Search Count: 1,593
All
Selected
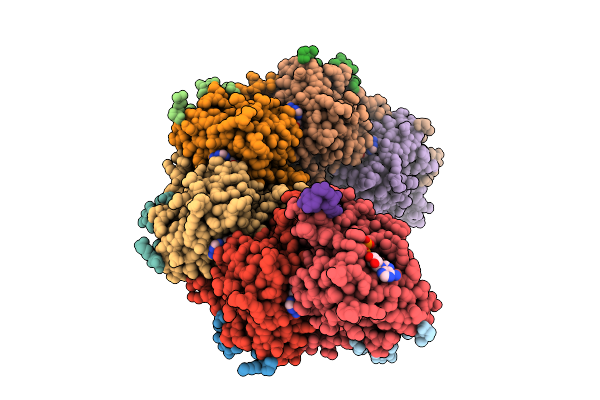 |
Rad51-Ssdna Filament In Complex With Calcium And Atp Bound By The Rad54B N-Terminus
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.61 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: ATP, CA |
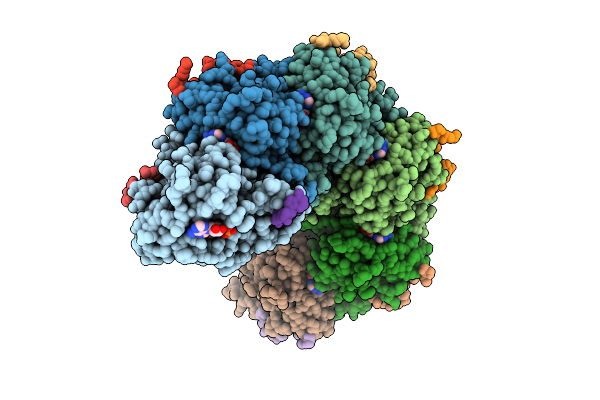 |
Rad51-Ssdna Filament In Complex With Magnesium And Atp Bound By The Rad54B N-Terminus (Peptide)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: K, ATP, MG |
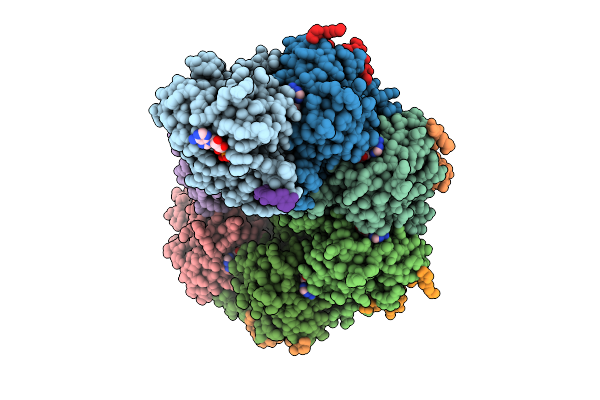 |
Rad51-Ssdna Filament In Complex With Magnesium And Atp Bound By The Rad54B N-Terminus (Beta-Barrel)
Organism: Homo, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: K, ATP, MG |
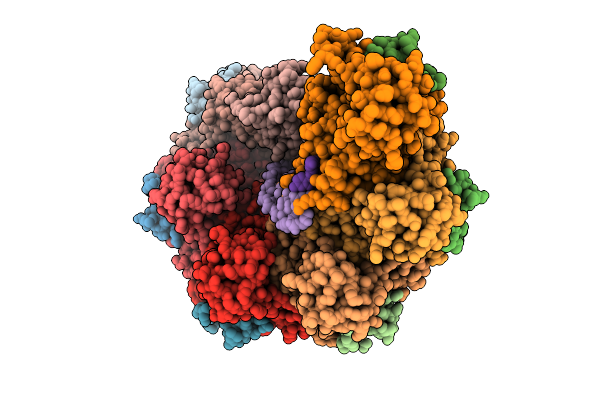 |
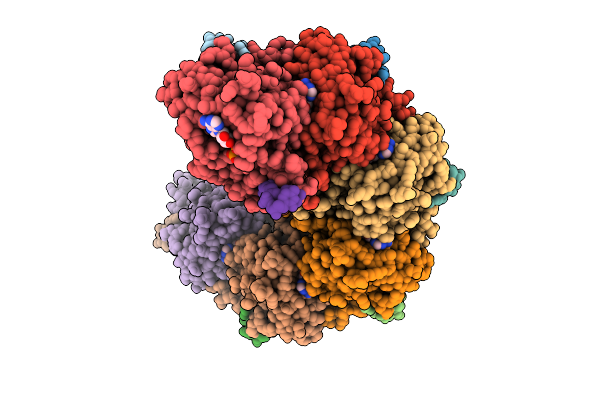
Rad51-Dsdna Filament In Complex With Calcium And Atp Bound By The Rad54B N-Terminus
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: ATP, CA |
 |
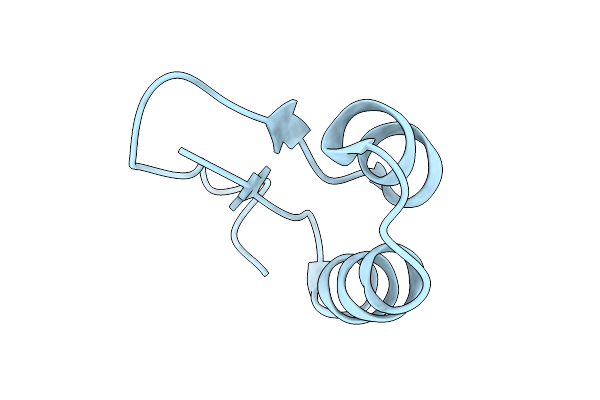
Rad51-Ssdna Filament In Complex With Calcium And Atp Bound By The Rad54 N-Terminus
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: ATP, CA |
 |
Organism: Crambe hispanica subsp. abyssinica
Method: ELECTRON CRYSTALLOGRAPHY Resolution:0.85 Å Release Date: 2026-02-25 Classification: PLANT PROTEIN |
 |
Room-Temperature X-Ray Structure Of Thermus Thermophilus Shmt In Complex With Tetrahydrofolate (Thf)
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: TRANSFERASE Ligands: SO4, THG |
 |
Room-Temperature Joint X-Ray/Neutron Structure Of Thermus Thermophilus Shmt In Complex With Plp-Gly External Aldimine And 5-Methyl-Tetrahydrofolate (5Mthf)
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.10 Å, 1.70 Å Release Date: 2026-02-04 Classification: TRANSFERASE Ligands: SO4, PLG, C2F, DOD |
 |
Organism: Aspergillus fumigatus
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: GN1, CL |
 |
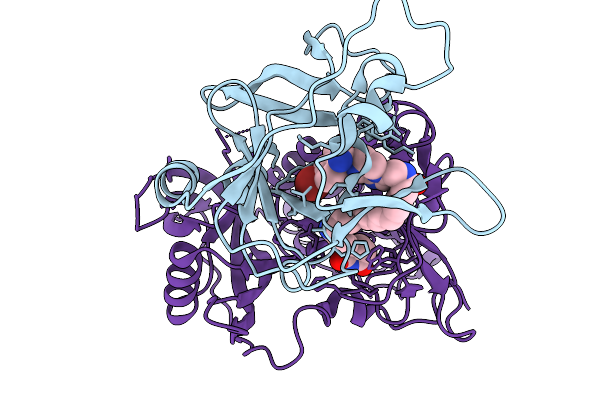
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2026-01-14 Classification: TRANSCRIPTION Ligands: A1CNQ |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-01-14 Classification: LIGASE Ligands: ZN, A1CN1 |
 |
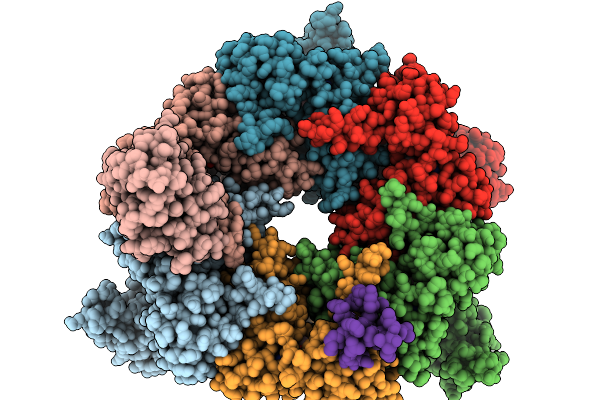
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-12-24 Classification: CHAPERONE Ligands: ADP |
 |
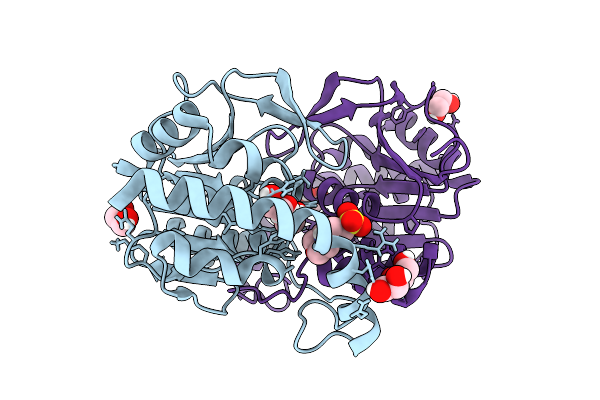
Joint X-Ray/Neutron Room Temperature Structure Of Perdeuterated Leca Lectin In Complex With Deuterated Galactose
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.49 Å, 1.9 Å Release Date: 2025-12-24 Classification: SUGAR BINDING PROTEIN Ligands: GLA, CA |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: CHAPERONE Ligands: ADP |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:0.89 Å Release Date: 2025-10-29 Classification: LYASE Ligands: EDO, MES |
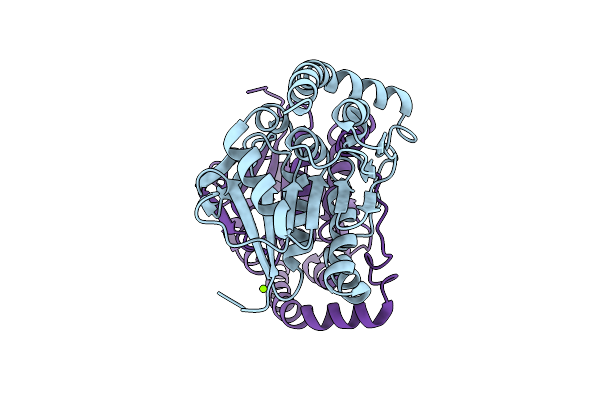 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-10-29 Classification: LYASE Ligands: MG, CL |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2025-10-29 Classification: LYASE Ligands: CL |
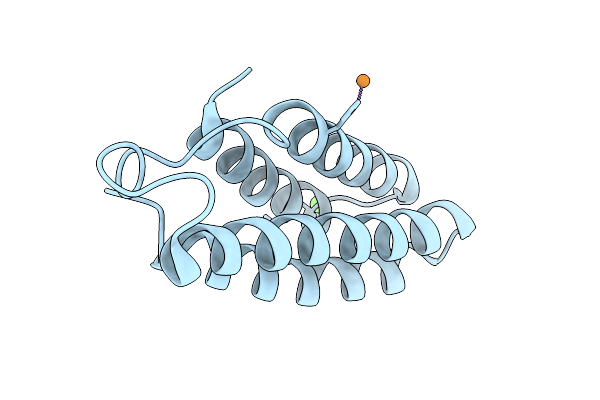 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-08-06 Classification: DE NOVO PROTEIN Ligands: CU, CA |
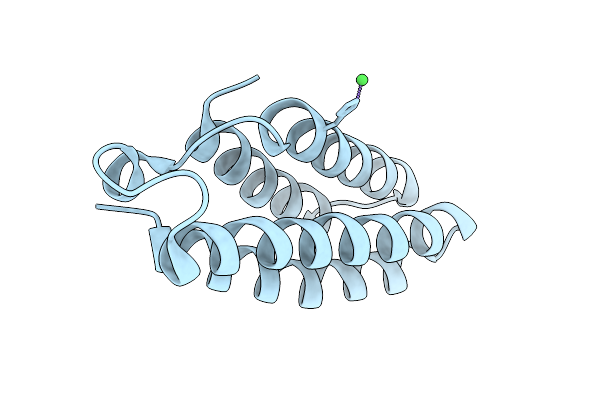 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2025-08-06 Classification: DE NOVO PROTEIN Ligands: NI |
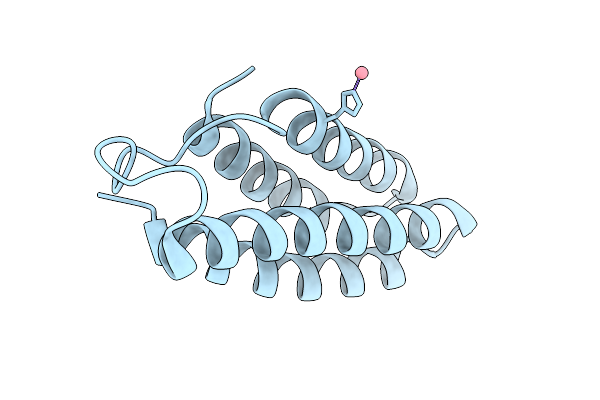 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-08-06 Classification: DE NOVO PROTEIN Ligands: CO |

