Search Count: 3,301
All
Selected
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: DSN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Structure Of Atd Truncated Glutamate Receptor Mglud1 Complexed With D-Serine
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: DSN |
 |
Structure Of Atd Truncated Glutamate Receptor Mglud1 Complexed With Gaba And Calcium
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
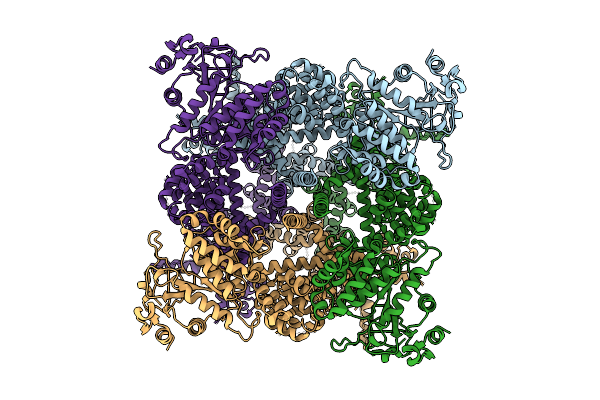
Avian Trpm8 (Parus Major) Desensitized, Fully-Swapped, Ligand-Free Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
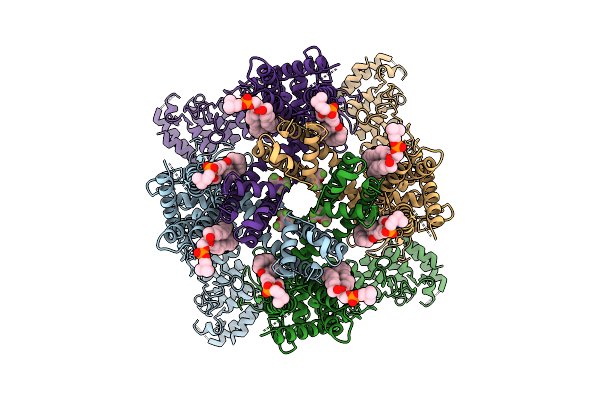
Avian Trpm8 (Parus Major) Menthol Bound Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: XUQ |
 |
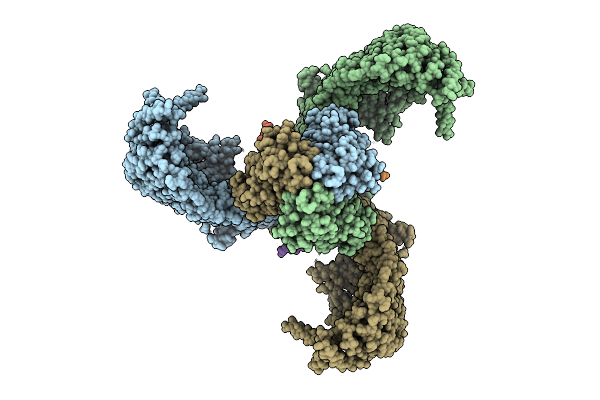
Cryo-Em Structure Of Human Trpv3 In Complex With Sevoflurane Determined In Msp2N2 Nanodisc
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: A1IPI, POV |
 |
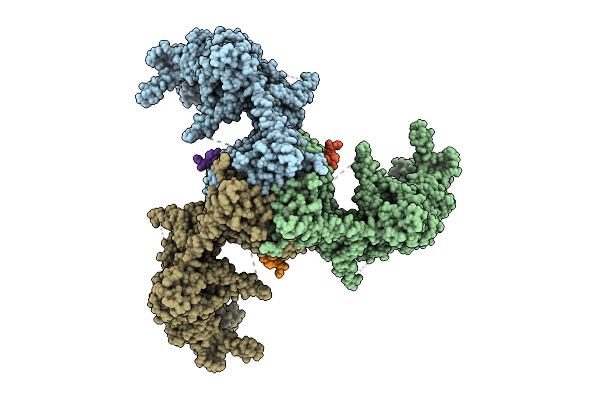
Organism: Mus musculus, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN |
 |
Avian Trpm8 (Parus Major) Semi-Swapped, Calcium Free, Menthol Bound Structure Resolved In Cell Vesicles
Organism: Parus major
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: XUQ |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN |
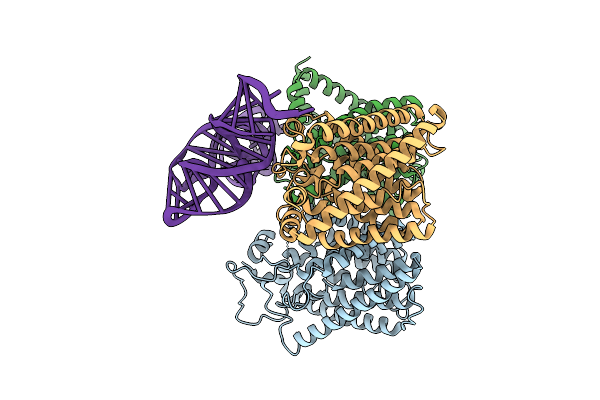 |
Organism: Arabidopsis thaliana, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN/RNA |
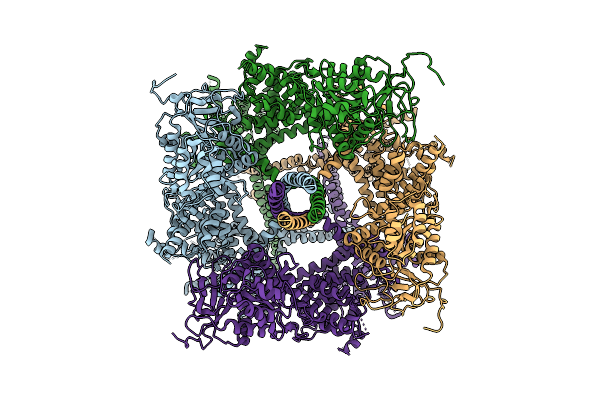 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN |
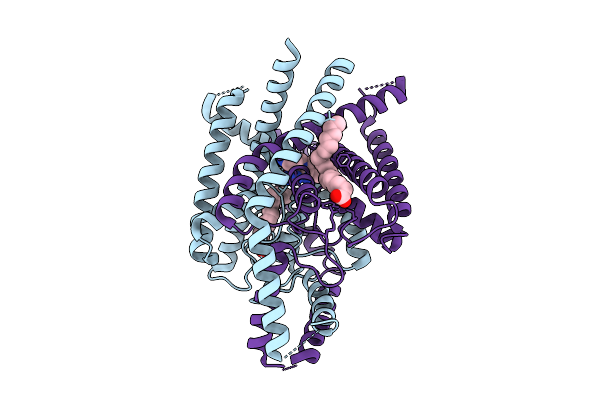 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: ACD, A1ENQ |
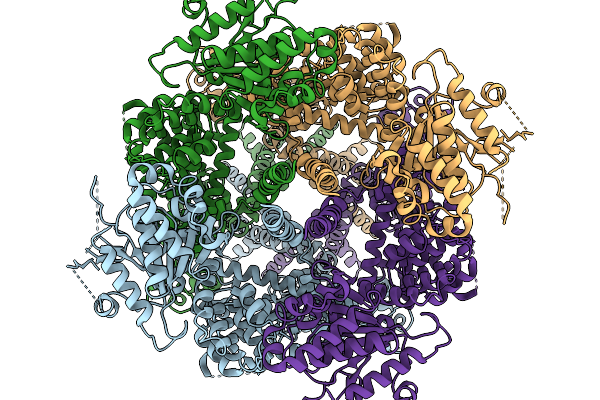 |
Avian Trpm8 (Parus Major) Closed, Ligand-Free Structure Resolved In Cell Vesicles Using Cryo-Em
Organism: Parus major
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-03-25 Classification: MEMBRANE PROTEIN |

