Search Count: 31,535
All
Selected
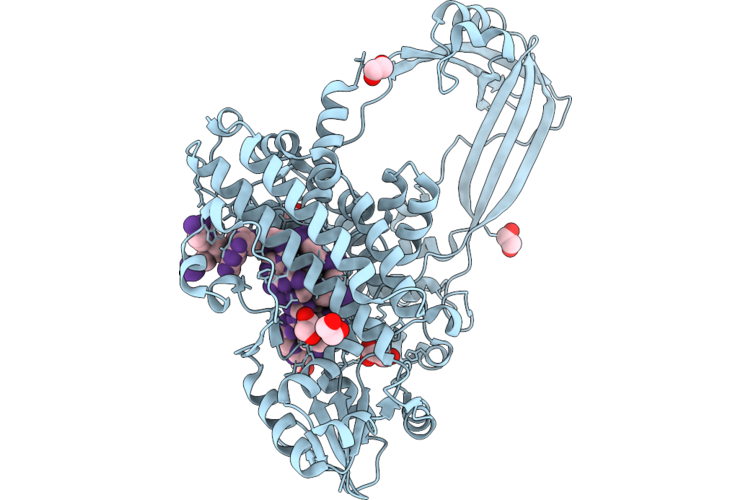 |
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-29 Classification: ISOMERASE/DNA Ligands: MLI, EDO |
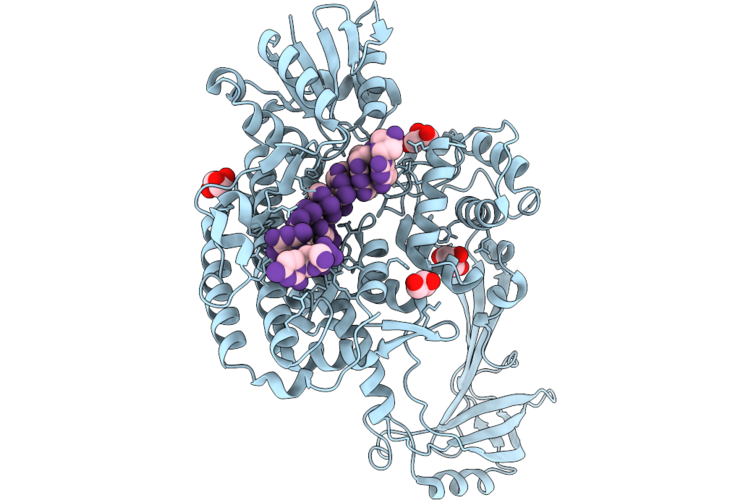 |
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2026-04-29 Classification: ISOMERASE/DNA Ligands: GOL |
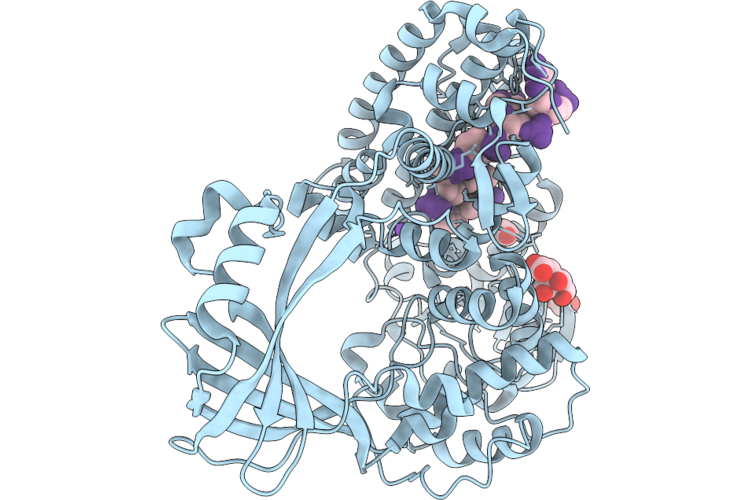 |
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-29 Classification: ISOMERASE/DNA Ligands: ACT, FLC, EDO, NA |
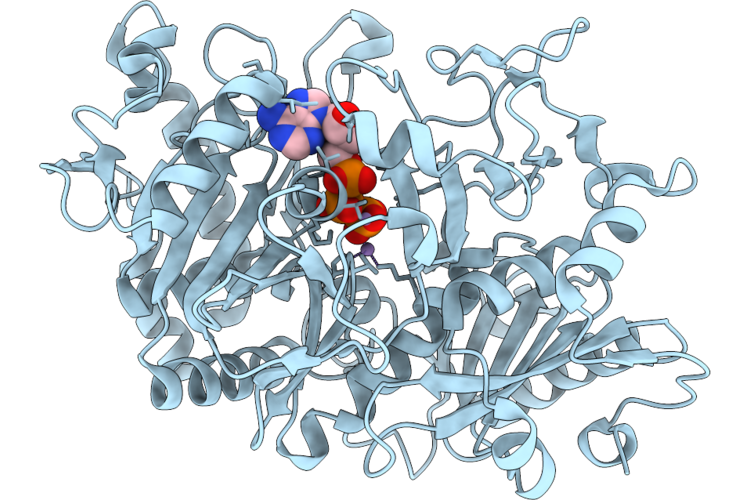 |
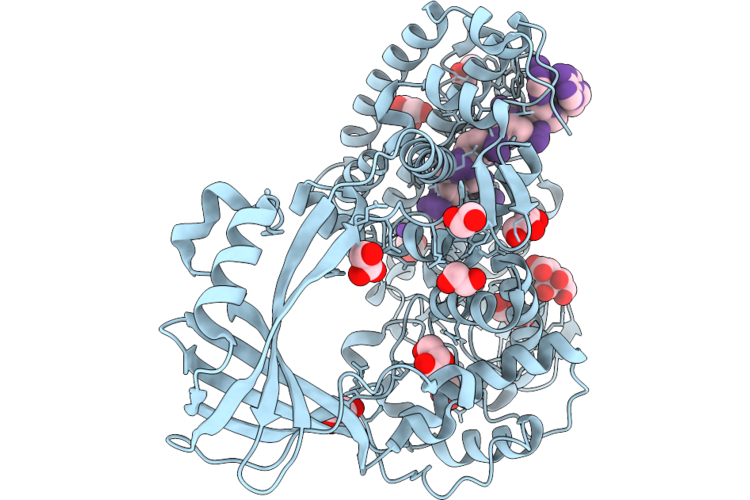
Structure Of Escherichia Coli Phosphoenolpyruvate Carboxykinase With Bound Atp, Mg2+, And Mn2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-29 Classification: LYASE Ligands: ATP, MN, MG |
 |
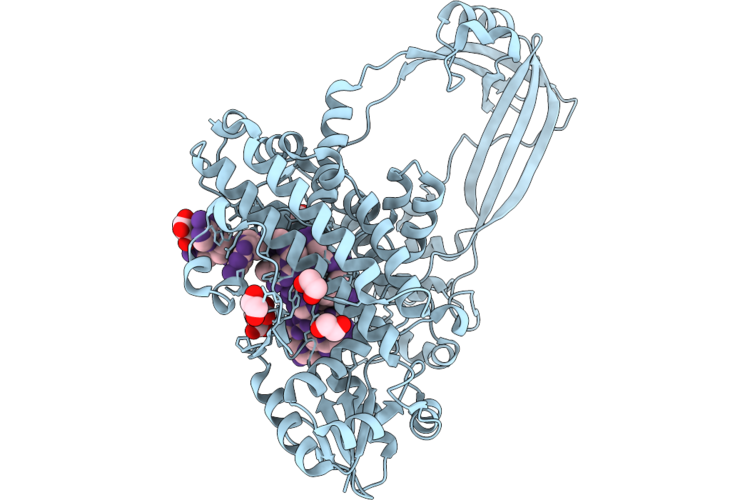
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-29 Classification: ISOMERASE/DNA Ligands: GOL, EDO, FLC, ACT, NA |
 |
Organism: Escherichia coli, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-29 Classification: ISOMERASE/DNA Ligands: EDO, SIN, CL, MLA |
 |
Fkbp12 In Complex With Bifunctional Ligand 1Ad And The First Bromodomain Of Brd4
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-29 Classification: ISOMERASE Ligands: A1JAZ |
 |
Crystal Structure Of Catalytic Domain Of Rv1625C Bound To Inhibitory Darpin G11
Organism: Mycobacterium tuberculosis, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-29 Classification: SIGNALING PROTEIN Ligands: GOL |
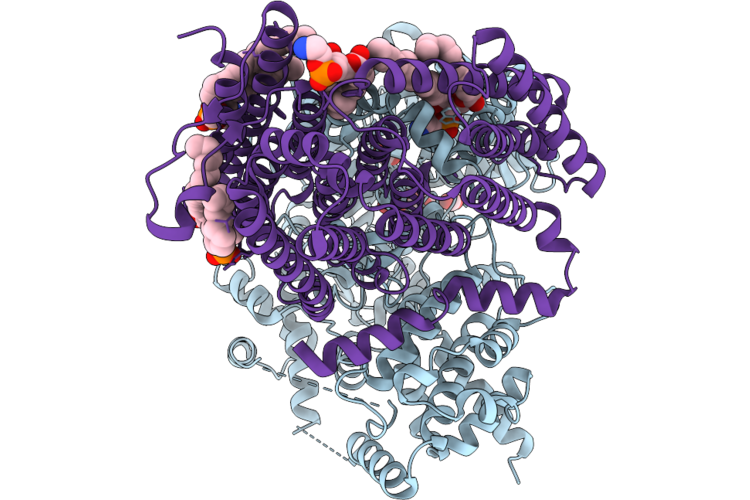 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
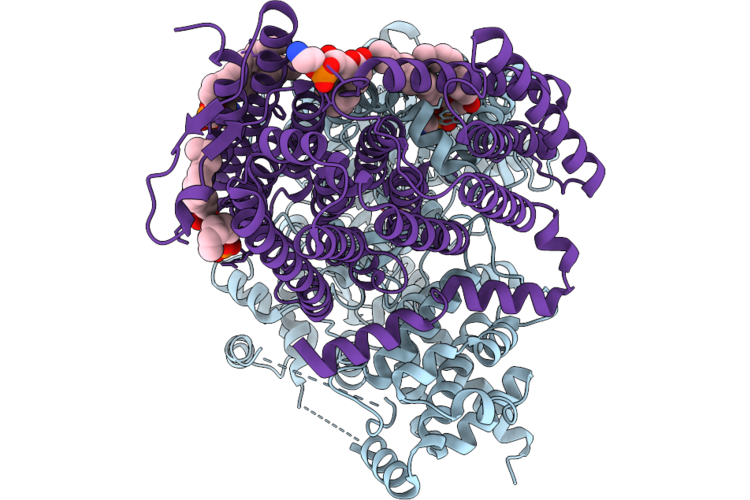 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, BCT, CO2, 6OU |
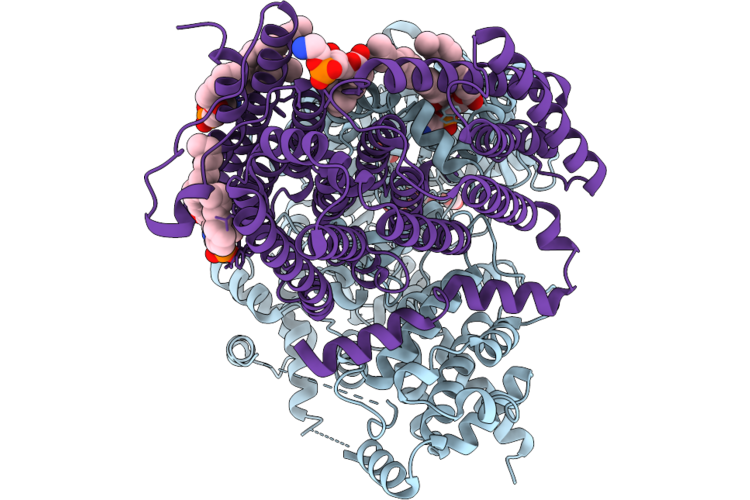 |
Organism: Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: ZN, CO2, 6OU |
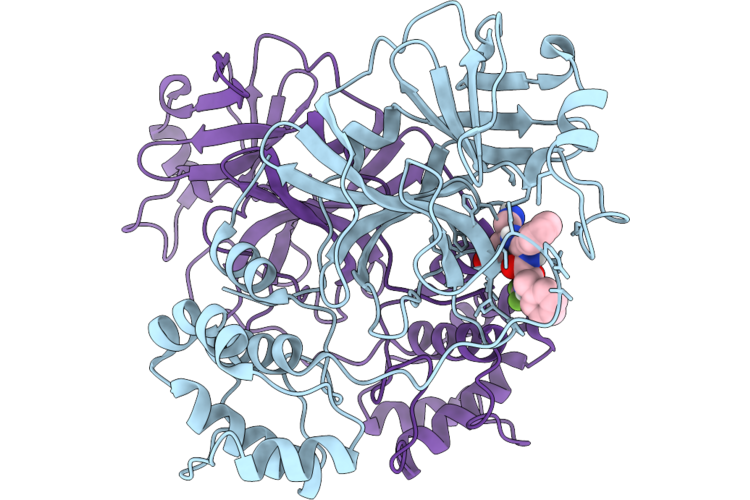 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: A1DAK |
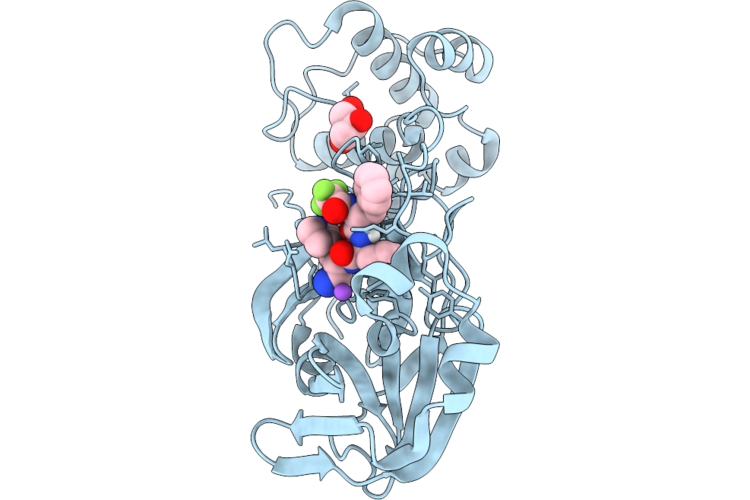 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: GOL, A1DA8, NA |
 |
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
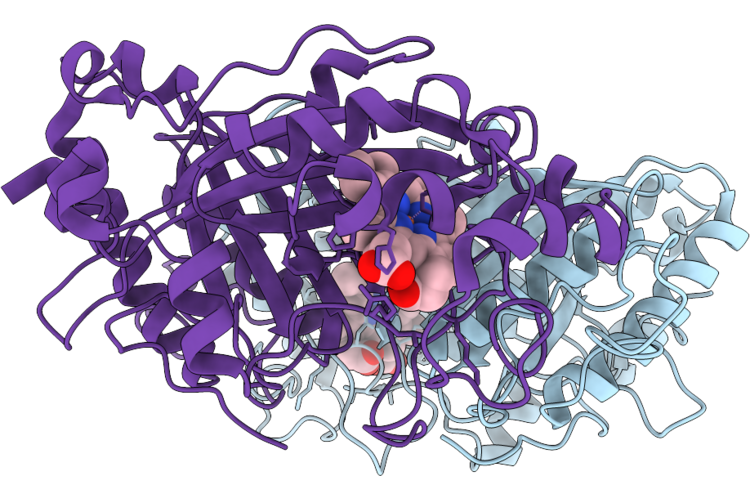 |
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
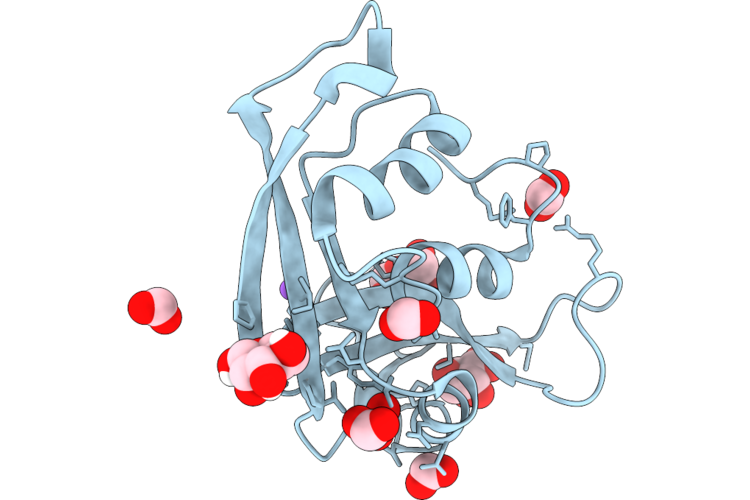 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: MAN, FMT, BMA, NA |
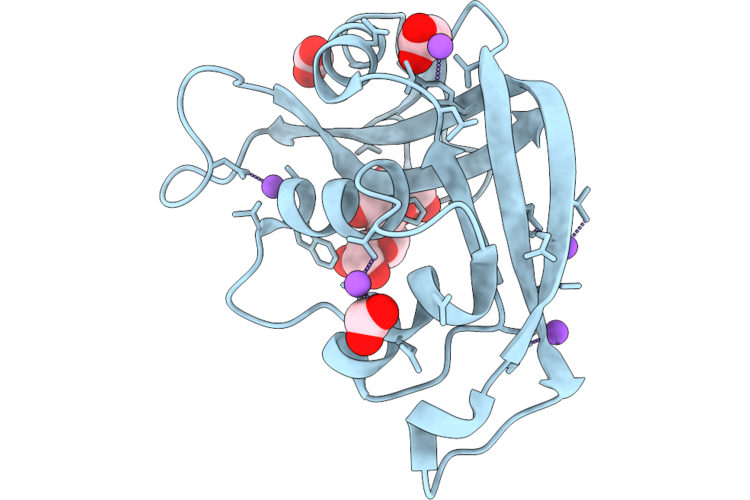 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:0.94 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, FMT, NA |
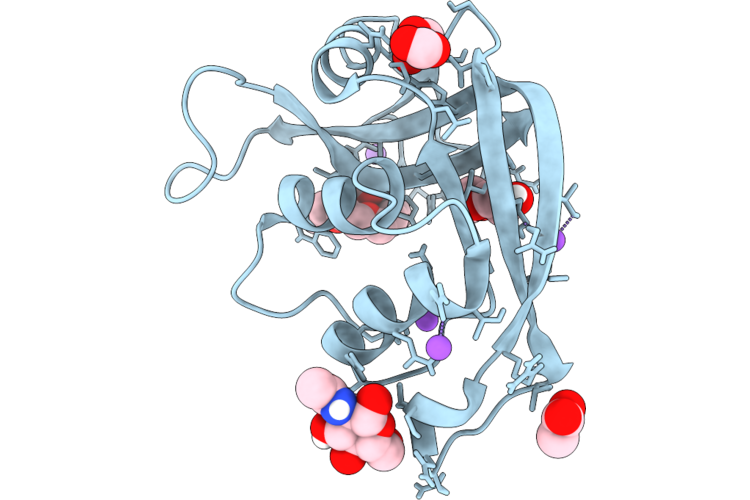 |
Structural Characterization Of A N-Acetyl-D-Glucosamine-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, NAG, PG4, FMT, PEG, NA |
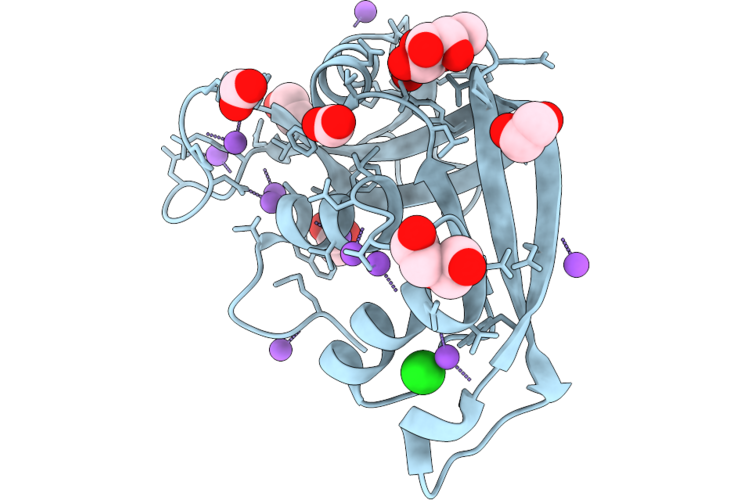 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: GLA, ACT, FMT, PEG, EDO, NA, CL |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |

