Search Count: 119
All
Selected
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
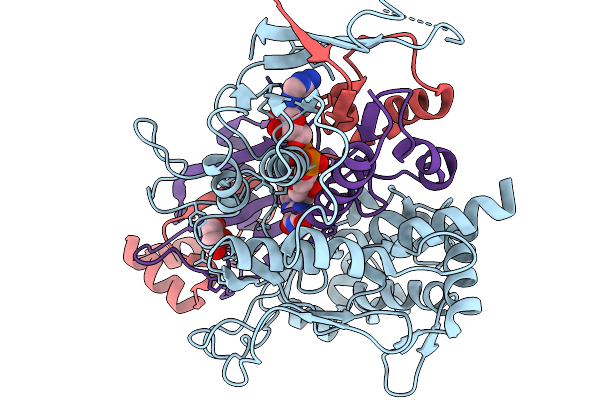 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
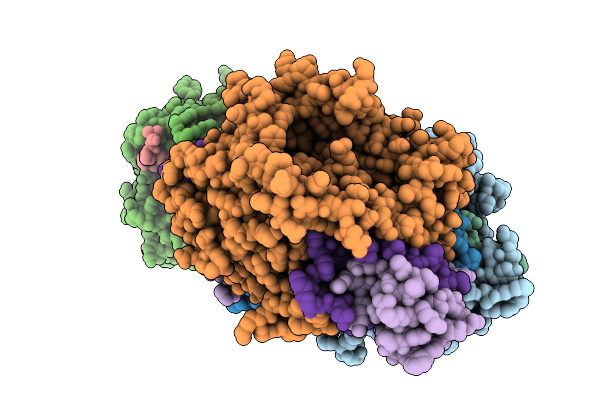 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
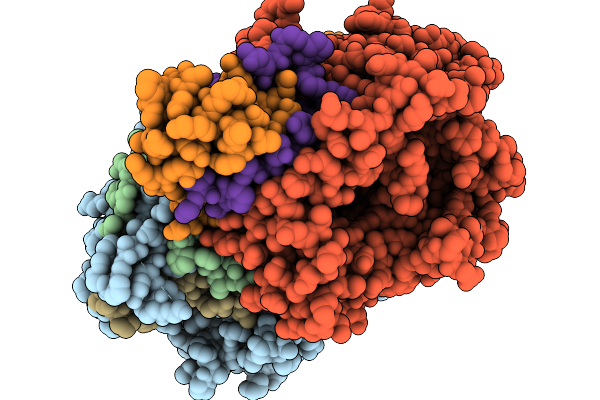 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:3.27 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
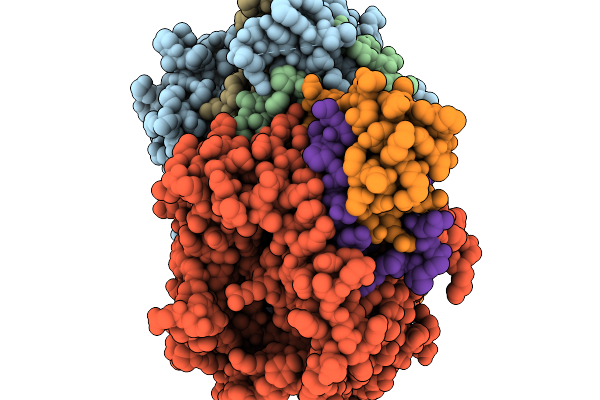 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
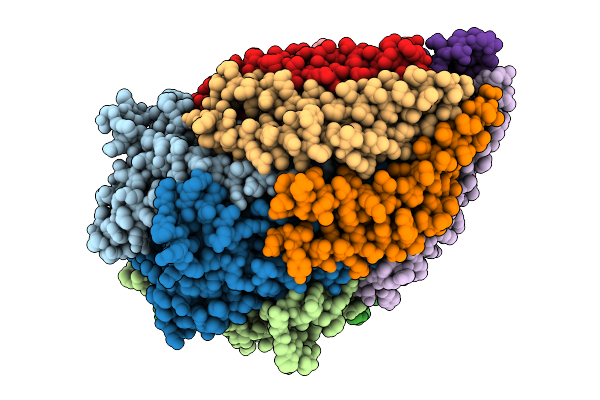 |
Structure Of The Salmonella Flagellar Flipqr Complex Reconstituted In The Peptidisc
Organism: Salmonella enterica subsp. enterica serovar typhimurium str. lt2
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: PROTEIN TRANSPORT |
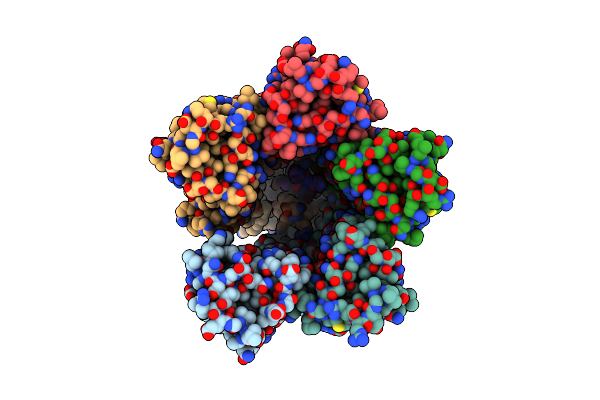 |
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 1
Organism: Paenibacillus sp. tca20, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN Ligands: AV0 |
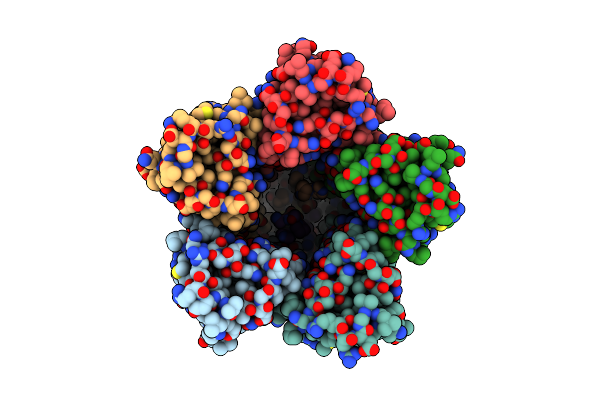 |
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 2
Organism: Paenibacillus sp. tca20, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN |
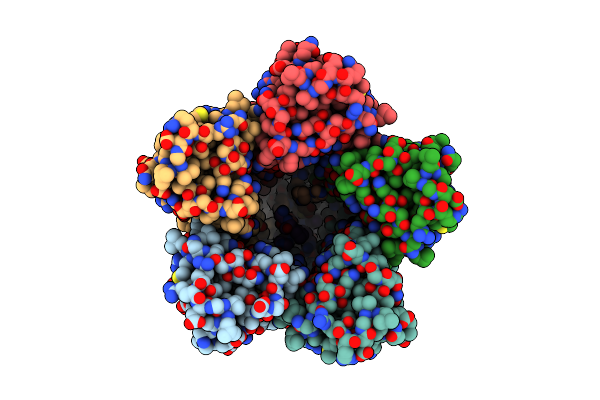 |
The Chimeric Flagellar Motor Complex Between Mota1B1 From Paenibacillus Sp. Tca20 And Motab From E.Coli, State 3
Organism: Paenibacillus sp. tca20
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2025-04-09 Classification: MOTOR PROTEIN |
 |
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-02-26 Classification: MOTOR PROTEIN Ligands: SO4 |
 |
Structure Of The Periplasmic Domain Of Mots From Bacillus Subtilis In 300 Mm Nacl
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-02-26 Classification: MOTOR PROTEIN |
 |
Structure Of The Periplasmic Domain Of Mots From Bacillus Subtilis In 300 Mm Kcl
Organism: Bacillus subtilis
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-02-26 Classification: MOTOR PROTEIN |
 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MOTOR PROTEIN |
 |
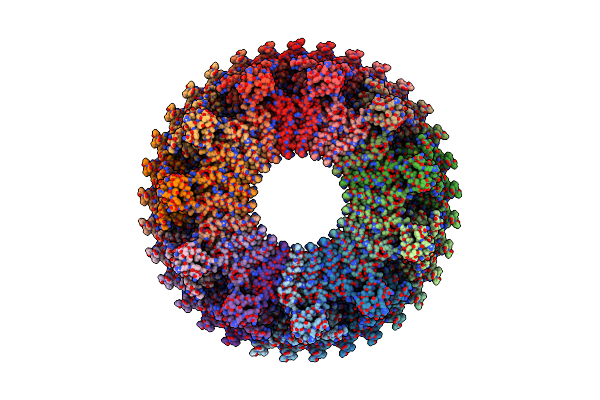
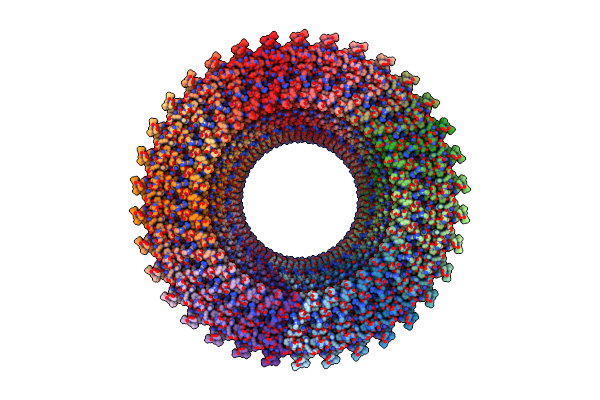
Structure Of The Rbm3 Ring Of Salmonella Flagellar Ms-Ring Protein Flif With C33 Symmetry Applied
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MOTOR PROTEIN |
 |
Structure Of The Rbm3 Ring Of Salmonella Flagellar Ms-Ring Protein Flif With C34 Symmetry Applied
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MOTOR PROTEIN |
 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MEMBRANE PROTEIN |
 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MEMBRANE PROTEIN |
 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MEMBRANE PROTEIN |
 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MEMBRANE PROTEIN |
 |
Bacterial Flagellar Sodium-Driven Stator Poma5Pomb2 With 100 Mm Nacl And 0.1 Mm Phenamil
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-01-01 Classification: MEMBRANE PROTEIN |

