Search Count: 13,498
All
Selected
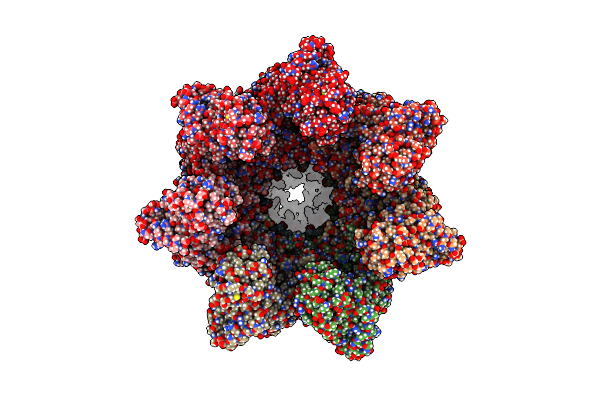 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-12-27 Classification: CHAPERONE Ligands: AF3, ADP, MG, K |
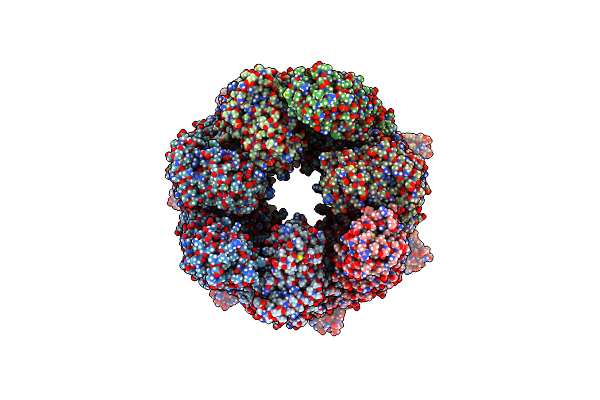 |
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2023-12-27 Classification: CHAPERONE Ligands: BEF, ADP, MG, K |
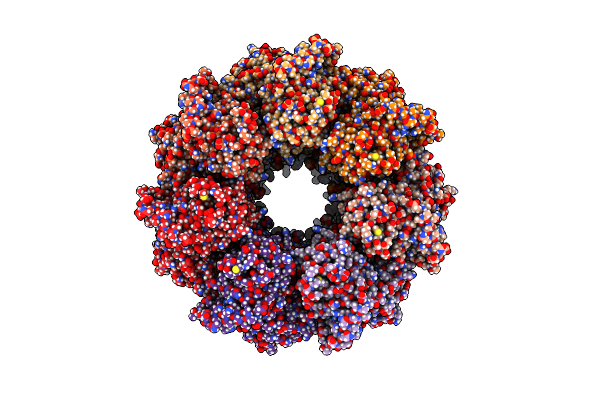 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: CHAPERONE |
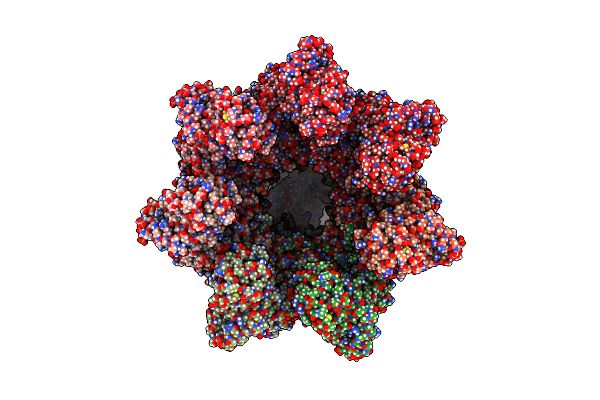 |
Organism: Escherichia coli (strain k12), Rhodospirillum rubrum (strain atcc 11170 / ath 1.1.1 / dsm 467 / lmg 4362 / ncimb 8255 / s1)
Method: ELECTRON MICROSCOPY Release Date: 2025-02-12 Classification: CHAPERONE Ligands: AF3, MG, ADP, K |
 |
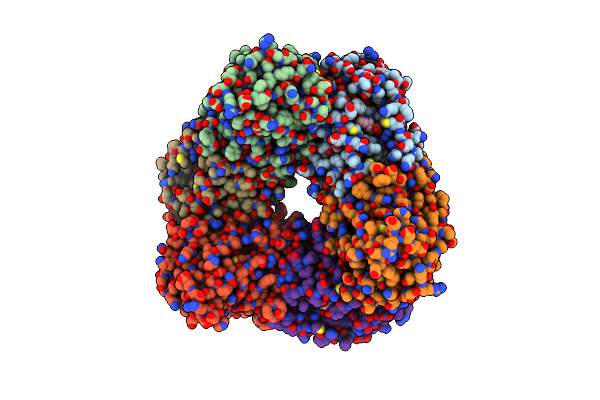
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp And Dsdna From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:2.30 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: ANP, MN |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 3)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN, ADP |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 1)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: TRANSLOCASE Ligands: ANP |
 |
The Cryo-Em Structure Of Hera-Nura Complex With Atpgammas And Dsdna From Thermococcus Kodakarensis (State 2)
Organism: Thermococcus kodakarensis, Dna molecule
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: AGS, MN, ADP |
 |
Crystal Structure Of Lactobacillus Rhamnosus 4-Deoxy-L-Threo-5-Hexosulose-Uronate Ketol-Isomerase Kdui Complexed With Mops
Organism: Lacticaseibacillus rhamnosus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-08-16 Classification: ISOMERASE Ligands: ZN, MPO |
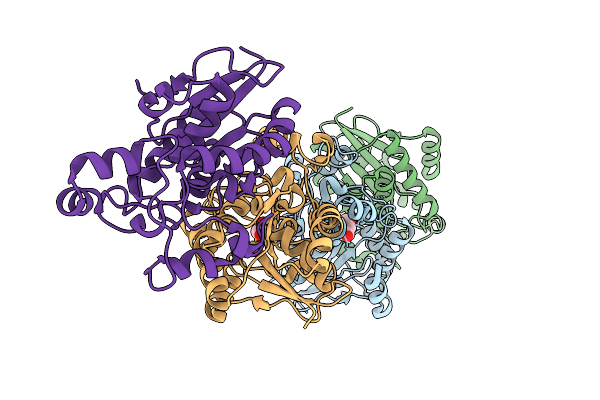 |
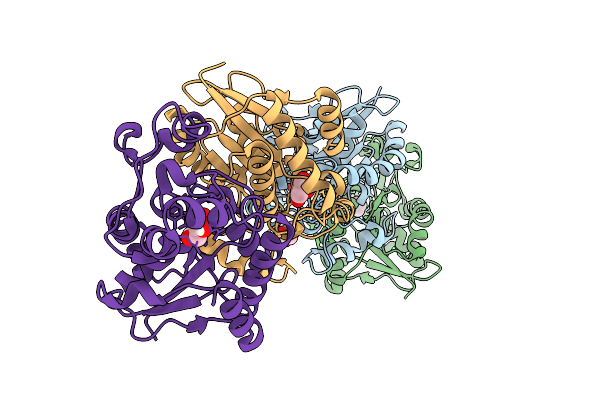
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: PEG, EDO |
 |
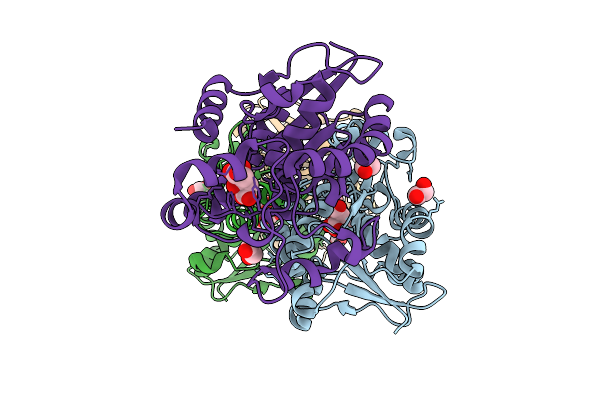
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: GOL, EDO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: CIT, EDO, ACT, GOL |
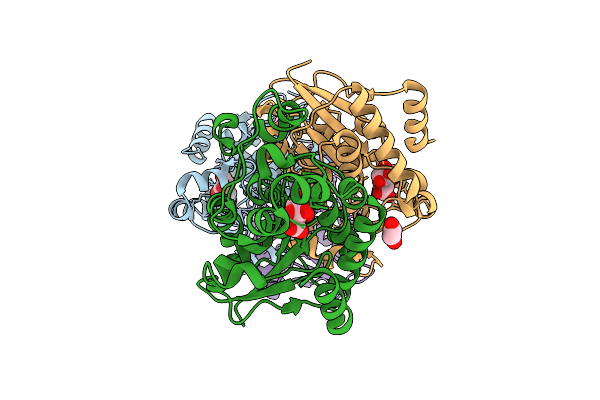 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-18 Classification: ISOMERASE Ligands: CIT, EDO |
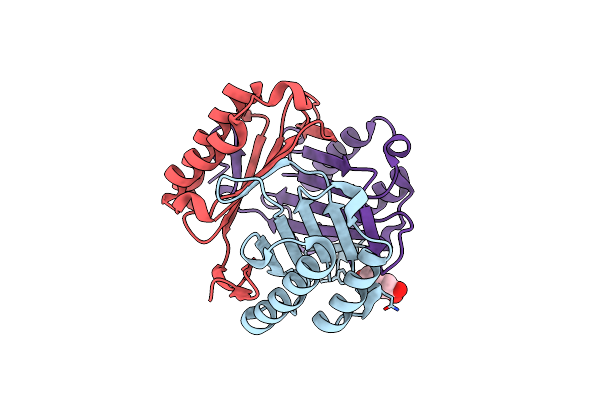 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2025-03-26 Classification: ISOMERASE Ligands: HFC, CL |
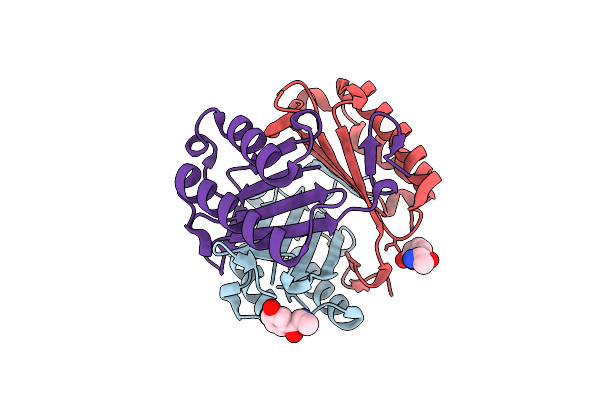 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2025-03-26 Classification: ISOMERASE Ligands: CL, 7Y8, J23 |
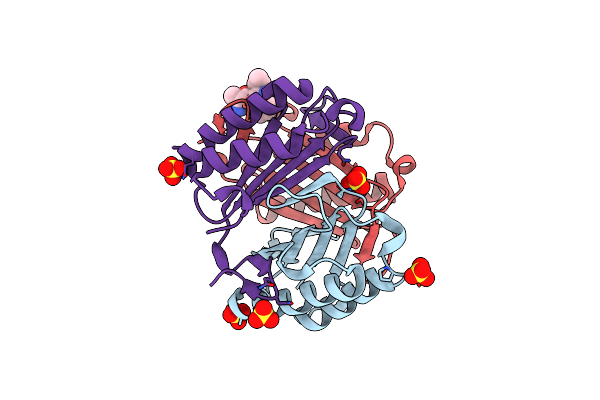 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2013-09-25 Classification: cytokine, Isomerase Ligands: SO4, CL, AVR |
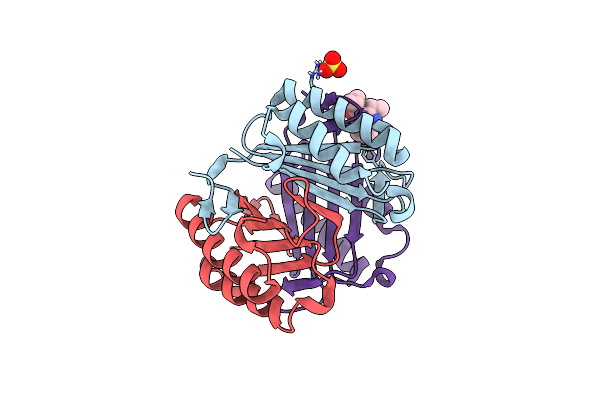 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2013-10-02 Classification: cytokine, Isomerase Ligands: CL, SO4, AVL |
 |
Crystallographic And Biological Characterization Of N- And C- Terminus Mutants Of Human Mif
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2013-10-02 Classification: cytokine, isomerase Ligands: SO4, CL |
 |
Organism: Candida albicans
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: GOL, A1JCW, A1JCV, CL, NA |

