Search Count: 132
All
Selected
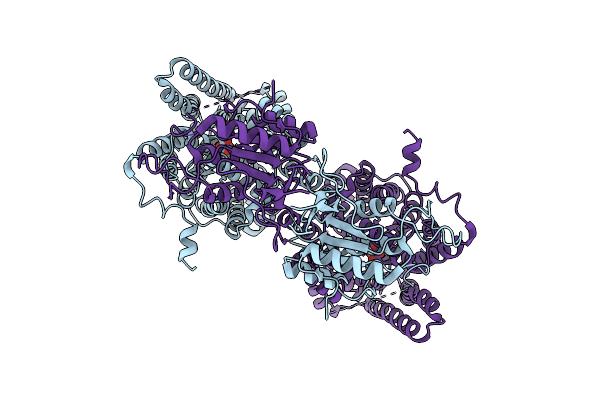 |
Cryo-Em Structure Of Serendipita Indica Sulfate Transporter Sisult In Sulphate Bound State
Organism: Serendipita indica
Method: ELECTRON MICROSCOPY Resolution:2.44 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: SO4 |
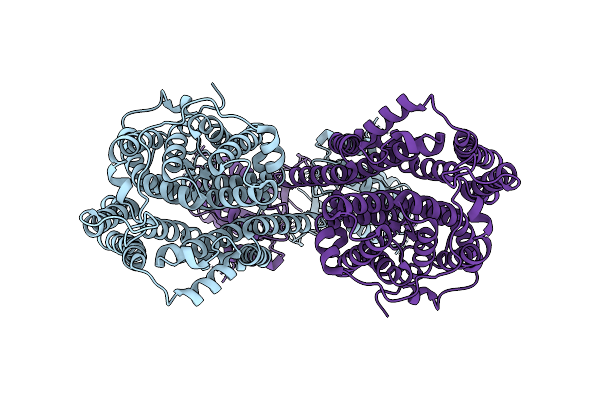 |
Cryo-Em Structure Of Serendipita Indica Sulfate Transporter Sisult In Apo State
Organism: Serendipita indica
Method: ELECTRON MICROSCOPY Resolution:2.96 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN |
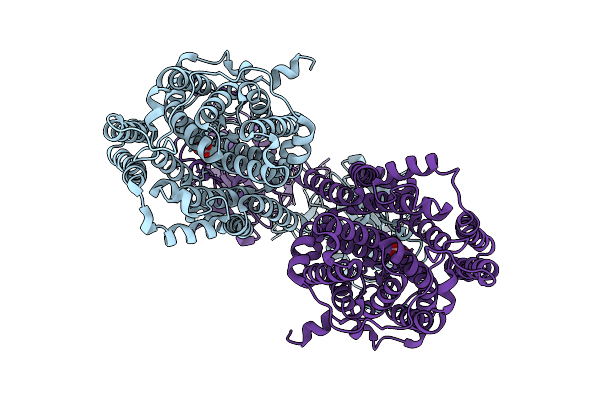 |
Cryo-Em Structure Of Serendipita Indica Sulfate Transporter Sisult In Oxlate Bound State
Organism: Serendipita indica
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-03-11 Classification: MEMBRANE PROTEIN Ligands: OXL |
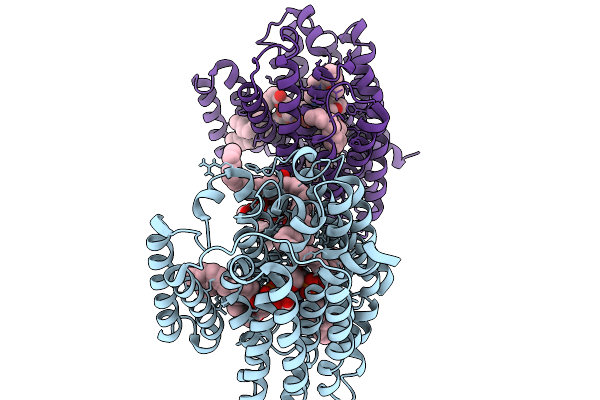 |
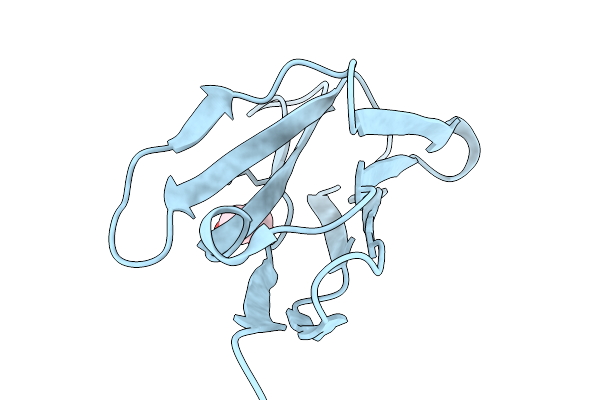
Organism: Mycobacterium tuberculosis h37rv
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: A1ATG, A1BXT, A1BXW |
 |
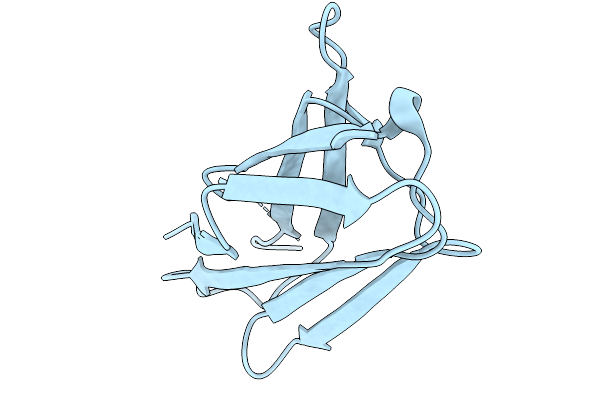
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM Ligands: EDO |
 |
Organism: Chiloscyllium plagiosum
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-07 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.13 Å Release Date: 2025-12-17 Classification: TRANSPORT PROTEIN Ligands: A1CAN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.33 Å Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN |
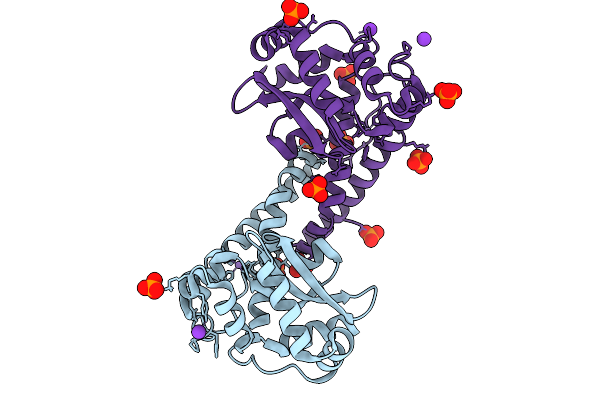 |
Finding The Exit Route Of Hydrogen Peroxide From The Manganese Superoxide Dismutase (Mnsod) Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: K, PO4, MN |
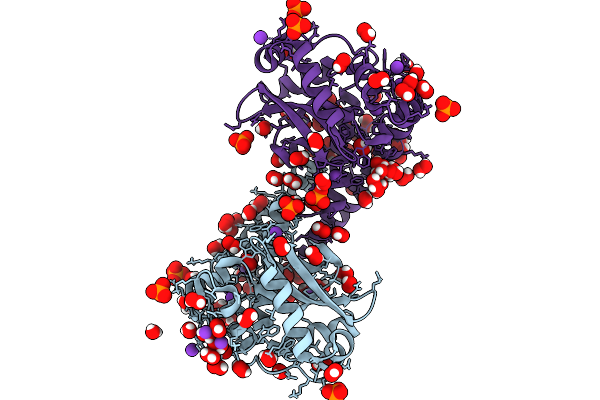 |
Finding The Exit Route Of Hydrogen Peroxide From The Manganese Superoxide Dismutase (Mnsod) Active Site
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: PEO, PO4, K, MN |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0A).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, EDO |
 |
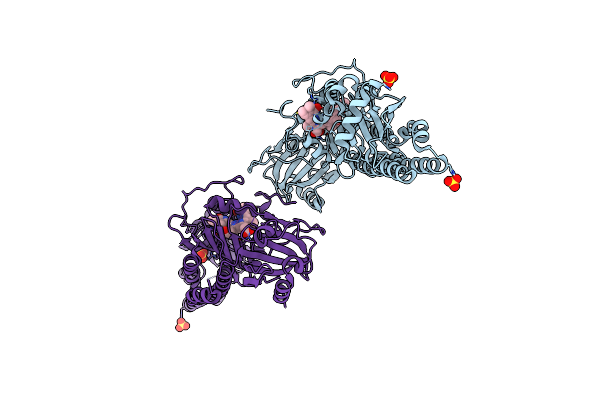
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0B).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
 |
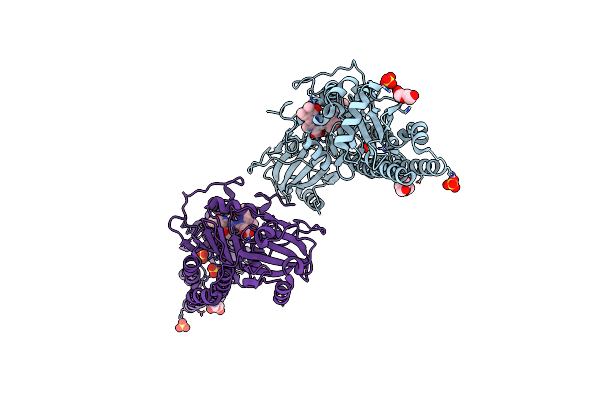
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I1 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I2 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PGE, PEG, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I3 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, GOL, PEG |
 |
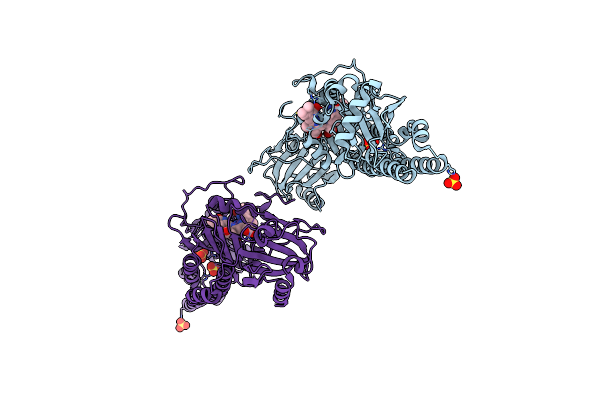
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I4 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, PEG |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I5 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I6 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
 |
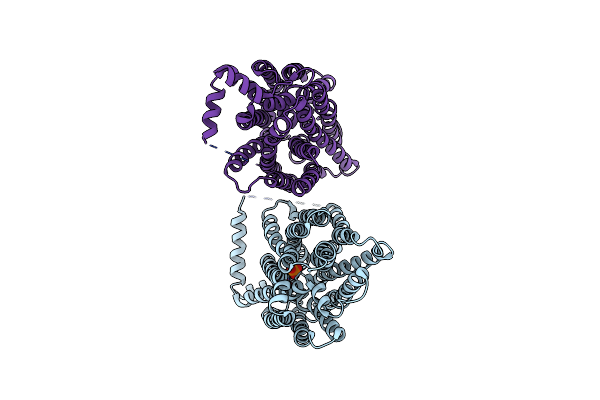
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I7 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PEG |
 |
Organism: Serendipita indica
Method: X-RAY DIFFRACTION Resolution:3.70 Å Release Date: 2025-03-26 Classification: TRANSPORT PROTEIN Ligands: PO4 |

