Search Count: 84
All
Selected
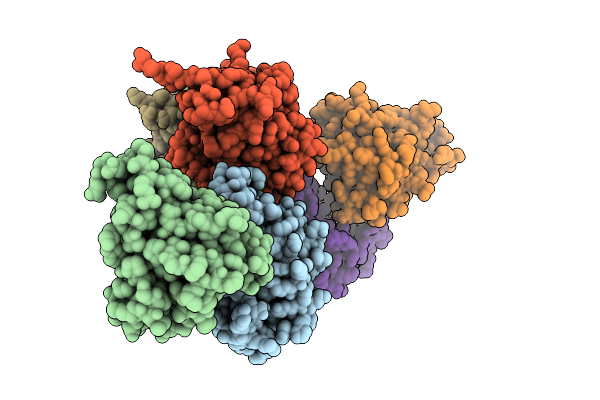 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: GSP, MG |
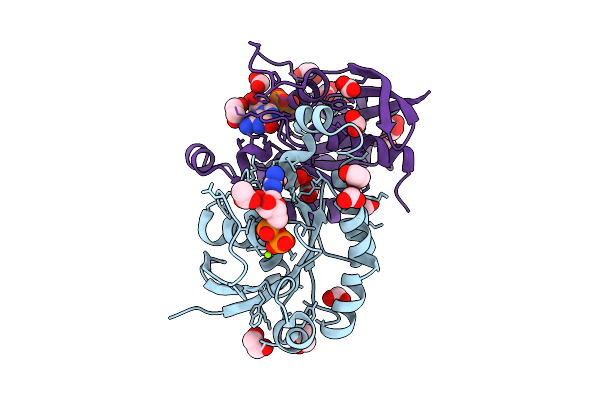 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: GDP, MG, FMT, ACT, EDO |
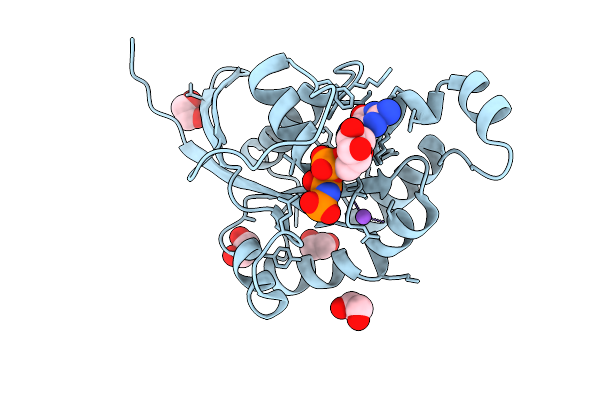 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: GNP, EDO, NA, F |
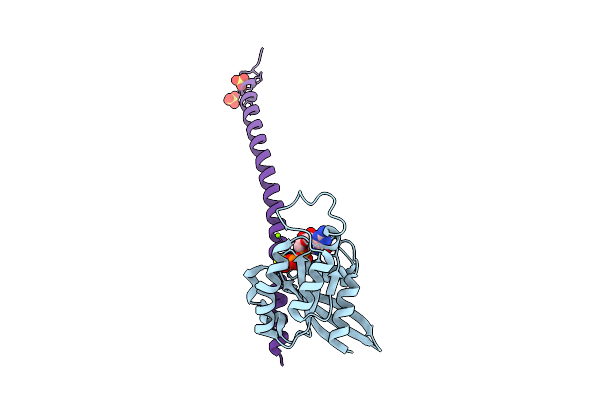 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: MG, GSP, SO4 |
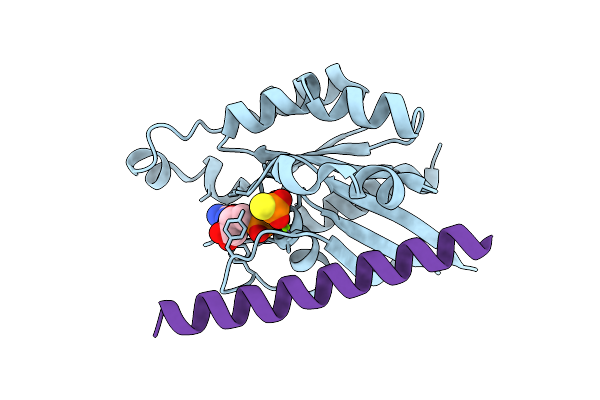 |
Organism: Hordeum vulgare
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-03-18 Classification: PLANT PROTEIN Ligands: GSP, MG |
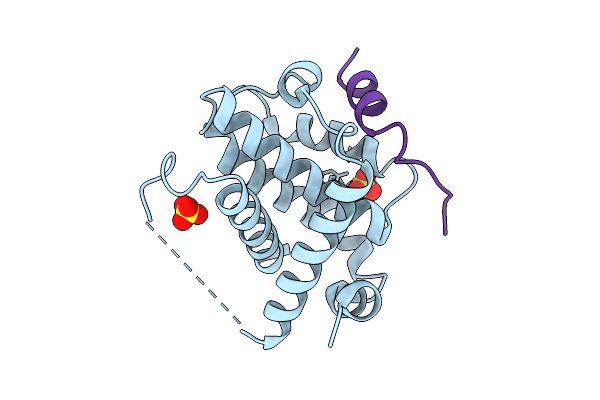 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-09-17 Classification: APOPTOSIS Ligands: SO4 |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-07-02 Classification: HYDROLASE |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL |
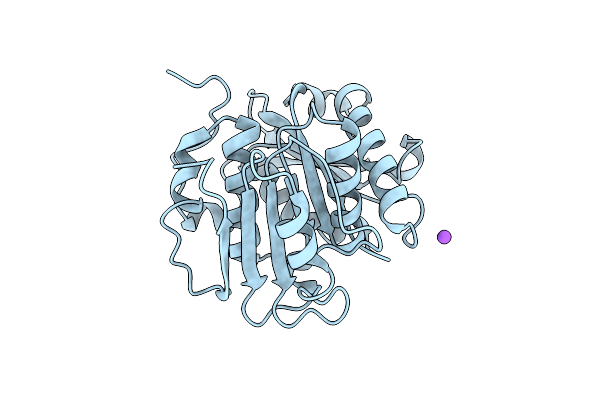 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: CL, NA |
 |
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: EDO, NA |
 |
Crystal Structure Of Leaf Branch Compost Cutinase Quintuple Variant Iccg L50Y
Organism: Uncultured bacterium
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2025-07-02 Classification: HYDROLASE Ligands: EDO, CL |
 |
Crystal Structure Of An Nadh-Accepting Ene Reductase Variant Nostocer1-L1,5 (Engineered Loop Swap From Achromobacter Sp. Ja81)
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2025-04-02 Classification: OXIDOREDUCTASE Ligands: EDO, FMN, ACT, NA, CL |
 |
Crystal Structure Of An Nadh-Accepting Ene Reductase Variant Nostocer1-L1,5 Mutant Q204K
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2025-04-02 Classification: OXIDOREDUCTASE Ligands: FMN, EDO, ACT |
 |
Crystal Structure Of An Nadh-Accepting Ene Reductase Variant Nostocer1-L1,5 Mutant Q350K
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2025-04-02 Classification: OXIDOREDUCTASE Ligands: FMN, EDO, ACT, CL |
 |
Crystal Structure Of An Nadh-Accepting Ene Reductase Variant Nostocer1-L1,5 Mutant D352K
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2025-04-02 Classification: OXIDOREDUCTASE Ligands: FMN, EDO, CL, NA |
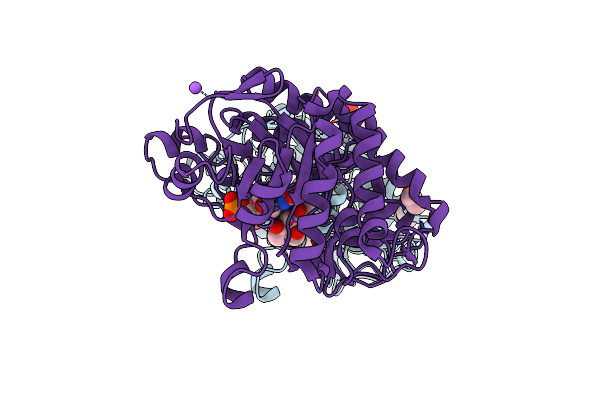 |
Crystal Structure Of An Nadh-Accepting Ene Reductase Variant Nostocer1-L1,5 Mutant T354K
Organism: Nostoc sp. pcc 7120 = fachb-418
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-04-02 Classification: OXIDOREDUCTASE Ligands: FMN, EDO, A1I6V, NA, CL |
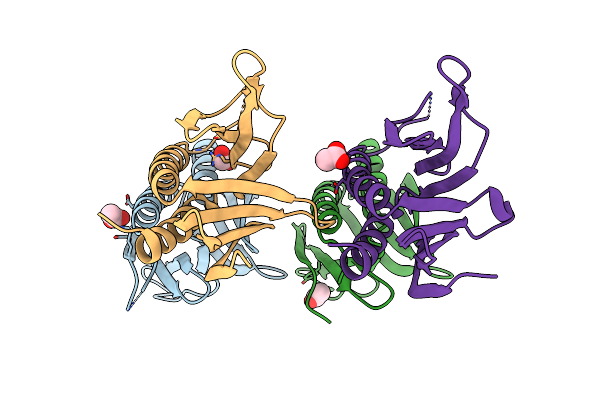 |
Crystal Structure Of Human Pura (Fragment Glu57-Glu212, Pur Repeat I And Ii)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-02-21 Classification: RNA BINDING PROTEIN Ligands: ACT, EDO |
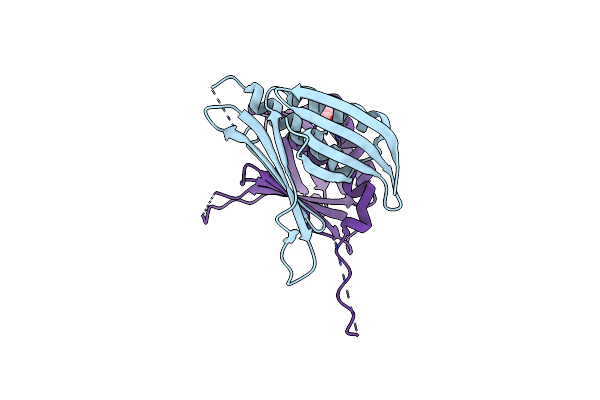 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2024-02-21 Classification: RNA BINDING PROTEIN Ligands: EDO, MG |
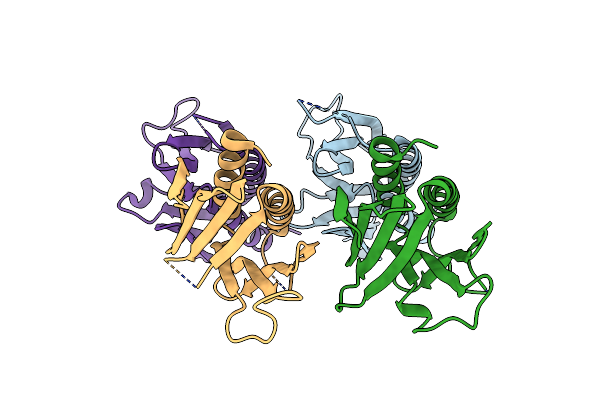 |
Crystal Structure Of Human Pura (Fragment Glu57-Glu212, Pur Repeat I And Ii) R140P Mutant
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2024-02-21 Classification: RNA BINDING PROTEIN |

