Search Count: 27
All
Selected
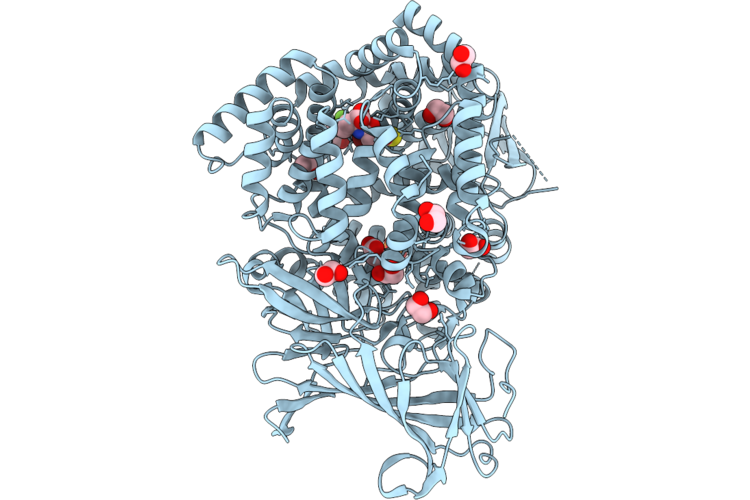 |
Erap1 In Complex With 1-[2-(5-Bromo-7-Fluoro-2-Oxo-2,3-Dihydro-1,3-Benzothiazol-3-Yl)Acetamido]-4,4-Difluorocyclohexane-1-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-05-06 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN, SIN, PO4, EDO, A1JVE |
 |
Erap1 In Complex With 1-[2-(6-Bromo-2,2-Difluoro-3-Oxo-3,4-Dihydro-2H-1,4-Benzoxazin-4-Yl)Acetamido]Cyclohexane-1-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-04-22 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN, PO4, EDO, A1JVD |
 |
Erap1 In Complex With 1-[2-(2-Oxo-5-Phenyl-2,3-Dihydro-1,3-Benzoxazol-3-Yl)Acetamido]Cyclohexane-1-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-04-22 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN, MLT, EDO, A1JVG |
 |
Erap1 In Complex With 1-[2-(5-Chloro-2-Oxo-2,3-Dihydro-1,3-Benzothiazol-3-Yl)Acetamido]Cyclohexane-1-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN, PO4, EDO, A1JVH |
 |
Erap1 In Complex With 1-[2-(3-Oxo-6-Phenyl-3,4-Dihydro-2H-1,4-Benzoxazin-4-Yl)Acetamido]Cyclohexane-1-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-04-22 Classification: PEPTIDE BINDING PROTEIN Ligands: ZN, PO4, EDO, A1JVF |
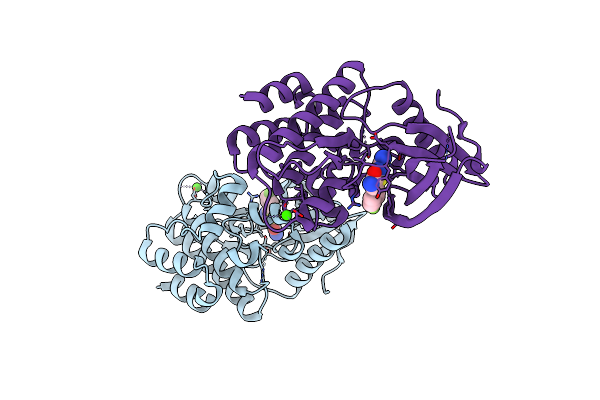 |
Human Tyk2 Pseudokinase Domain (575-869) In Complex With 5-(4-Fluoro-Phenyl)-2-Ureido-Thiophene-3-Carboxylic Acid Amide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2021-04-14 Classification: SIGNALING PROTEIN Ligands: NM7, CA |
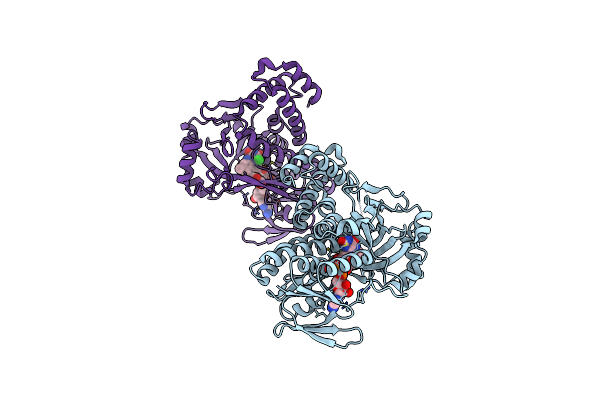 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, CL, GOL |
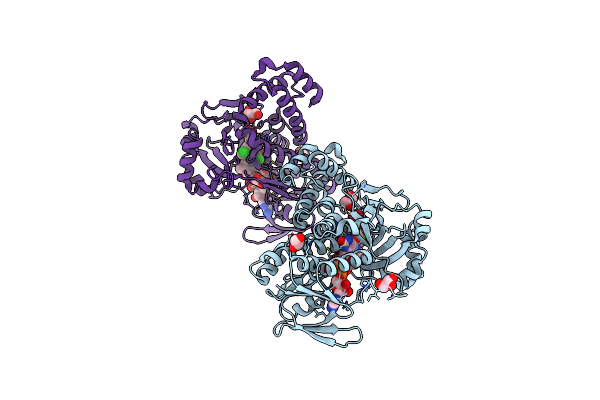 |
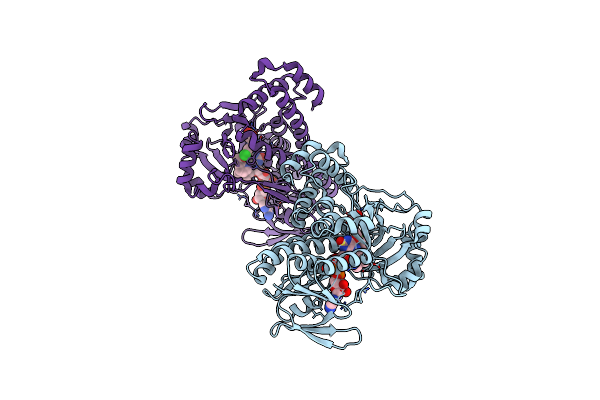
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-(5-Chloro-6-Methyl-2-Oxo-2,3-Dihydro-1,3-Benzoxazol-3-Yl)Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, CL, 8RK, GOL |
 |
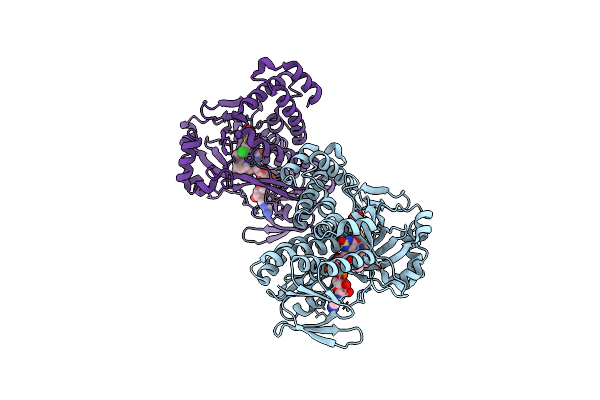
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-{5-Chloro-2-Oxo-6-[(1R)-1-(Pyridin-2-Yl)Ethoxy]-2,3-Dihydro-1,3-Benzoxazol-3-Yl}Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, 8R8 |
 |
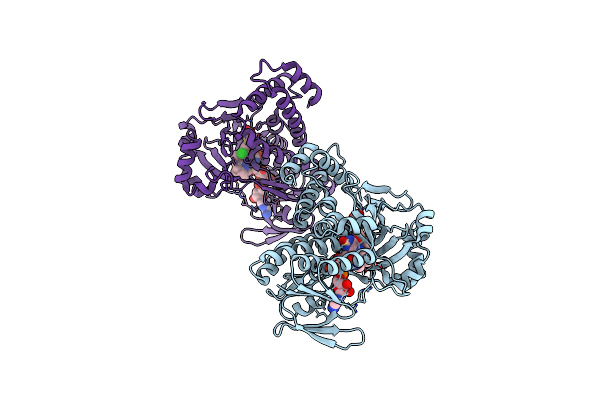
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-{5-Chloro-6-[(1R)-1-(Pyridin-2-Yl)Ethoxy]-1,2-Benzoxazol-3-Yl}Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, 8R5 |
 |
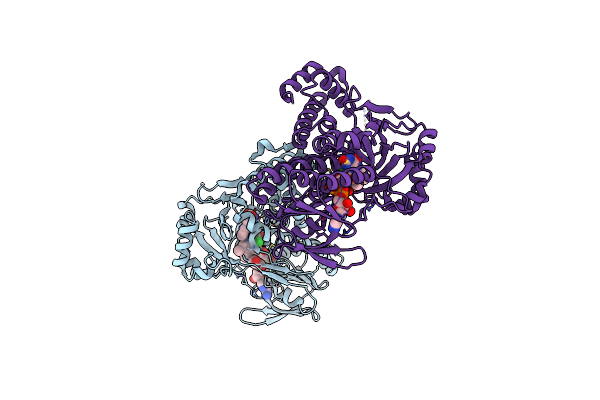
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-{5-Chloro-6-[(1R)-1-(6-Methylpyridazin-3-Yl)Ethoxy]-1,2-Benzoxazol-3-Yl}Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, 8RB |
 |
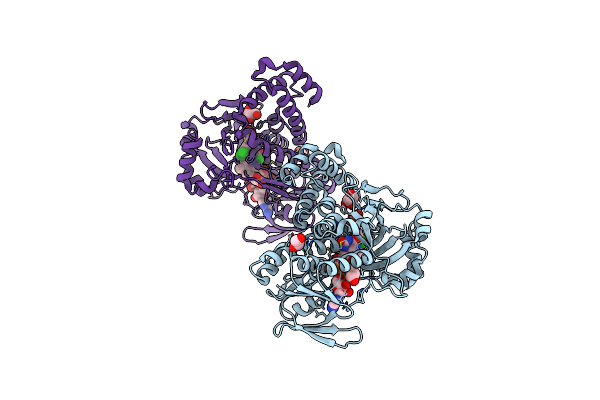
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With The Enzyme Substrate L-Kynurenine
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2017-06-21 Classification: OXIDOREDUCTASE Ligands: FAD, CL, GOL, KYN |
 |
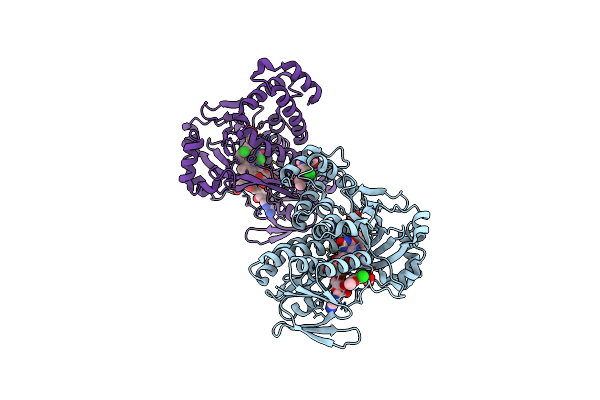
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-(5,6-Dichloro-2-Oxo-2,3-Dihydro-1,3-Benzoxazol-3-Yl)Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2017-06-14 Classification: OXIDOREDUCTASE Ligands: FAD, CL, JHY, GOL |
 |
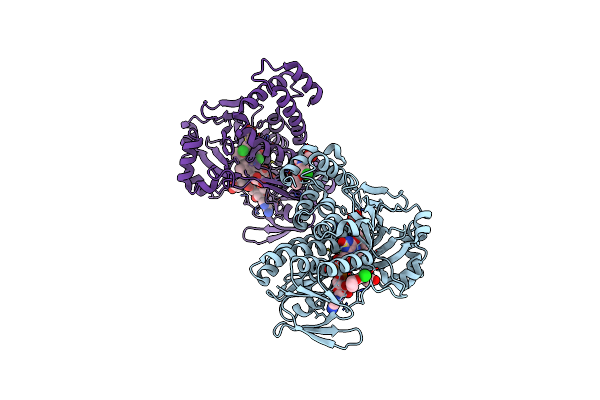
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-(5-Chloro-6-Ethoxy-2-Oxo-2,3-Dihydro-1,3-Benzoxazol-3-Yl)Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2017-04-19 Classification: OXIDOREDUCTASE Ligands: FAD, CL, 8EQ, GOL |
 |
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-(5-Chloro-6-Cyclopropoxy-2-Oxo-2,3-Dihydro-1,3-Benzoxazol-3-Yl)Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2017-04-19 Classification: OXIDOREDUCTASE Ligands: FAD, CL, FYK, GOL |
 |
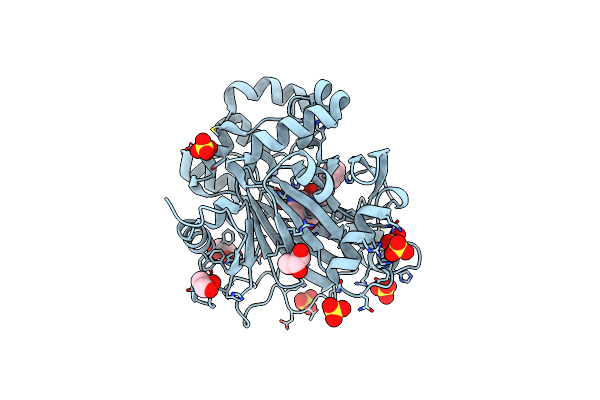
Pseudomonas Fluorescens Kynurenine 3-Monooxygenase (Kmo) In Complex With 3-[5-Chloro-6-(Cyclobutylmethoxy)-2-Oxo-2,3-Dihydro-1,3-Benzoxazol-3-Yl]Propanoic Acid
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2017-04-19 Classification: OXIDOREDUCTASE Ligands: FAD, CL, OK1, GOL |
 |
Crystal Structure Of Human Kdm4D In Complex With 3-4-Methylthiophen-2- Yl Methylaminopyridine-4-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2016-11-09 Classification: INHIBITOR Ligands: ZN, FE2, GOL, YC8, SO4 |
 |
Cell Penetrant Inhibitors Of The Jmjd2 (Kdm4) And Jarid1 (Kdm5) Families Of Histone Lysine Demethylases
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2016-01-27 Classification: OXIDOREDUCTASE Ligands: YC8, ZN, MG, BCN, CO |
 |
Crystal Structure Of Human Kdm4D In Complex With 3-(4- Phenylbutanamido)Pyridine-4-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2016-01-27 Classification: TRANSCRIPTION Ligands: ZN, FE2, GOL, YC8, SO4 |
 |
Crystal Structure Of Human Kdm4D In Complex With 3-4-Methylthiophen-2- Ylmethylaminopyridine-4-Carboxylic Acid
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2016-01-27 Classification: OXIDOREDUCTASE Ligands: ZN, CO, AUY, SO4 |

