Search Count: 664
All
Selected
 |
A Novel Histone H3K27 Reader, Cbfa2T2, Inhibits H3K27Me3 Demethylation And Tumor Growth By Regulating Metabolic Genes And Metabolite Levels
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.81 Å Release Date: 2026-04-22 Classification: ANTITUMOR PROTEIN |
 |
Crystal Structure Of Terpeniod Cyclase Spsods From Rhizobacterium Serratia Plymuthica
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:2.22 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GOL |
 |
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-03-18 Classification: LYASE Ligands: POP, MG, TRS |
 |
Organism: Serratia plymuthica 4rx13
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-18 Classification: LYASE Ligands: GPP, MG, TRS, POP |
 |
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, IMD, PEG, TRS, CL |
 |
Crystal Structure Of A P450 Bm3 Heme Domain Mutant In Complex With Zearalenone
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, ZER, TRS |
 |
Crystal Structure Of A P450 Bm3 Heme Domain Mutant In Complex With Alpha-Zearalanol
Organism: Priestia megaterium (strain nbrc 15308)
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: HEM, 36J, PEG, NI, TRS |
 |
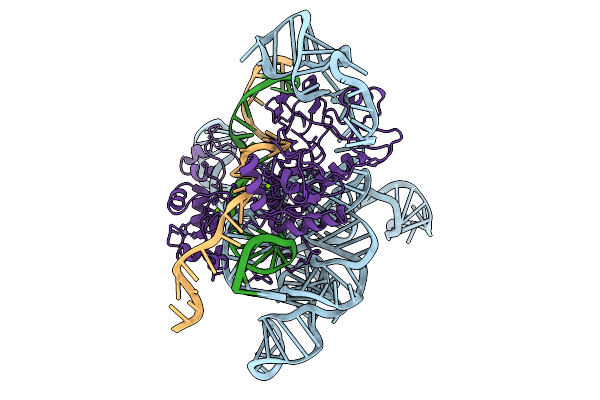
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.04 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
 |
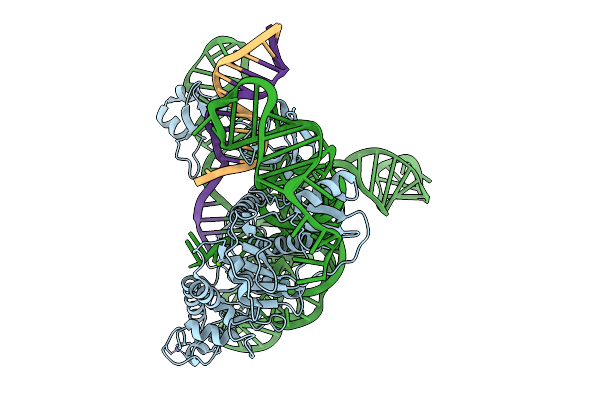
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: MG, ZN |
 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2026-01-21 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
 |
Cryoem Structures Uncover The Unexpected Hinges Of Iscb For Enhanced Gene Editing
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.09 Å Release Date: 2026-01-21 Classification: DNA BINDING PROTEIN Ligands: ZN, MG |
 |
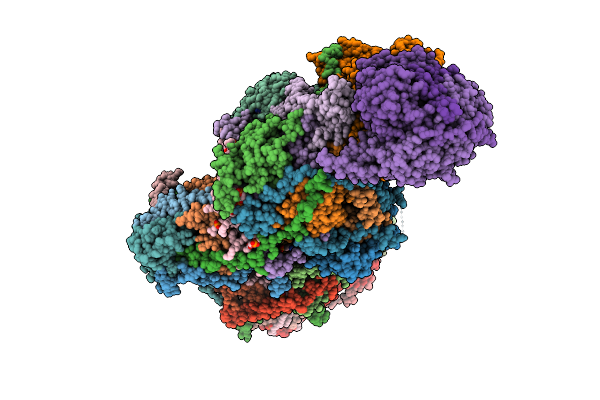 |
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: MEMBRANE PROTEIN Ligands: CDL, PEE, NDP, ZMP, ADP, 3PE, PC1, PLX, XEW, SF4, FES, MG, ZN, FMN |
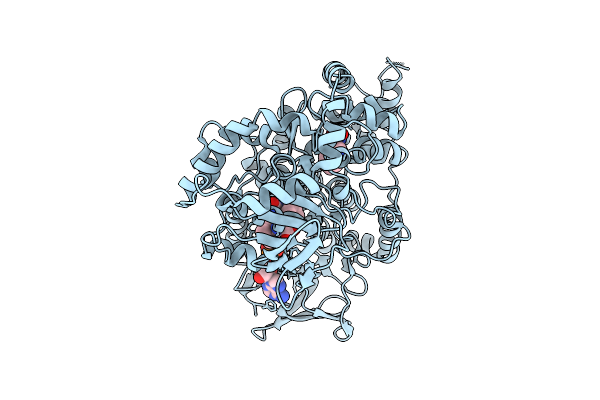 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: FAD, TRP |
 |
Crystal Structure Of Aetf-L183F/V220I/S523A In Complex With Fad And L-Tryptophan
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-12-10 Classification: FLAVOPROTEIN Ligands: TRP, FAD |
 |
 |
 |
Complex I From Respirasome Closed State 1 Bound By Metformin And Coq10, Alternative Orientation (Sc-Metc1-Ii)
Organism: Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: MEMBRANE PROTEIN Ligands: CDL, PC1, NDP, PEE, ZMP, ADP, PLX, 3PE, PX2, U10, SF4, FES, MG, MF8, ZN, FMN |

