Search Count: 62
All
Selected
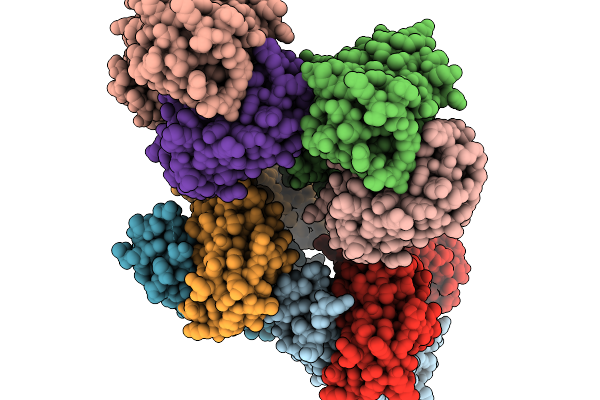 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
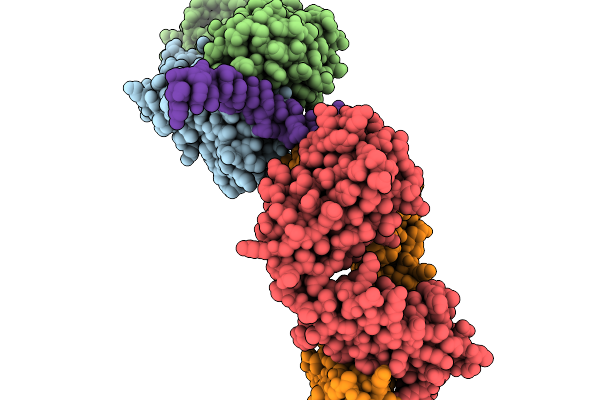 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.27 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM |
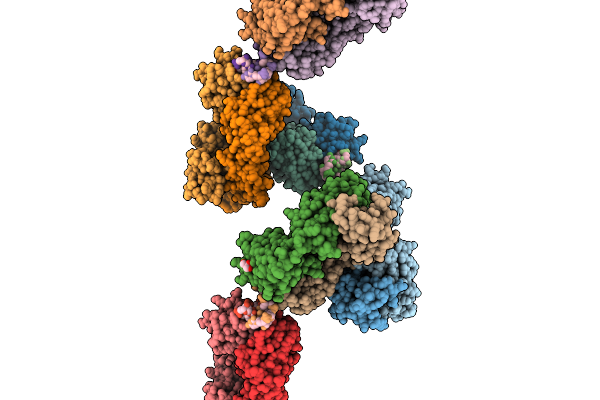 |
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-02-18 Classification: IMMUNE SYSTEM Ligands: GOL |
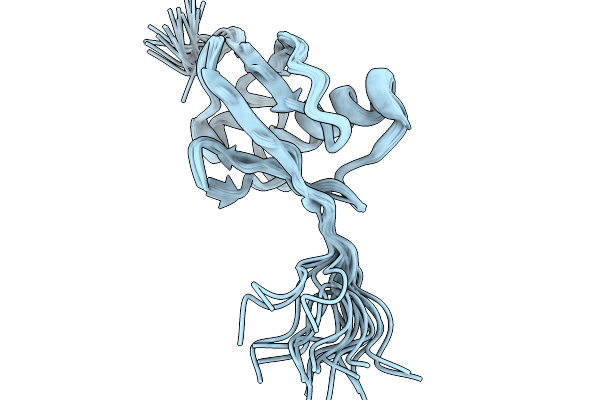 |
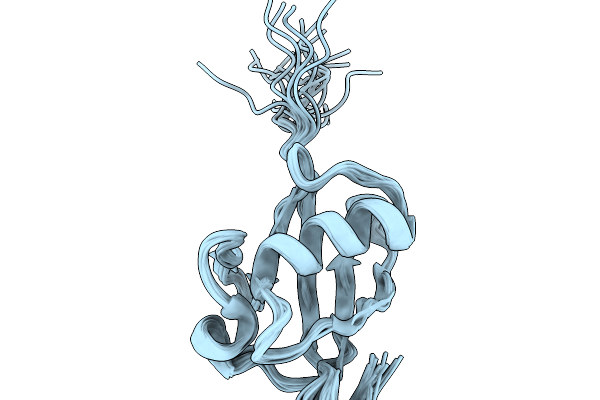
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Nmr Structure Of Proteinmpnn-Desighed Ubiquitin Variant R4 At Ph 6.3 With 8 M Urea
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Escherichia coli bl21(de3)
Method: SOLUTION NMR Release Date: 2026-02-11 Classification: DE NOVO PROTEIN |
 |
Organism: Caulobacter vibrioides (strain atcc 19089 / cip 103742 / cb 15)
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-03-12 Classification: HYDROLASE |
 |
Crystal Structure Of Synthetic Ubiquitin Variant R10 Designed By Proteinmpnn
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-02-12 Classification: DE NOVO PROTEIN Ligands: GOL |
 |
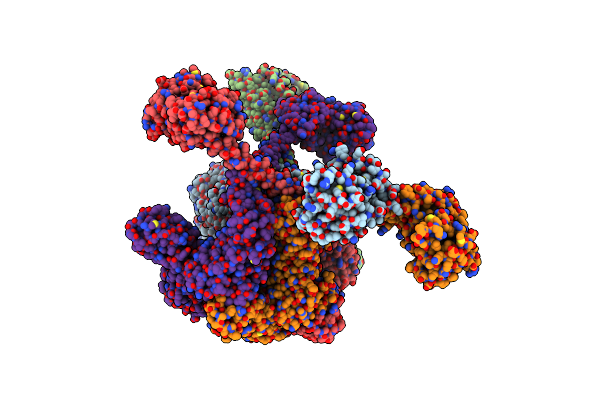
Crystal Structure Of Synthetic Ubiquitin Variant R4 Designed By Proteinmpnn
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2025-02-12 Classification: DE NOVO PROTEIN Ligands: SO4 |
 |
Open-Spiral Pentamer Of The Substrate-Free Lon Protease With A Y224S Mutation
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE Ligands: AGS |
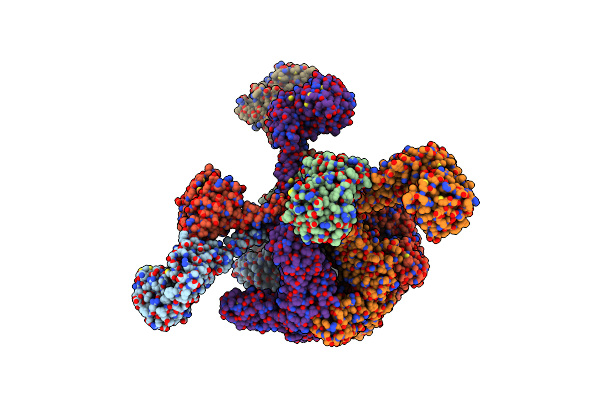 |
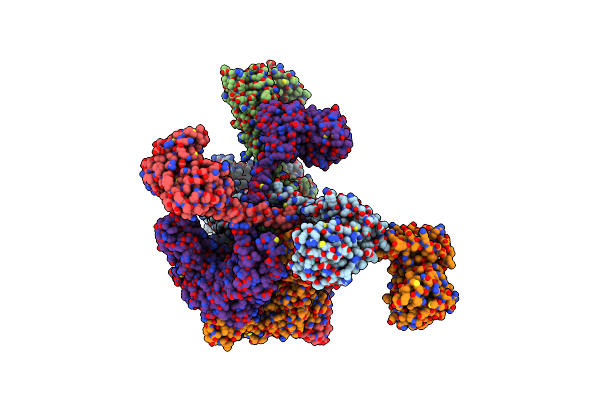
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE Ligands: AGS |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE Ligands: ADP |
 |
Close-Ring Hexamer Of The Substrate-Bound Lon Protease With An S678A Mutation
Organism: Meiothermus taiwanensis, Bos taurus
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE Ligands: ADP |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE |
 |
Organism: Meiothermus taiwanensis
Method: ELECTRON MICROSCOPY Release Date: 2023-10-25 Classification: HYDROLASE Ligands: ADP |

