Search Count: 281
All
Selected
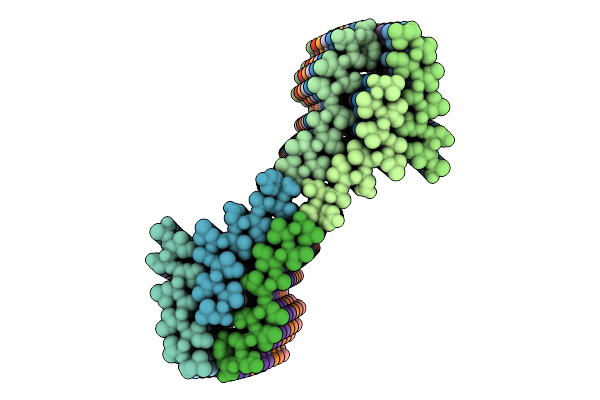 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
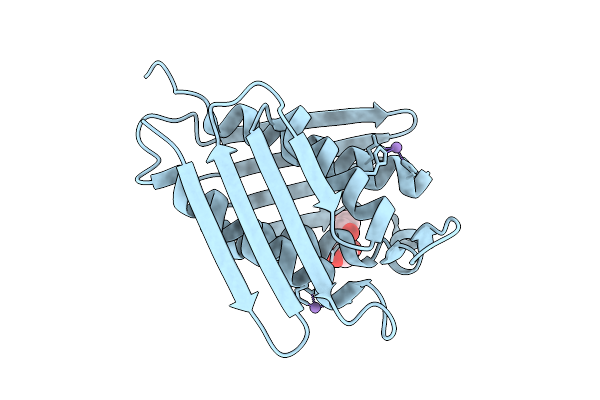 |
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-03-18 Classification: LYASE Ligands: MN, EDO, CL |
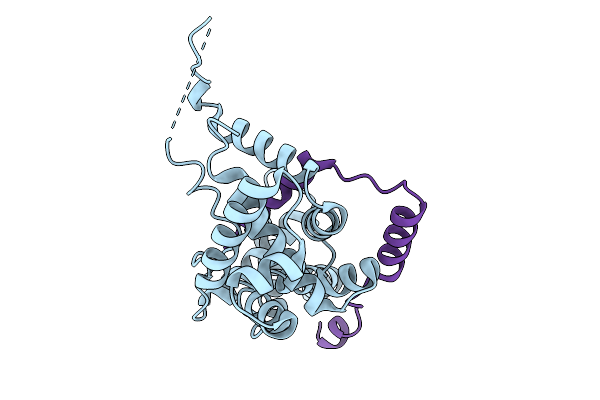 |
Crystal Structure Of Sigma28/Flgm Complex From Pseudomonas Aeruginosa At 1.95 Angstrom Resolution
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION |
 |
Cryo-Em Structure Of Sigma28-Rnap Open Promoter Complex From Pseudomonas Aeruginosa
|
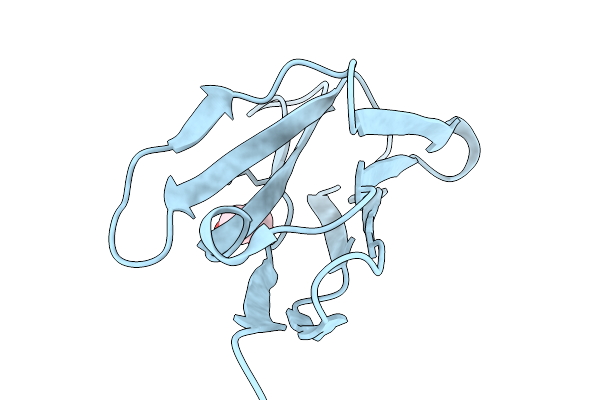 |
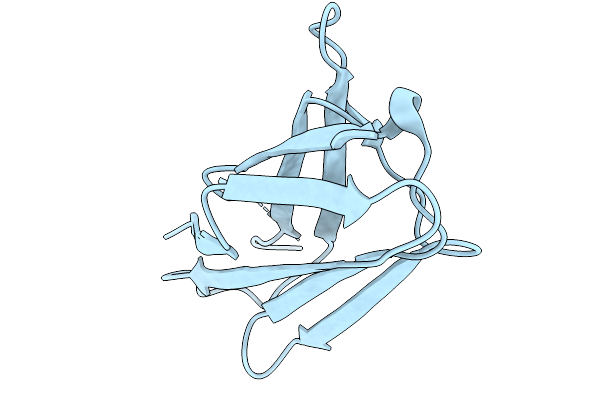
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM Ligands: EDO |
 |
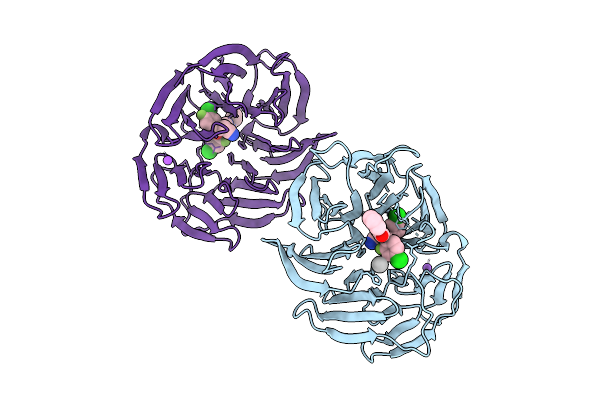
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM |
 |
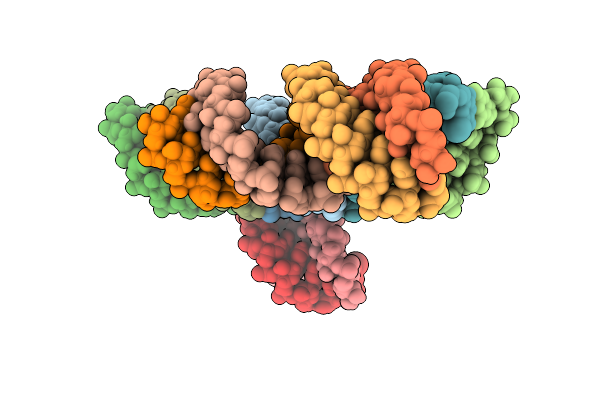
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-11-26 Classification: TRANSPORT PROTEIN Ligands: A1CY9, NA, UNX |
 |
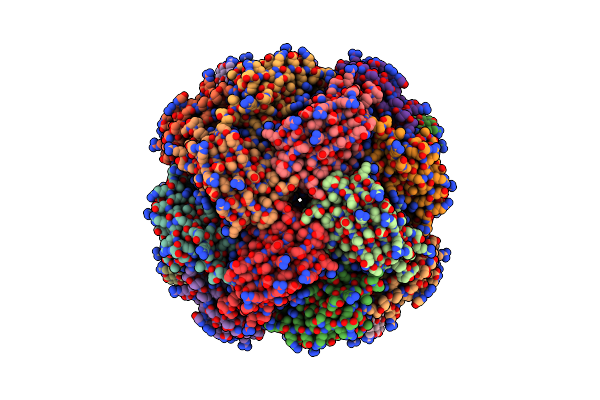
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-11-05 Classification: RNA |
 |
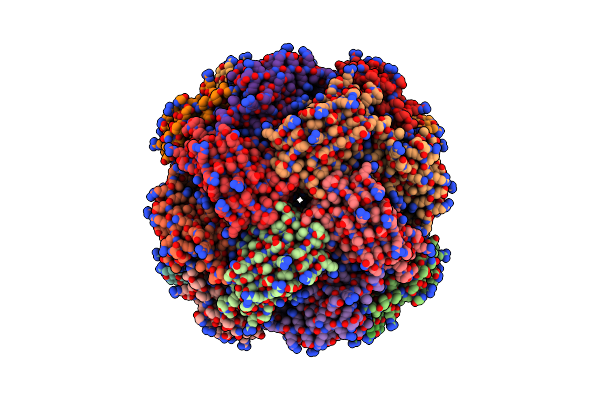
Imidazole Glycerol Phosphate Dehydratase From Mycobacterium Tuberculosis, Apo Structure
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2025-06-18 Classification: LYASE Ligands: MN |
 |
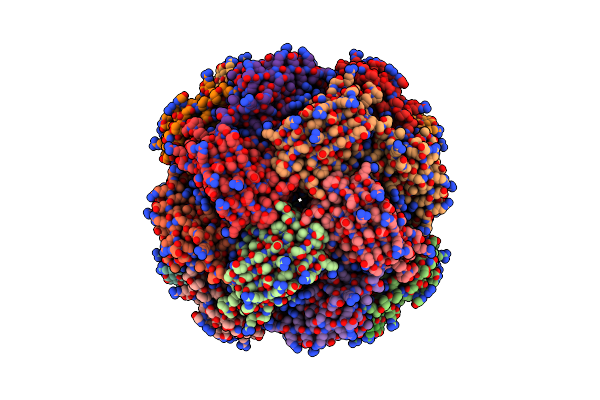
Imidazole Glycerol Phosphate Dehydratase From Mycobacterium Tuberculosis, In Complex With Aminotriazole
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2025-06-18 Classification: LYASE Ligands: MN, 3TR |
 |
Imidazole Glycerol Phosphate Dehydratase From Mycobacterium Tuberculosis, Apo Structure
Organism: Mycobacterium tuberculosis
Method: ELECTRON MICROSCOPY Resolution:2.20 Å Release Date: 2025-06-18 Classification: LYASE Ligands: MN |
 |
Organism: Oscillatoria sp. n9dm
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-04-16 Classification: PHOTOSYNTHESIS Ligands: CYC, GOL |
 |
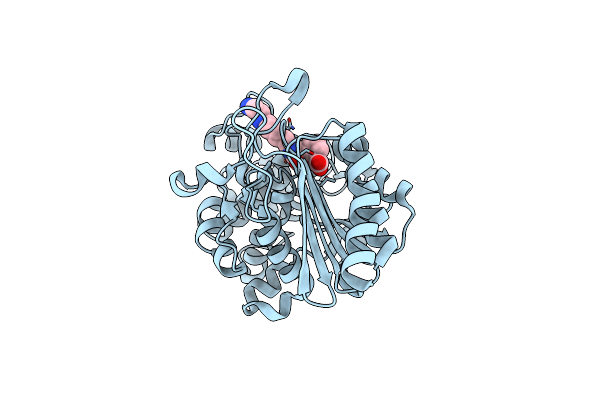
Structure Of Phosphopantetheine Adenylyltransferase (Ppat) From Enterobacter Spp. With The 17-Mer Expression Tag Bound In The Substrate Binding Site Of A Neighbouring Molecule At 2.10 A Resolution.
Organism: Enterobacter sp. 638
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-04-16 Classification: TRANSFERASE Ligands: CIT |
 |
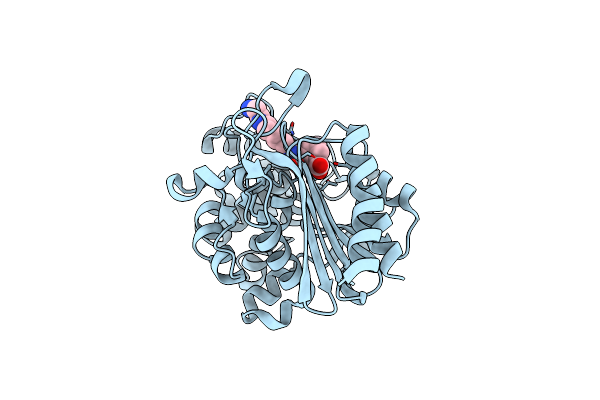
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-03-19 Classification: HYDROLASE/ANTIBIOTIC Ligands: KJK |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-03-19 Classification: HYDROLASE/ANTIBIOTIC Ligands: KJK |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-03-19 Classification: HYDROLASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2025-02-05 Classification: TRANSFERASE Ligands: A1BLX |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2025-01-15 Classification: VIRAL PROTEIN Ligands: A1AV3, DMS |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-01-15 Classification: VIRAL PROTEIN Ligands: A1AV7 |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2025-01-15 Classification: VIRAL PROTEIN Ligands: TLA, NA, A1AV5, A1AV6 |

