Search Count: 182
All
Selected
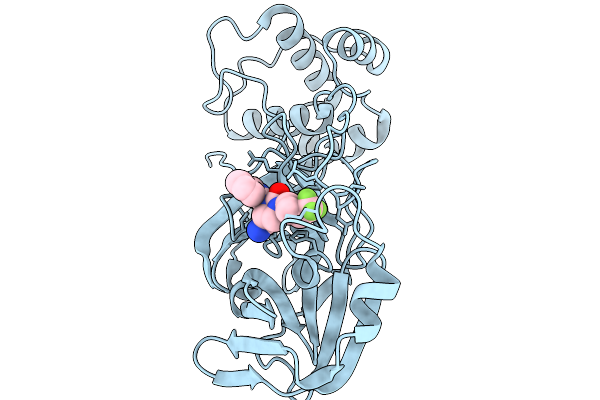 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-04 Classification: HYDROLASE Ligands: A1CAR |
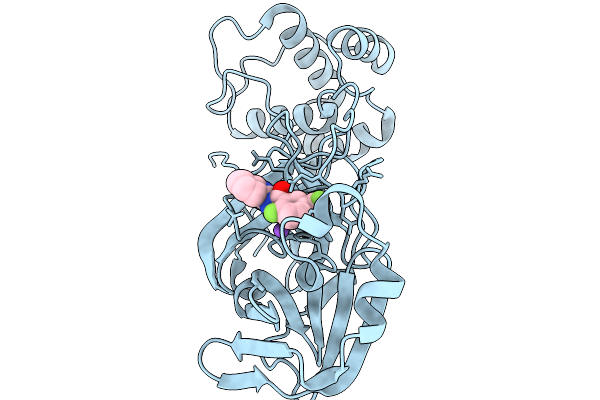 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: A1CAS, NA |
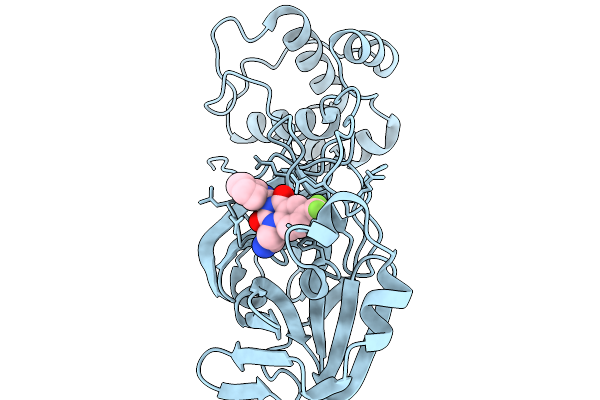 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: A1CBN |
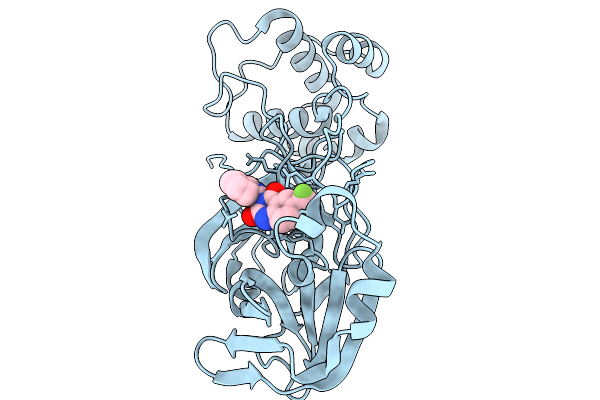 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: A1CBY |
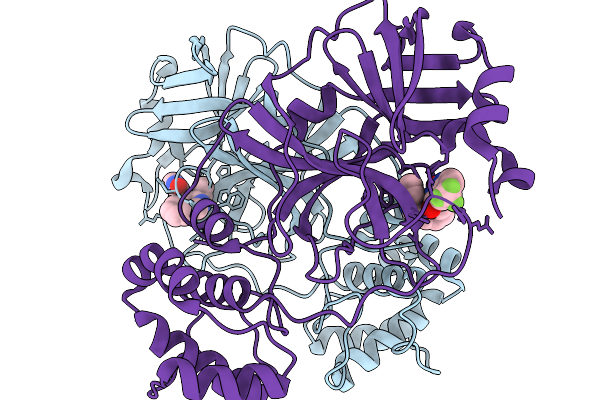 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-02-18 Classification: HYDROLASE Ligands: A1CB2 |
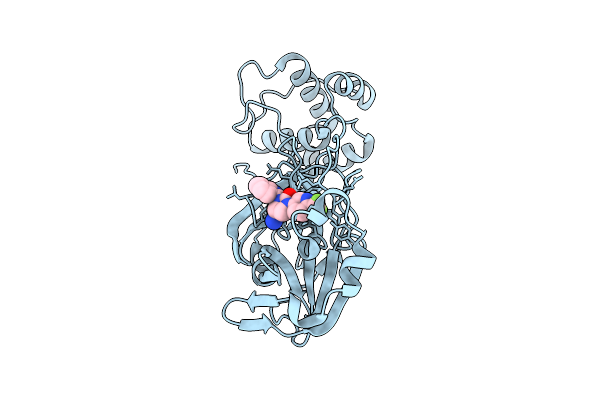 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-12-24 Classification: HYDROLASE Ligands: A1CBU |
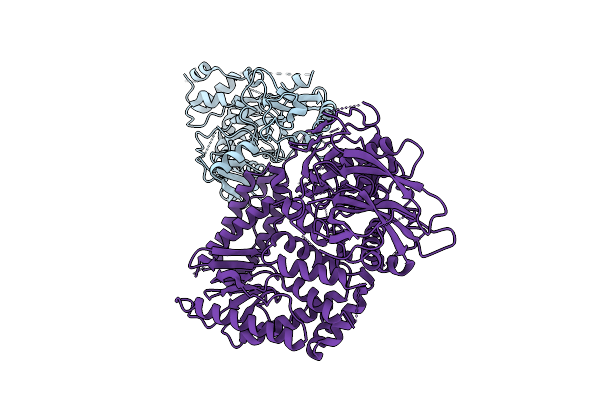 |
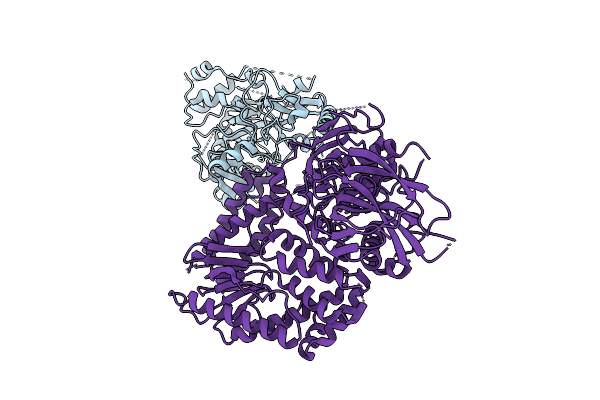
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
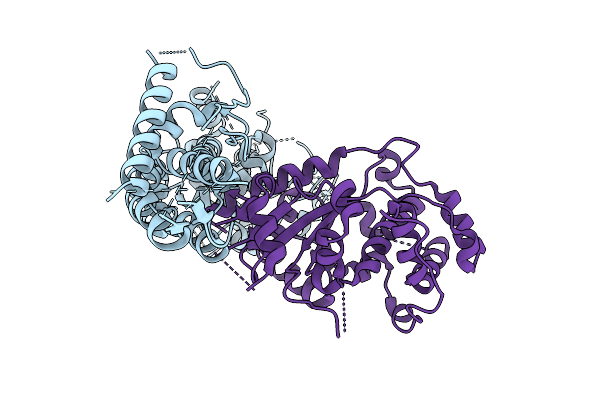 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna And Atp-Gamma-S
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
 |
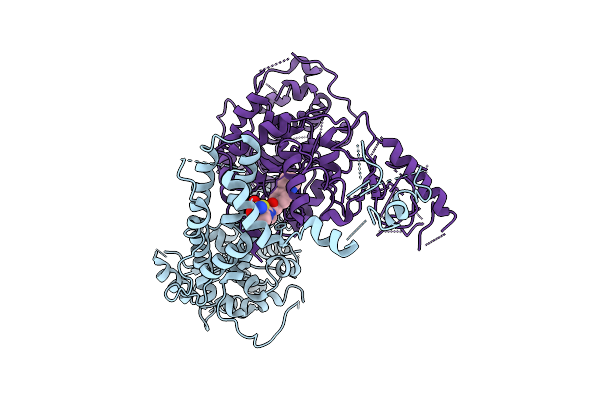
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE |
 |
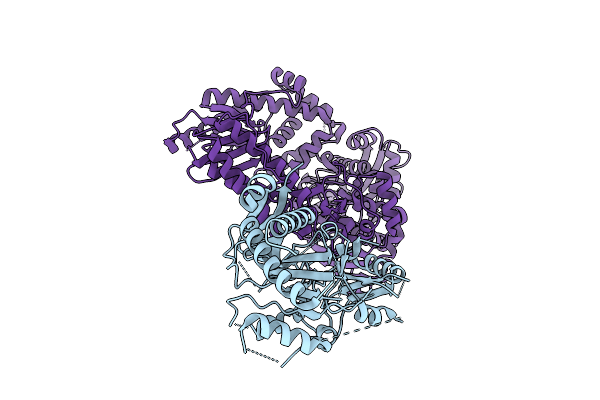
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex Prepared With Forked Dna, Atp-Gamma-S And Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
 |
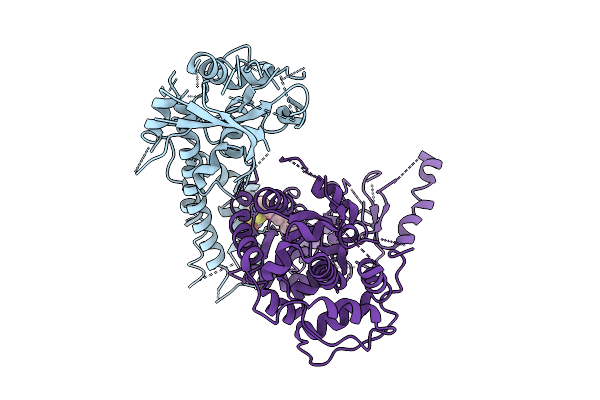
The Rigid Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR |
 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Pritelivir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXB |
 |
The Flexible Portion Of Cryo-Em Structure Of Herpesvirus Helicase-Primase Complex With Amenamevir
Organism: Herpesviridae
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: REPLICATION, TRANSFERASE/INHIBITOR Ligands: A1BXD |
 |
Crystal Structure Of Acyl-Coa Dehydrogenase Fade2 Mutant (Pa0508 M130G E296A Q303A) From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2025-02-19 Classification: OXIDOREDUCTASE Ligands: FAD, GOL, PGE |
 |
Crystal Structure Of The Acyl-Coa Dehydrogenase Fade1(Pa0506) E441A From Pseudomonas Aeruginosa Complexed With C16Coa
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.44 Å Release Date: 2025-02-12 Classification: OXIDOREDUCTASE Ligands: PKZ, EDO, PGE, NO3, TRS |
 |
Crystal Structure Of The Apo Acyl-Coa Dehydrogenase Fade2 (Pa0508) From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2025-01-15 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, 1PE, GOL |
 |
Crystal Structure Of The Acyl-Coa Dehydrogenase Pa0506 (Fade1) From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2025-01-15 Classification: OXIDOREDUCTASE Ligands: FAD, PEG, CO8 |
 |
Organism: Thermococcus kodakarensis
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-11-06 Classification: RNA BINDING PROTEIN |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2023-02-01 Classification: TRANSFERASE Ligands: AU |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-02-01 Classification: TRANSFERASE Ligands: AU |

