of Sars-Cov-2 Main Protease K90R Mutant In Complex With Leritrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: A1EN0 |
 |
Crystal Structure Of Sars-Cov-2 Main Protease E166N Mutant In Complex With Leritrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: A1EN0 |
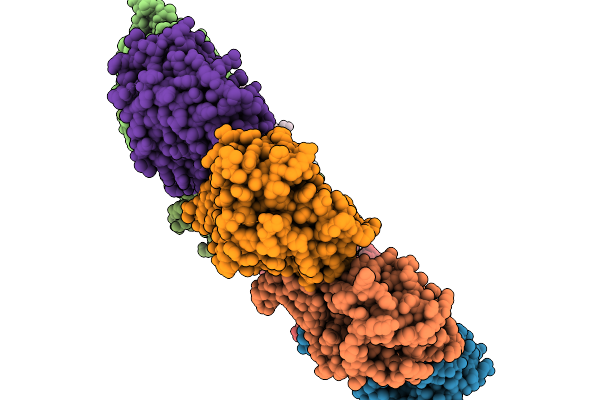 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: CHAPERONE |
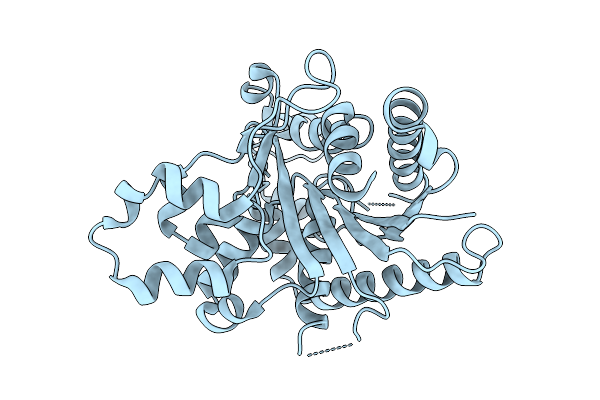 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-03-18 Classification: LIPID TRANSPORT |
 |
Crystal Structure Of Sars-Cov-2 Main Protease K90R Mutant In Complex With Ibuzatrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-03-11 Classification: VIRAL PROTEIN Ligands: YDL |
 |
Crystal Structure Of Sars Main Protease In Complex With Ibuzatrelvir Pomotrelvir
Organism: Severe acute respiratory syndrome-related coronavirus
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-03-04 Classification: VIRAL PROTEIN Ligands: ZQB |
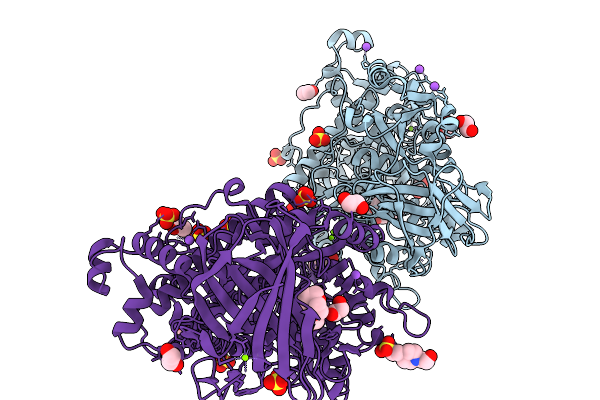 |
Organism: Catenovulum maritimum
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: EDO, O4B, MG, SO4, NA, CL, EPE, PEG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.00 Å Release Date: 2026-03-04 Classification: PROTEIN TRANSPORT |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.20 Å Release Date: 2026-03-04 Classification: PROTEIN TRANSPORT |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:6.60 Å Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:6.80 Å Release Date: 2026-03-04 Classification: TRANSPORT PROTEIN |
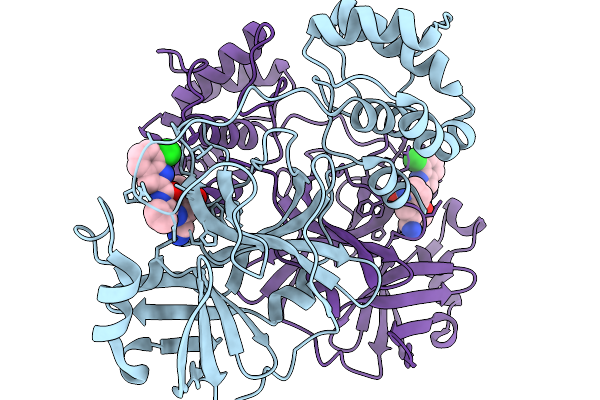 |
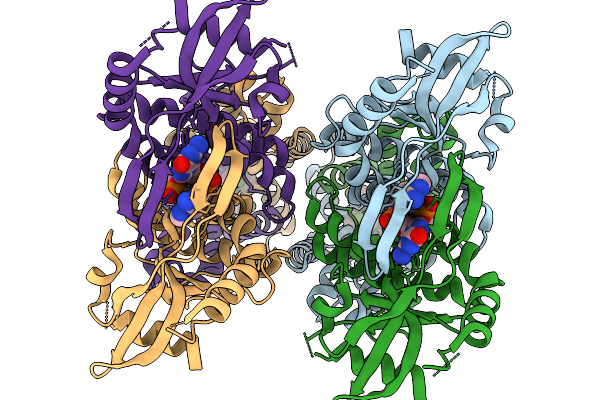
Crystal Structure Of Sars-Cov-2 Main Protease E166R Mutant In Complex With Pomotrelvir
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: ZQB |
 |
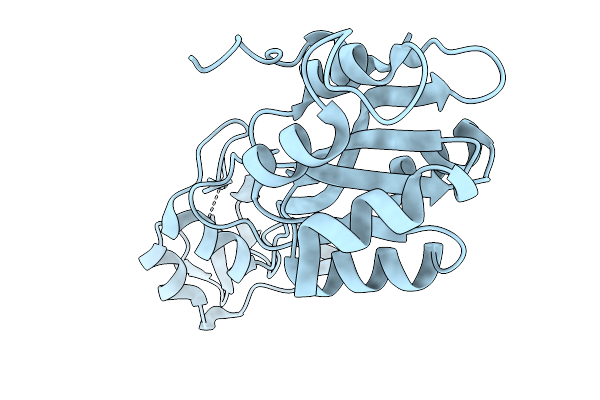
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.93 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM Ligands: 9IM, 1SY |
 |
Organism: Chaetomium thermophilum (strain dsm 1495 / cbs 144.50 / imi 039719)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN |
 |
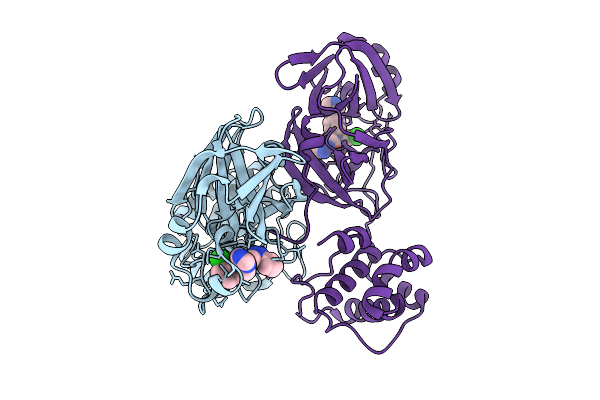
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-28 Classification: PROTEIN BINDING Ligands: NAD |
 |
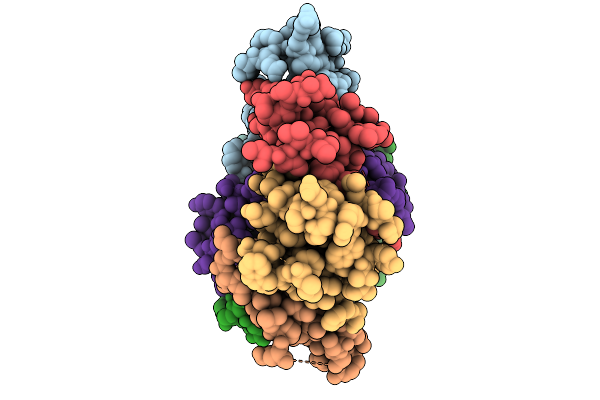
Organism: Severe acute respiratory syndrome-related coronavirus
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2026-01-21 Classification: VIRAL PROTEIN Ligands: XIU |
 |
Structure Of The Bacteroides Fragilis Nctc9343 T6Ss Hcp2-Hcp3 Heterohexamer In Complex With The Effector Bte1
Organism: Bacteroides fragilis nctc 9343
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN |
 |
Organism: Bacteroides fragilis nctc 9343
Method: ELECTRON MICROSCOPY Resolution:3.11 Å Release Date: 2026-01-21 Classification: STRUCTURAL PROTEIN |

