Search Count: 27
All
Selected
 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.26 Å Release Date: 2026-01-28 Classification: SIGNALING PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-01-21 Classification: SIGNALING PROTEIN,TRANSFERASE Ligands: ZN, A1BUO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-01-21 Classification: SIGNALING PROTEIN Ligands: ZN, A1BVB |
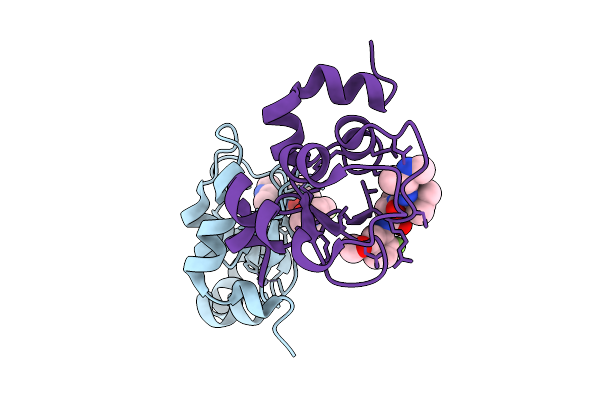 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-21 Classification: SIGNALING PROTEIN Ligands: ZN, A1BVC |
 |
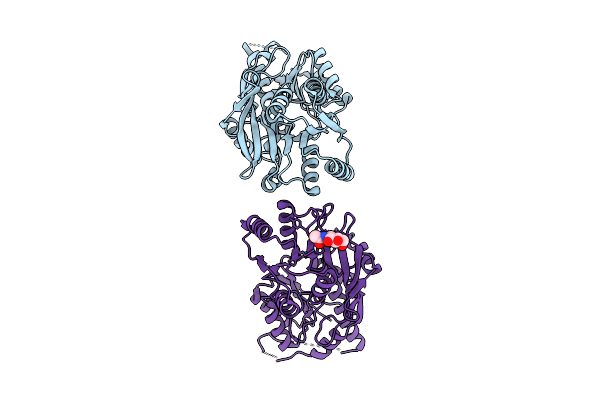
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.55 Å Release Date: 2025-12-31 Classification: SIGNALING PROTEIN Ligands: E2Q |
 |
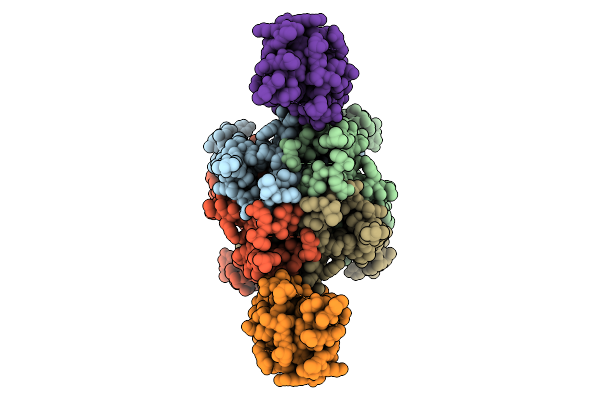
Cerebellar Glua2/4 Ntd Heterophilic Tetramer Interface (Focused Refinement)
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2025-12-24 Classification: SIGNALING PROTEIN Ligands: NAG |
 |
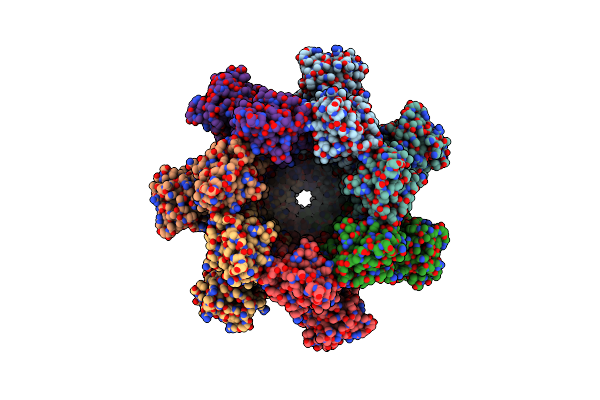
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.66 Å Release Date: 2025-12-24 Classification: SIGNALING PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2022-12-28 Classification: METAL BINDING PROTEIN Ligands: CA, PG4, IMD, ZN |
 |
1.68 Angstrom Crystal Structure Of Ca/Cam-E140G:Camkiidelta Peptide Complex
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2022-12-28 Classification: METAL BINDING PROTEIN Ligands: CA, GOL |
 |
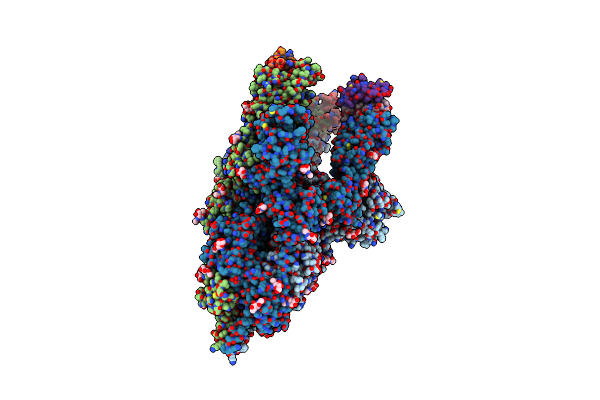
Cryo-Em Structure Of Complete Transmembrane Channel E289A Mutant Vibrio Cholerae Cytolysin
Organism: Vibrio cholerae
Method: ELECTRON MICROSCOPY Release Date: 2022-08-31 Classification: TOXIN |
 |
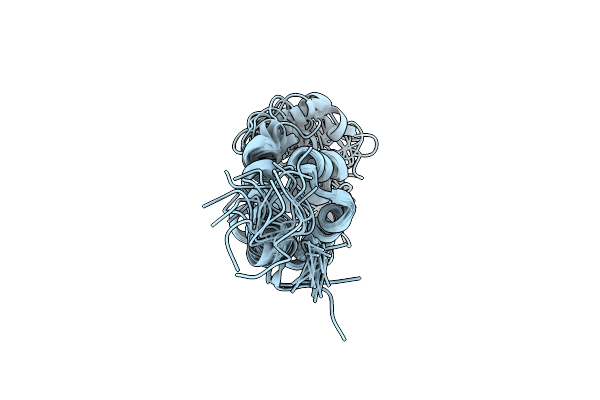
3D Model Of The 3-Rbd Up Single Trimeric Spike Protein Of Sars-Cov2 In The Presence Of Synthetic Peptide Sih-5.
Organism: Severe acute respiratory syndrome coronavirus 2, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-04-27 Classification: VIRAL PROTEIN Ligands: NAG |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2021-08-04 Classification: MEMBRANE PROTEIN |
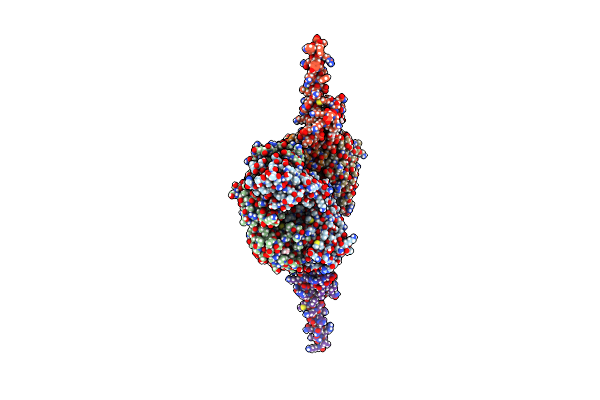 |
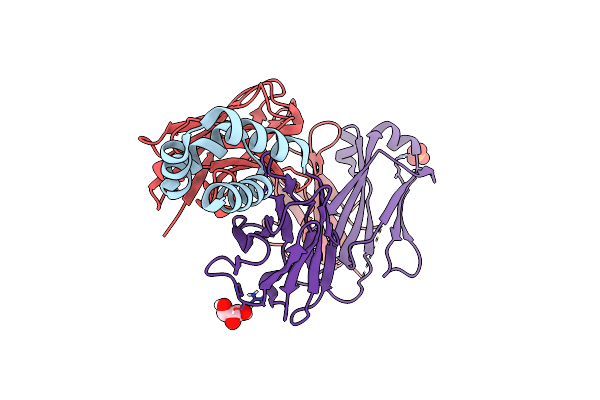
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2021-08-04 Classification: IMMUNE SYSTEM Ligands: EOH, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2021-08-04 Classification: IMMUNE SYSTEM Ligands: PO4, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2021-08-04 Classification: IMMUNE SYSTEM Ligands: GOL, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2021-08-04 Classification: IMMUNE SYSTEM Ligands: GOL, MES, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2021-08-04 Classification: IMMUNE SYSTEM Ligands: FLC, EOH, GOL, NH4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2021-02-10 Classification: METAL BINDING PROTEIN Ligands: CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2021-02-10 Classification: METAL BINDING PROTEIN Ligands: CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2021-02-10 Classification: METAL BINDING PROTEIN Ligands: CA |

