Search Count: 1,888
All
Selected
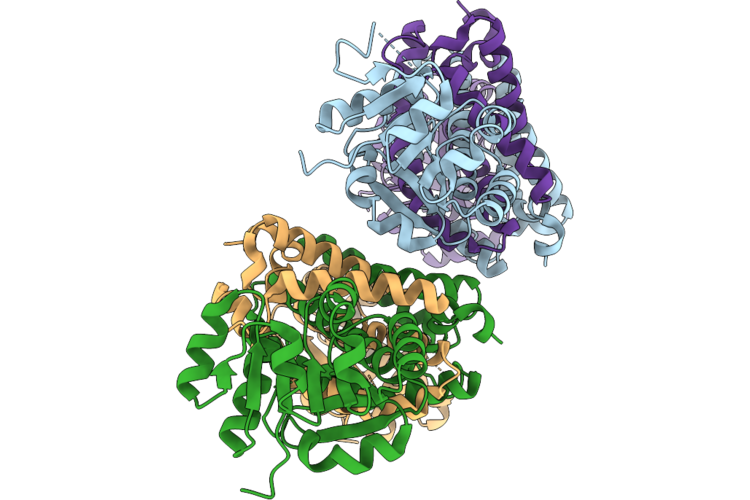 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE |
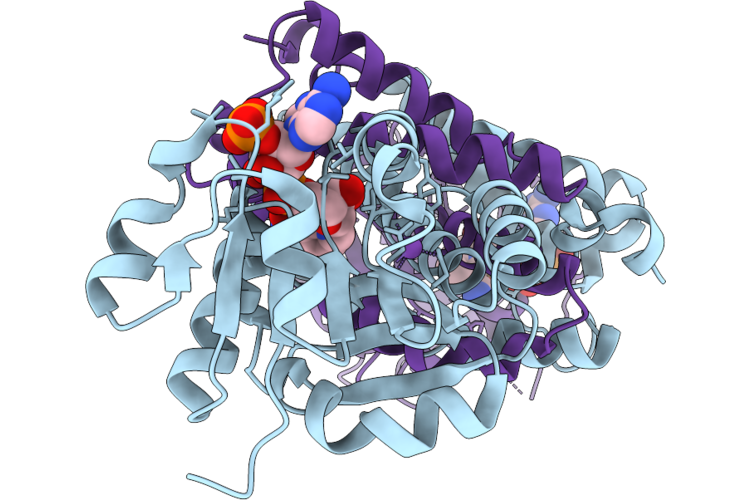 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: NAP, NA |
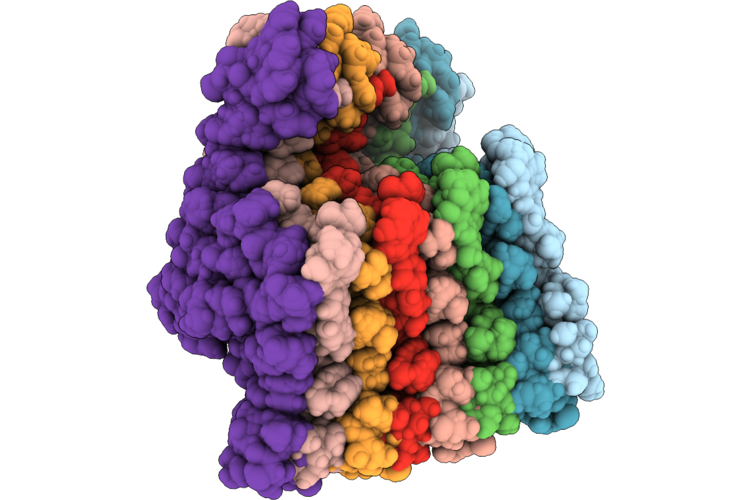 |
Ssnmr Structure Of Anti-Necroptosis Viral:Human Functional Hetero-Amyloid M45:Ripk3
Organism: Homo sapiens, Murid betaherpesvirus 1
Method: SOLID-STATE NMR Release Date: 2026-04-22 Classification: PROTEIN FIBRIL |
 |
Organism: Haloarcula taiwanensis
Method: ELECTRON MICROSCOPY Resolution:4.90 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: RET |
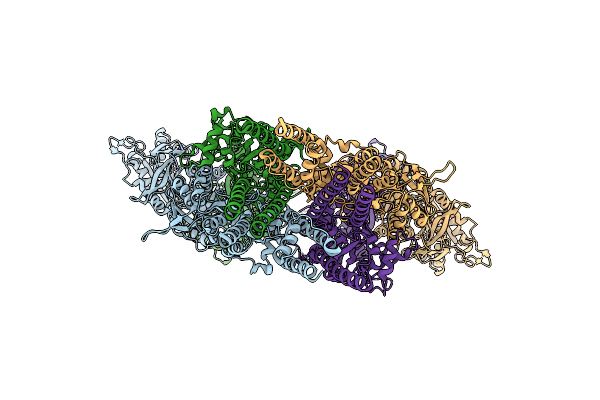 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
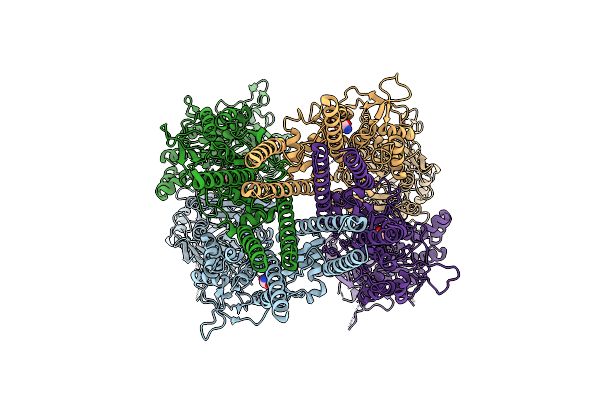 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: DSN |
 |
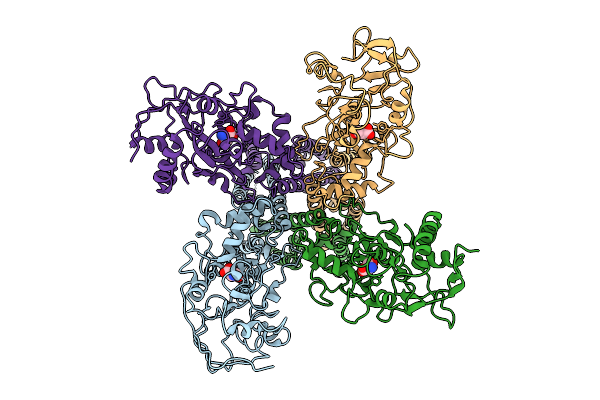
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
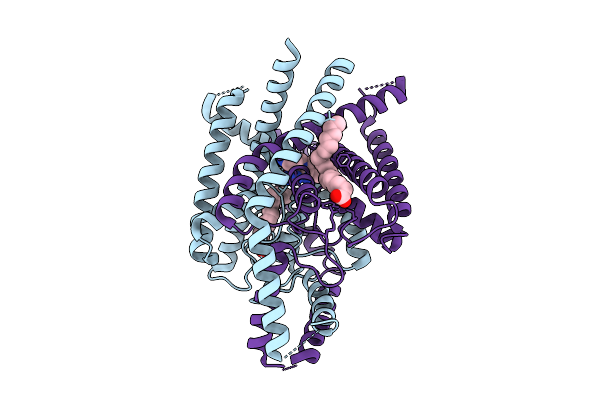
Structure Of Atd Truncated Glutamate Receptor Mglud1 Complexed With D-Serine
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: DSN |
 |
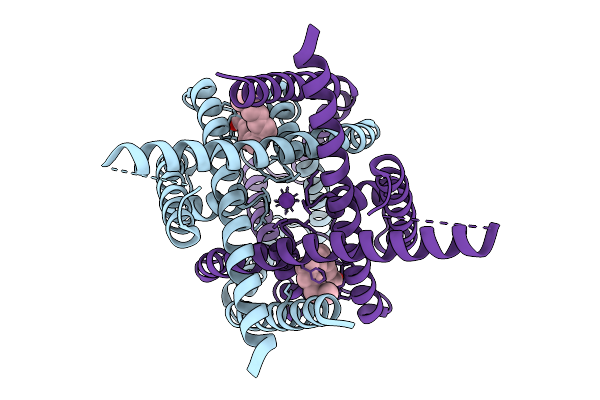
Structure Of Atd Truncated Glutamate Receptor Mglud1 Complexed With Gaba And Calcium
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: ACD, A1ENQ |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: A1MBI |
 |
Organism: Homo sapiens, Severe acute respiratory syndrome coronavirus 2, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: ACD, K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: HLT |
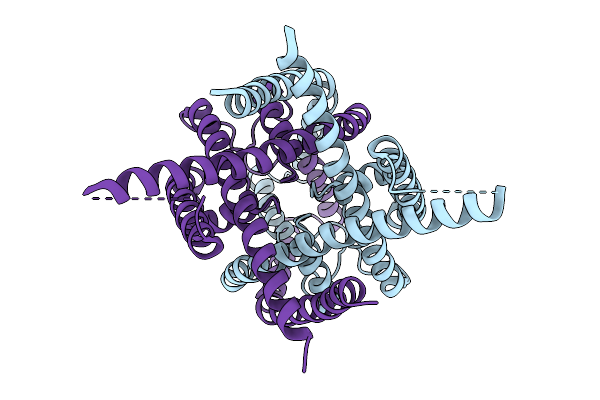 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
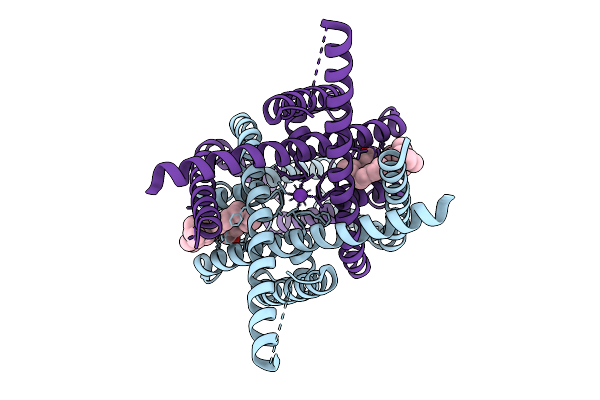 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: ACD, CLR, K |
 |
Organism: Severe acute respiratory syndrome coronavirus 2, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.33 Å Release Date: 2026-03-18 Classification: VIRAL PROTEIN |
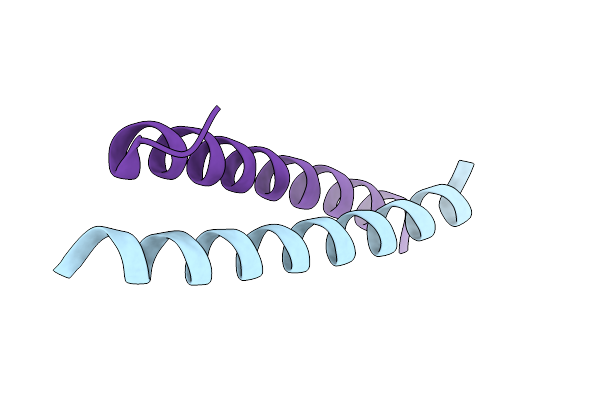 |
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-03-18 Classification: TRANSCRIPTION |
 |
Backbone Modification In The Gcn4 Leucine Zipper: Calpha-Methyl-Glu At Position 11
Organism: Saccharomyces cerevisiae
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: TRANSCRIPTION |

