Search Count: 253
All
Selected
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: PEG, GOL |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: GOL |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: GOL, PEG |
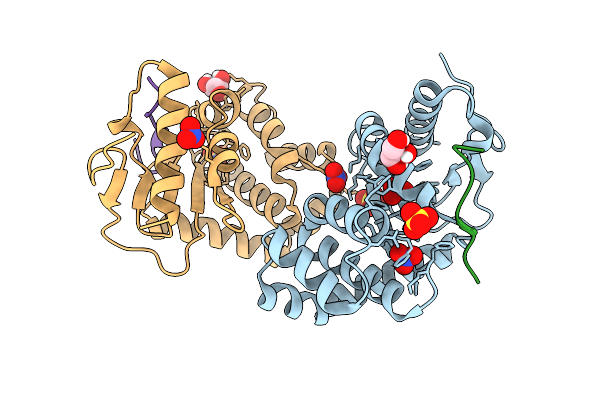 |
Crystal Structure Of Escherichia Coli Dsba C33A Mutant In Complex With A Peptide Derived From Lptd - Binding Mode I
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: GOL, NO3, SO4 |
 |
Crystal Structure Of Escherichia Coli Dsba C33A Mutant In Complex With A Peptide Derived From Lptd - Binding Mode Ii
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: GOL |
 |
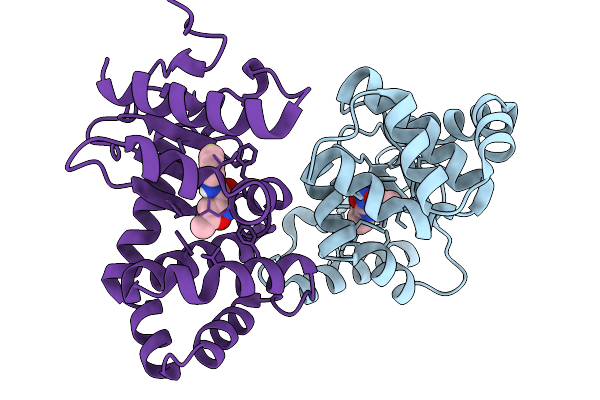
E.Coli Dsba In Complex With N-(2-Aminophenyl)-5-Methylisoxazole-3-Carboxamide
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: A1BX8, CU |
 |
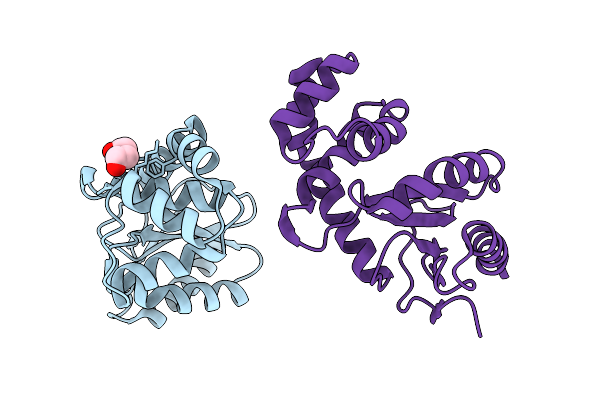
E.Coli Dsba In Complex With N-(2-Amino-3-Fluorophenyl)-5-Methylisoxazole-3-Carboxamide
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: A1BYA, CU |
 |
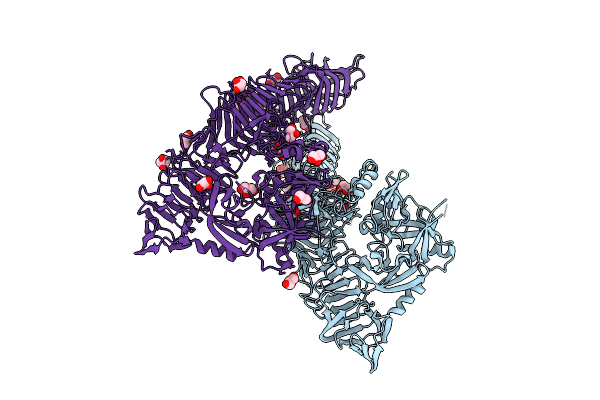
Organism: Bordetella pertussis
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE Ligands: PEG |
 |
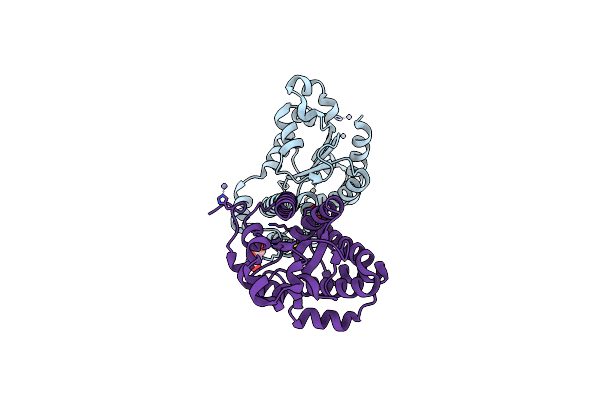
Organism: Escherichia coli o127:h6
Method: X-RAY DIFFRACTION Resolution:2.94 Å Release Date: 2025-10-29 Classification: HYDROLASE Ligands: P6G, PE8, GOL, PEG |
 |
Organism: Francisella tularensis subsp. tularensis schu s4
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2024-12-18 Classification: OXIDOREDUCTASE Ligands: ZN, PEG |
 |
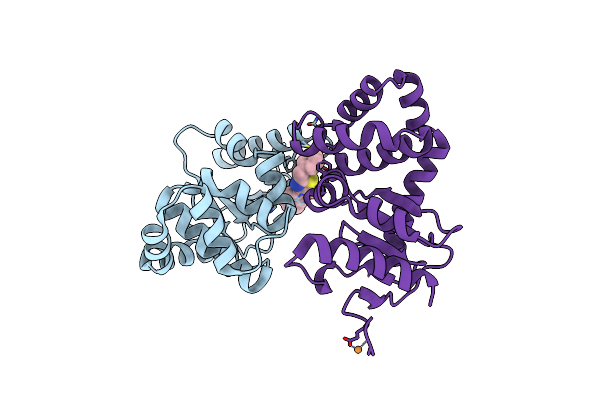
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2024-12-11 Classification: OXIDOREDUCTASE Ligands: A1BH7 |
 |
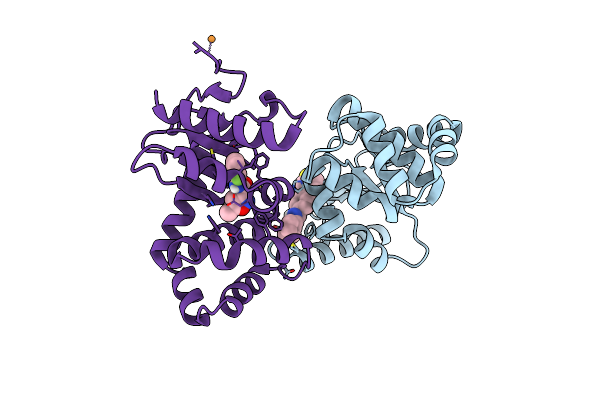
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2024-10-16 Classification: OXIDOREDUCTASE Ligands: W9H, CU |
 |
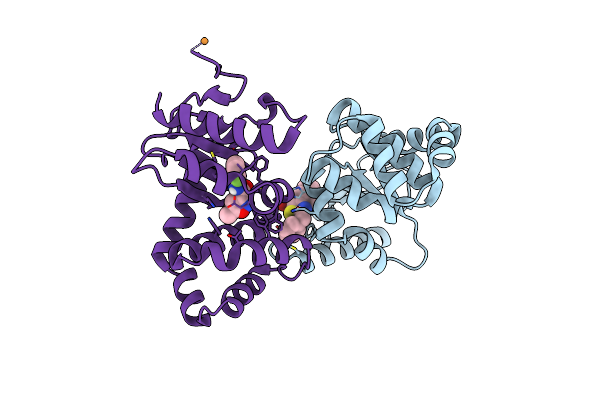
Ecdsba Soaked With N-(2-Fluorophenyl)-5-Methylisoxazole-3-Carboxamide And N-(4-(Thiophen-3-Yl)Benzyl)Cyclohexanamine
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: 5V9, SW0, CU |
 |
Ecdsba Soaked With N-(2-Fluorophenyl)-5-Methylisoxazole-3-Carboxamide And 2-Benzyl-4-Phenylthiazole-5-Carboxylic Acid
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: W9H, CU, SW0 |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2024-09-11 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: 5V9, CU |
 |
E.Coli Dsba In Complex With N-(2-Fluorophenyl)-5-Methylisoxazole-3-Carboxamide
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2023-06-21 Classification: OXIDOREDUCTASE/INHIBITOR Ligands: SW0, EDO, NA, CU |
 |
E.Coli Dsba In Complex With 4-Phenyl-2-(3-Phenylpropyl)Thiazole-5-Carboxylic Acid
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2023-04-12 Classification: OXIDOREDUCTASE/OXIDOREDUCTASE INHIBITOR Ligands: QVP, CU |
 |
Organism: Serratia marcescens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-03-08 Classification: HYDROLASE Ligands: PEG, GOL, IOD, CA |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2023-02-15 Classification: OXIDOREDUCTASE Ligands: GOL, CU |
 |
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2023-02-15 Classification: OXIDOREDUCTASE Ligands: GOL, CU |

