Search Count: 47
All
Selected
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: PROTEIN FIBRIL |
 |
Cryo-Em Structure Of The Zeaxanthin-Bound Light-Driven Proton Pumping Rhodopsin, Nm-R1
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.46 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, R16, D12, K3I |
 |
Cryo-Em Structure Of The Myxol-Bound Light-Driven Proton Pumping Rhodopsin, Nm-R1
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.27 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, R16, D12, A1L4O |
 |
Cryo-Em Structure Of The Myxol-Bound Light-Driven Chloride Ion-Pumping Rhodopsin, Nm-R3
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.48 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, A1L4O, CL, PC1, PLC, R16, 8K6, D12, C14, D10 |
 |
Cryo-Em Structure Of The Light-Driven Chloride Ion-Pumping Rhodopsin, Nm-R3
Organism: Nonlabens marinus s1-08
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2025-07-30 Classification: MEMBRANE PROTEIN Ligands: RET, CL, PC1, PLC, D12, R16, 8K6, C14 |
 |
Organism: Zaire ebolavirus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-01-29 Classification: VIRAL PROTEIN/RNA |
 |
The Crystal Structure Of Cyanorhodopsin-Ii (Cyr-Ii) P7104R From Nodosilinea Nodulosa Pcc 7104
Organism: Nodosilinea nodulosa pcc 7104
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2023-10-25 Classification: MEMBRANE PROTEIN Ligands: RET, PG4, HEX, OCT, C14, R16, SO4, CL |
 |
Organism: Lloviu cuevavirus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-04-19 Classification: VIRAL PROTEIN |
 |
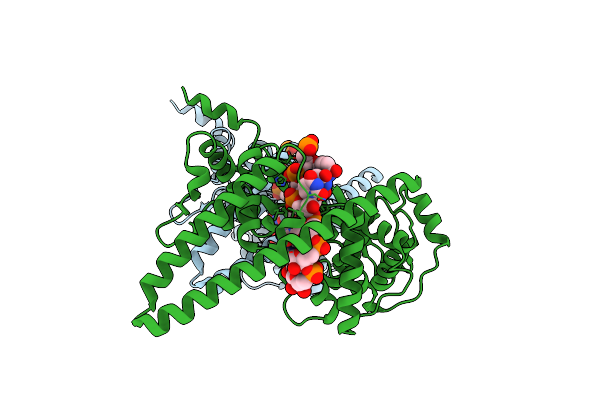
Organism: Lloviu cuevavirus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-04-19 Classification: VIRAL PROTEIN |
 |
Organism: Lake victoria marburgvirus (strain angola/2005), Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-03-09 Classification: VIRAL PROTEIN |
 |
Organism: Escherichia coli (strain k12)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-04-14 Classification: LYASE Ligands: EDO |
 |
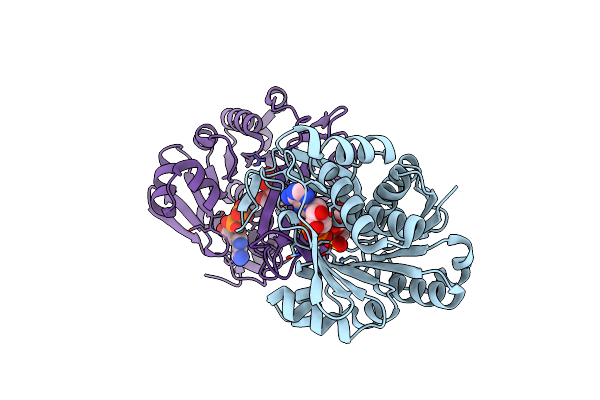
Crystal Structure Of Active Site Mutant Of Sq Isomerase (Yihs-H248A) From Salmonella Enterica In Complex With Sulfofructose (Sf)
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2021-04-14 Classification: ISOMERASE Ligands: RB8 |
 |
Crystal Structure Of Sf Kinase Yihv From E. Coli In Complex With Sulfofructose (Sf), Adp-Mg
Organism: Escherichia coli (strain k12)
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2021-04-14 Classification: TRANSFERASE Ligands: ADP, RB8, MG |
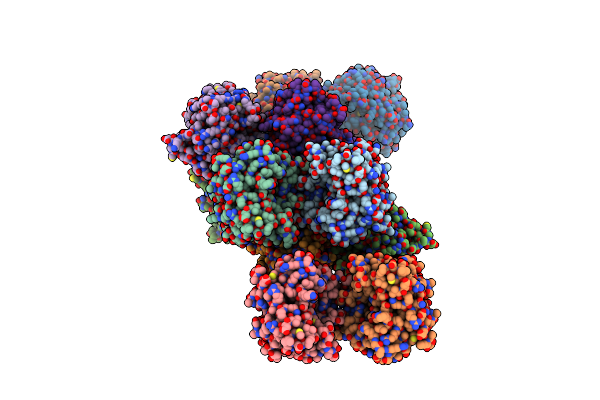 |
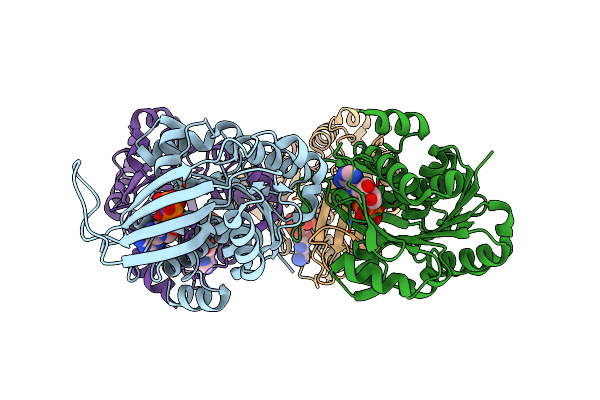
Crystal Structure Of Sfp Aldolase Yiht From Salmonella Enterica In Complex With Sulfate Bound At The Active Site
Organism: Salmonella typhimurium (strain lt2 / sgsc1412 / atcc 700720)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2021-04-14 Classification: LYASE Ligands: SO4 |
 |
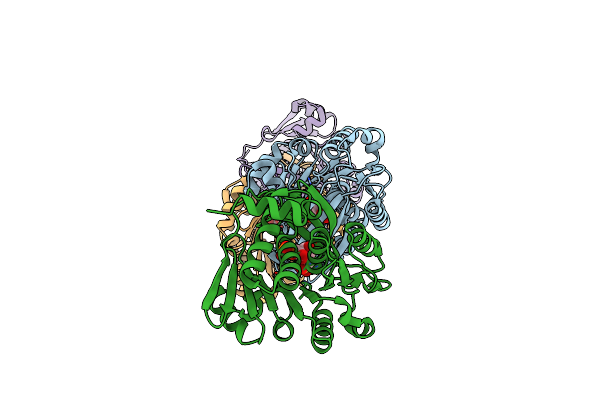
Organism: Escherichia coli (strain k12)
Method: X-RAY DIFFRACTION Resolution:2.93 Å Release Date: 2021-04-14 Classification: TRANSFERASE Ligands: ANP, MG |
 |
Crystal Structure Of E. Coli Sf Kinase (Yihv) In Complex With Product Sulfofructose Phosphate (Sfp)
Organism: Escherichia coli (strain k12)
Method: X-RAY DIFFRACTION Resolution:2.97 Å Release Date: 2021-04-14 Classification: TRANSFERASE Ligands: RAH |
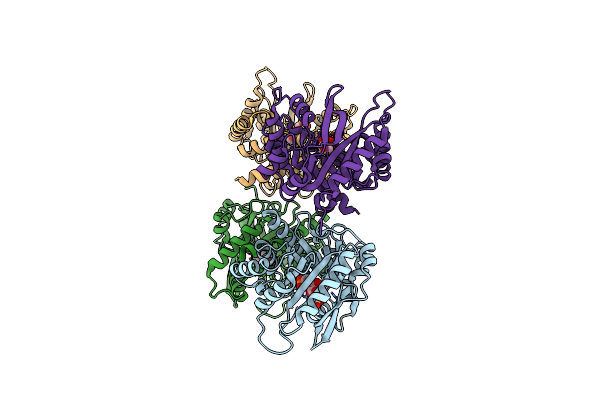 |
Crystal Structure Of Class I Sfp Aldolase Yiht From Salmonella Enterica With Sfp/ Dhap (Schiff Base Complex With Active Site Lys193)
Organism: Salmonella enterica
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2021-04-14 Classification: LYASE Ligands: U8W, U8Z |
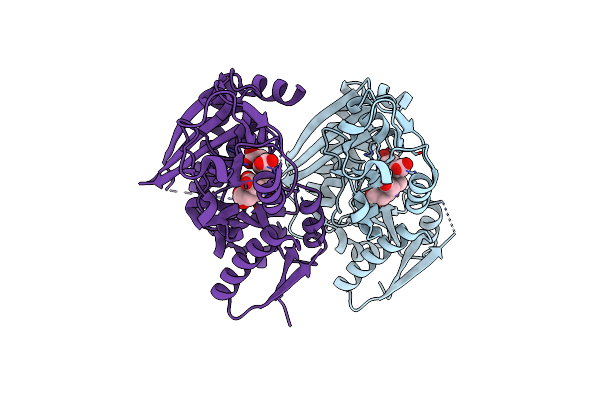 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2021-03-17 Classification: TRANSFERASE Ligands: DPM |
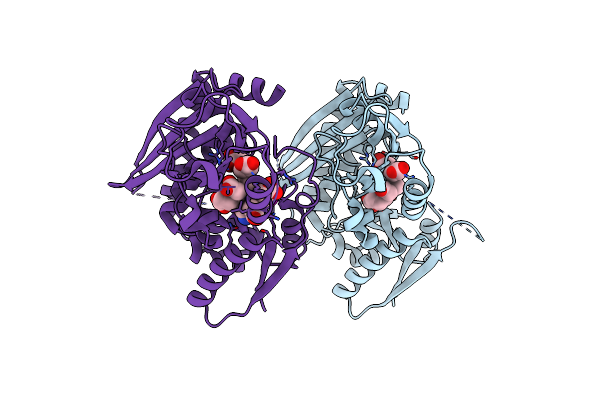 |
Crystal Structure Of The 2-Iodoporphobilinogen-Bound Holo Form Of Human Hydroxymethylbilane Synthase
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2021-03-17 Classification: TRANSFERASE Ligands: DPM, FWL |
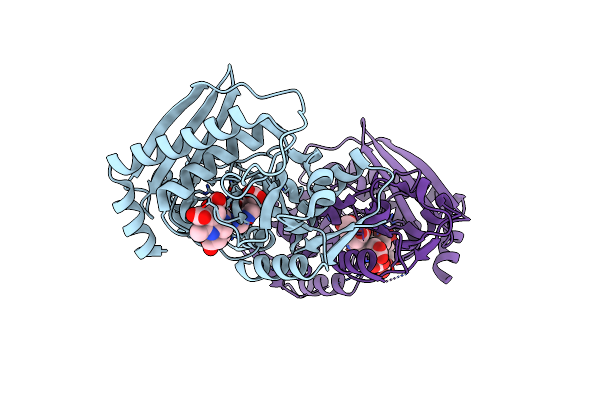 |
Crystal Structure Of The Es2 Intermediate Form Of Human Hydroxymethylbilane Synthase
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2021-03-17 Classification: TRANSFERASE Ligands: 7J8 |

