Search Count: 25
All
Selected
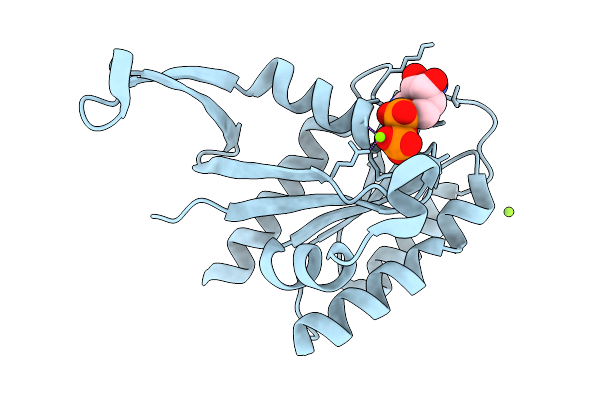 |
Crystal Structure Of Gdp-Bound Kras G12D/I55E: Suppressing G12D Oncogenicity Via Second-Site I55E Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2026-03-25 Classification: ONCOPROTEIN Ligands: GDP, MG, CL |
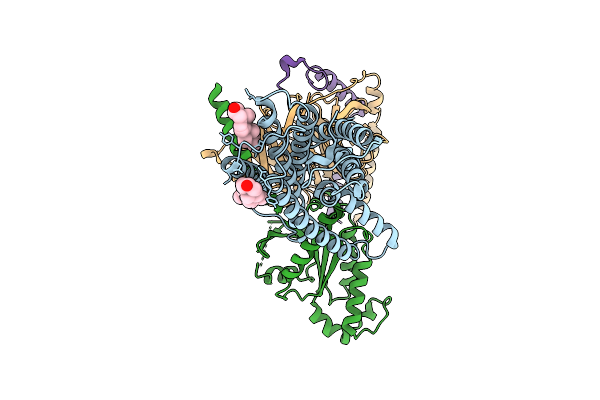 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-12 Classification: SIGNALING PROTEIN Ligands: CLR |
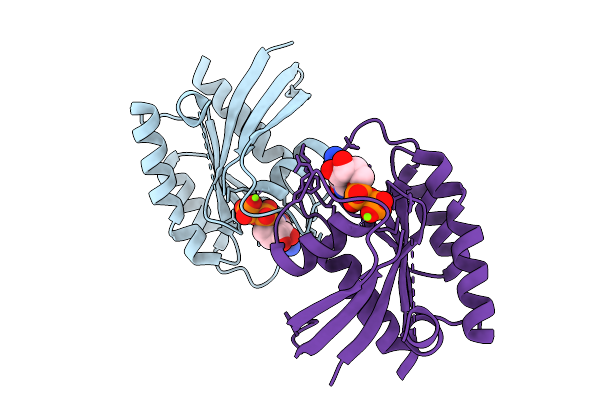 |
Crystal Structure Of Gdp-Bound Kras G12D/M67R: Suppressing G12D Oncogenicity Via Second-Site M67R Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: GDP, MG |
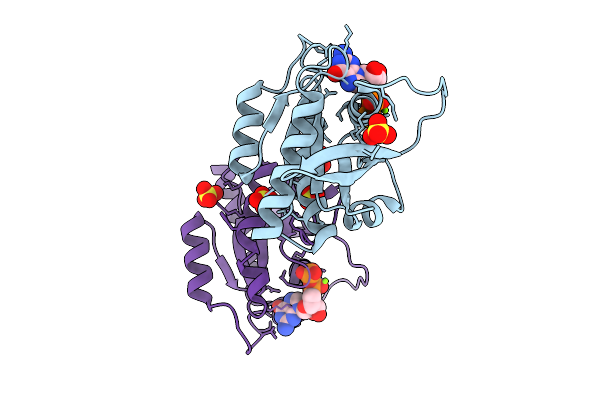 |
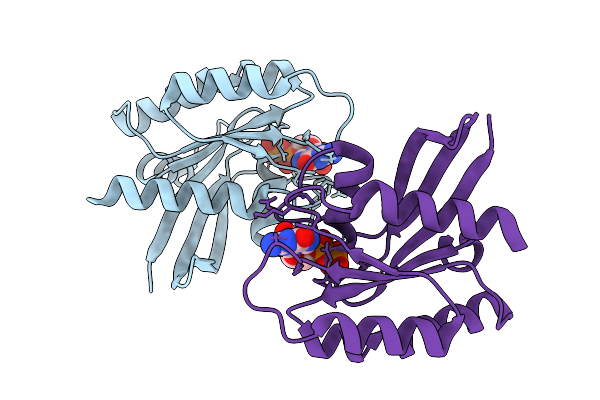
Crystal Structure Of Gdp-Bound Kras G12D/F28K: Suppressing G12D Oncogenicity Via Second-Site F28K Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: GDP, MG, SO4 |
 |
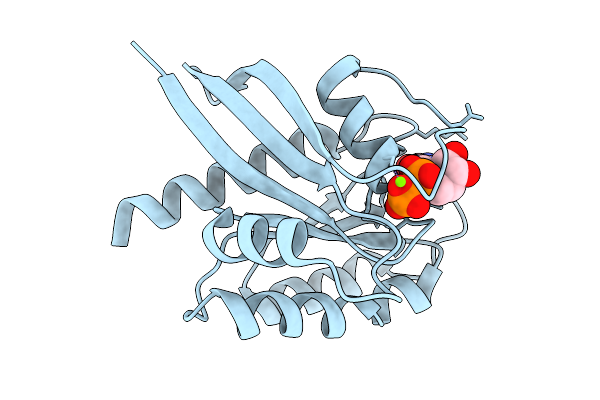
Crystal Structure Of Gdp-Bound Kras G12D/P34R: Suppressing G12D Oncogenicity Via Second-Site P34R Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: MG, GDP |
 |
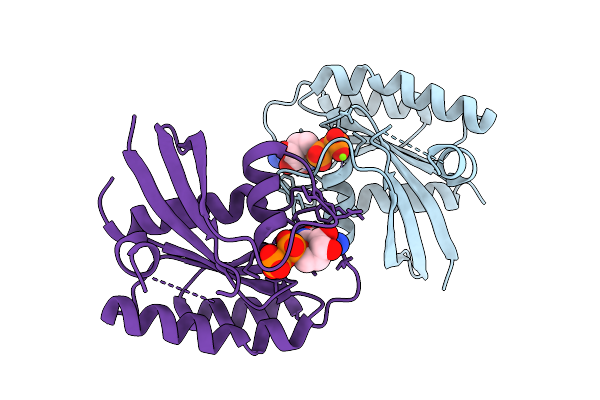
Crystal Structure Of Gdp-Bound Kras G12D/R41Q: Suppressing G12D Oncogenicity Via Second-Site R41Q Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: GDP, MG |
 |
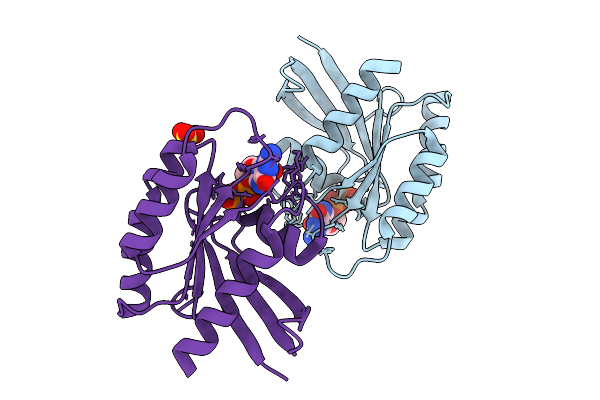
Crystal Structure Of Gdp-Bound Kras G12D/V45E: Suppressing G12D Oncogenicity Via Second-Site V45E Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: GDP, MG |
 |
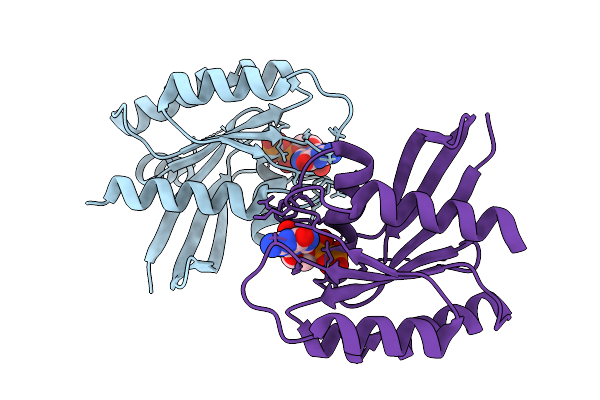
Crystal Structure Of Gdp-Bound Kras G12D/D54R: Suppressing G12D Oncogenicity Via Second-Site D54R Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: GDP, MG, SO4 |
 |
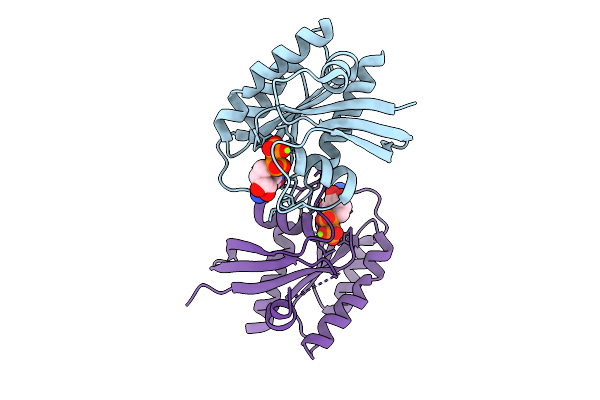
Crystal Structure Of Gdp-Bound Kras G12D/G60R: Suppressing G12D Oncogenicity Via Second-Site G60R Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: MG, GDP |
 |
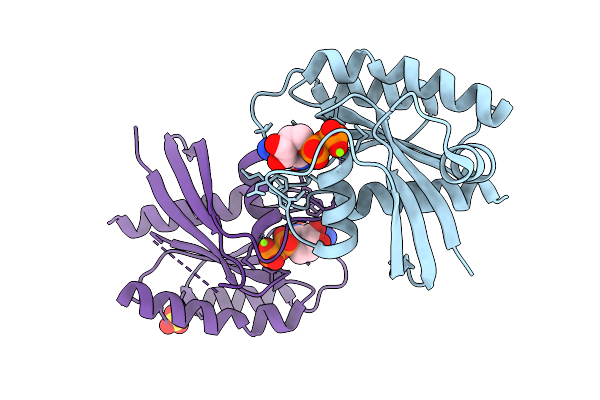
Crystal Structure Of Gdp-Bound Kras G12D/V103Y: Suppressing G12D Oncogenicity Via Second-Site V103Y Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: MG, GDP |
 |
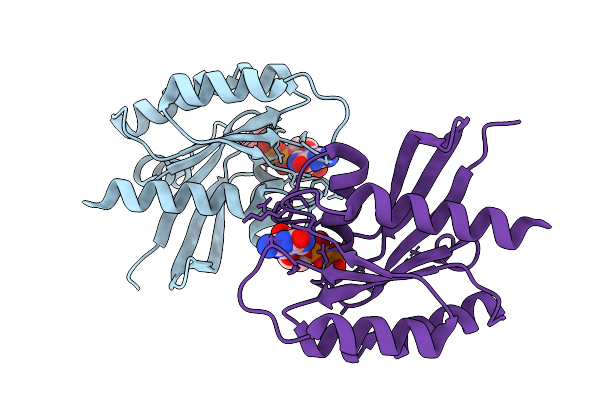
Crystal Structure Of Gdp-Bound Kras G12D/E62Q: Suppressing G12D Oncogenicity Via Second-Site E62Q Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: MG, GDP, SO4 |
 |
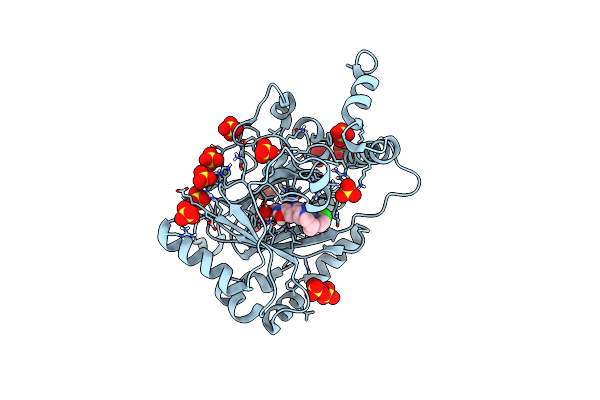
Crystal Structure Of Gdp-Bound Kras E3K/G12D: Suppressing G12D Oncogenicity Via Second-Site E3K Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-10-08 Classification: ONCOPROTEIN Ligands: MG, GDP, GOL |
 |
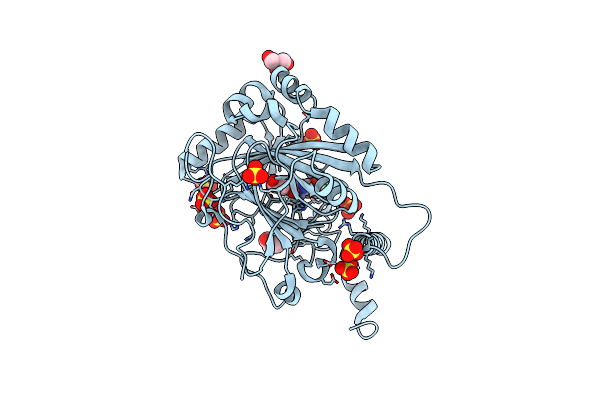
Factor Inhibiting Hif-1 Alpha In Complex With (5-(1-(3-(4-Chlorophenyl)Propyl)-1H-1,2,3-Triazol-4-Yl)-3-Hydroxypicolinoyl)Glycine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2024-02-28 Classification: OXIDOREDUCTASE Ligands: SO4, ZN, P1X |
 |
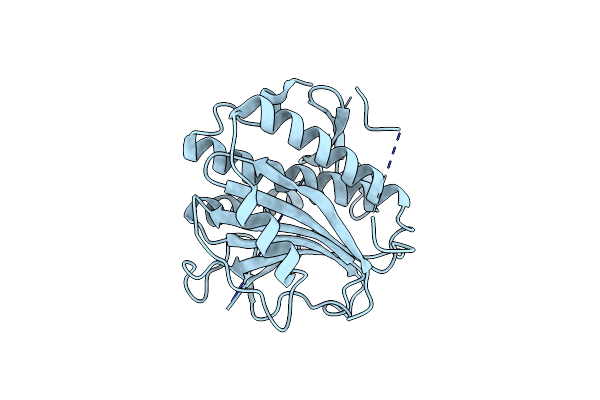
Factor Inhibiting Hif-1 Alpha In Complex With (5-(3-(3-Chlorophenyl)Isoxazol-5-Yl)-3-Hydroxypicolinoyl)Glycine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2024-02-28 Classification: OXIDOREDUCTASE Ligands: P5I, GOL, ZN, SO4 |
 |
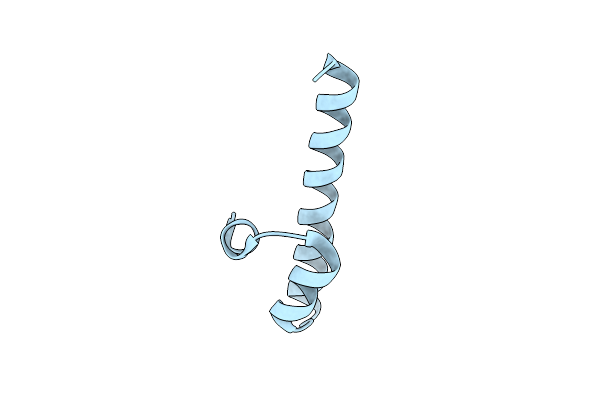
Organism: Caenorhabditis elegans
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2023-05-17 Classification: PEPTIDE BINDING PROTEIN |
 |
Organism: Escherichia coli str. k-12 substr. mg1655
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-03-29 Classification: ANTITOXIN |
 |
Structure Of Apo, Semet-Labeled Cobalamin-Dependent S-Adenosylmethionine Radical Enzyme Oxsb
Organism: Bacillus megaterium
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2017-04-19 Classification: METAL BINDING PROTEIN Ligands: EDO |
 |
Structure Of Cobalamin-Dependent S-Adenosylmethionine Radical Enzyme Oxsb With Aqua-Cobalamin Bound
Organism: Bacillus megaterium
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2017-04-19 Classification: METAL BINDING PROTEIN Ligands: SF4, B12, DTT, MES, EDO |
 |
Structure Of Cobalamin-Dependent S-Adenosylmethionine Radical Enzyme Oxsb With Aqua-Cobalamin And S-Adenosylmethionine Bound
Organism: Bacillus megaterium
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2017-04-19 Classification: METAL BINDING PROTEIN Ligands: SF4, B12, SAM, MES, EDO |
 |
Structure Of The Hd-Domain Phosphohydrolase Oxsa With Oxetanocin-A Diphosphate Bound
Organism: Bacillus megaterium
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2016-11-23 Classification: METAL BINDING PROTEIN Ligands: MG, 7D3 |

